Abstract
Staphylococci were isolated from veterinary staff, hospitalized animals, and medical equipment from 2 major tertiary veterinary hospitals in South Korea to investigate antimicrobial resistance and genetic relatedness. The detection rate for staphylococci was 55.2% (111/201 samples), and 11 species were identified among the collected staphylococcal strains. The most prevalent species were Staphylococcus pseudintermedius (52/111, 46.8%), Staphylococcus epidermidis (21/111, 18.9%), and Staphylococcus aureus (19/111, 17.1%). The methicillin-resistance rates of staphylococci isolated from veterinary staff and medical equipment were higher than those from hospitalized animals. The genotype of methicillin-resistant S. aureus (MRSA) strains in the current study was sequence type (ST)72-SCCmec IVc-t324, which is similar to the genotype of prevalent MRSA strains in human beings and food animals in South Korea. Among the mecA-positive S. pseudintermedius isolates, SCCmec V was most prevalent in strains originating from both veterinary staff and hospitalized animals. SCCmec IVa was detected in methicillin-resistant S. epidermidis, whereas SCCmec IVc was found in other methicillin-resistant, coagulase-negative staphylococci. The SCCmec typing, antimicrobial susceptibility tests, and pulsed field gel electrophoresis results showed that methicillin-resistant staphylococci dissemination between hospitalized animals and veterinary staff is possible in South Korean veterinary hospitals.
Introduction
Methicillin-resistant staphylococci (MRS) are important zoonotic pathogens that cause serious public health problems by increasing both the failure rate of antibiotic therapy and mortality rates among human beings and animals. 4 The microorganisms can colonize the skin or mucous membranes, or attach to the surface of medical equipment, and cause a variety of pathologic complications ranging from mild infections to severe systemic diseases. 4 The β-lactam resistance of MRS is due to penicillin-binding protein (PBP2a), which is encoded by the methicillin-resistant gene, mecA. The mecA gene is located in the staphylococcal cassette chromosome mec (SCCmec), a 21- to 67-kb mobile genetic element that can be transferred between different species of staphylococci. 13
SCCmec types I–III are usually detected in hospital-associated, methicillin-resistant Staphylococcus aureus (MRSA), whereas SCCmec types IV and V are generally detected in community-associated MRSA. 21 However, several MRSA strains harboring SCCmec IV are frequently detected in nosocomial as well as community infections worldwide—for example, USA300 in the United States, EMRSA-15 in the United Kingdom, and sequence type (ST)72 clones in South Korea.8,20,25 In addition, SCCmec IV has the most diverse subtypes (IVa–h) 25 among the SCCmec types. Various SCCmec types also have been reported among methicillin-resistant, coagulase-negative staphylococci (MR-CNS) as well as MRSA. 13 Despite the importance of monitoring MR-CNS, veterinary medicine studies have been primarily limited to coagulase-positive staphylococci (CPS) such as Staphylococcus aureus, Staphylococcus intermedius, and Staphylococcus pseudintermedius. 12
There are growing concerns about the direct dissemination of MRS between human beings and companion animals through contact and the emergence of new MRS strains resulting from SCCmec transfer between staphylococci. 1 In several reports, antibiotic-resistant bacteria in cats and dogs were found to be transmitted to owners and veterinary staff or vice versa.6,14 Veterinary hospitals could be significant sources of pathogen dissemination between veterinary staff and hospitalized animals from direct human–animal contact and indirect exposure to contaminated environments or medical equipment.
Preventing the dissemination of MRS in veterinary hospitals requires better knowledge of MRS distribution. Thus, in the present study, staphylococci were collected from veterinary staff, hospitalized animals, and medical equipment in tertiary veterinary hospitals in South Korea. The genetic and phenotypic characteristics were evaluated based on antibiotic susceptibility tests, pulsed field gel electrophoresis (PFGE) analysis, multilocus sequence typing (MLST), SCCmec typing, and spa typing.
Materials and methods
Bacterial isolation and identification
Staphylococci were collected from 2 tertiary veterinary hospitals—1 veterinary teaching hospital and 1 referral hospital—in Seoul and Gyeonggi provinces (South Korea) between March and April 2008. A total of 201 swab samples were taken from 37 veterinary staff (37 nasal cavity and 37 hand samples), 18 hospitalized animals (31 ear, 17 skin, 17 anus, 14 nasal mucosa, 4 foot pad, 1 vulva, and 2 infected wound samples), and 41 different locations in the hospitals (door handles, telephones, floors, tables, computers in the veterinarians’ offices, cages, and medical devices). Medical devices and other objects that frequently had contact with the veterinary staff and hospitalized animals were selected for sampling. All samples were collected using culture swabs a and were kept refrigerated during transfer to the laboratory.
Swabs were streaked on 5% sheep blood agar b and incubated for 24–48 hr at 37°C. After incubation, 1 representative staphylococci-like colony was collected. The selected colonies were identified to the species level using an automated microbial identification system. c The results were confirmed by species-specific polymerase chain reaction (PCR) and sequencing analysis of the sodA gene for CNS. 31 Staphylococcus pseudintermedius was identified using a PCR–restriction fragment length polymorphism approach. 2 If different species or genotypes were identified within 1 sample, then the staphylococci isolates collected from that sample were considered to be different strains.
Antibiotic susceptibility tests
Antibiotic susceptibility tests were carried out using the disk diffusion method according to the Clinical Laboratory Standard Institute guidelines. 9 Ten antibiotics d were used: oxacillin, ampicillin, amoxicillin–clavulanic acid, erythromycin, gentamicin, amikacin, tetracycline, vancomycin, sulfamethoxazole–trimethoprim, and ciprofloxacin. The minimum inhibitory concentration (MIC) of oxacillin e was determined using a broth microdilution method 9 and was used to identify MRS, which were defined in the present study as strains showing MIC ≥4 µg/ml (for S. aureus and S. pseudintermedius) or MIC ≥0.5 µg/ml (for other staphylococci). For quality control, S. aureus ATCC (American Type Culture Collection) 25923 was used in the disk diffusion tests, and Escherichia coli ATCC 25922 and Enterococcus faecalis ATCC 29212 were used in the microdilution tests.
Methicillin-resistant staphylococci genotyping
Genomic DNA was isolated using a commercial DNA extraction kit f according to the manufacturer’s instructions with a modified cell lysis step performed with 50 U/ml of lysostaphin. g Identifications of MRS strains were confirmed by mecA amplification, and the SCCmec type was determined with 3 multiplex PCR strategies. The first multiplex PCR identified the mec class (A, B, or C), whereas the second determined the type of ccr complex genes (ccrAB1, ccrAB2, ccrAB3, ccrAB4, or ccrC). 22 The third multiplex PCR was used for SCCmec IV subtyping (IVa–h). 25
Multilocus sequence typing analysis was performed for S. aureus 10 and Staphylococcus epidermidis. 26 Pulsed field gel electrophoresis was performed as previously described to genotype all MRS strains. 23 The PFGE patterns were analyzed by commercial software, h and the percent similarity was expressed as a Dice coefficient at a 2% tolerance and 2% optimization. Spa typing 16 was performed on MRSA strains, and the results were analyzed by commercial software. i A based upon repeat pattern algorithm was used to group the spa types.
Statistical analysis
Statistical analyses were performed using SPSS 12.0 for Windows. j The prevalence of staphylococci according to staphylococcal species and origin, the comparison of MRS detection rates according to coagulase-negative and -positive results, and the comparison of multidrug resistance rates according to methicillin susceptibility were calculated using χ2 tests for categorical variables. P values <0.01 were considered statistically significant.
Results
Distribution of staphylococci in veterinary hospital settings
A total of 201 samples were collected from veterinary staff (n = 86), hospitalized animals (n = 74), and medical equipment (n = 41), with staphylococci detected in 111 samples (55.2%). Among the 111 staphylococcal isolates, 11 species were identified: S. aureus, S. epidermidis, S. pseudintermedius, S. intermedius, Staphylococcus saprophyticus, Staphylococcus warneri, Staphylococcus lentus, Staphylococcus cohnii subsp. urealyticus, Staphylococcus auricularis, Staphylococcus simulans, and Staphyloccis vitulinus (Table 1). The most prevalent species were S. pseudintermedius (52/111 strains, 46.8%), S. epidermidis (21/111 strains, 18.9%), and S. aureus (19/111 strains, 17.1%). Staphylococci were isolated at a significantly higher (P < 0.001) rate from veterinary staff (60 strains from 86 samples, 81.1%) than from medical equipment (18 strains from 41 samples, 43.9%) or hospitalized animals (33 strains from 86 samples, 38.4%). Significantly more (P < 0.001) species were observed on veterinary staff (9 species) than on hospitalized animals (6 species) or medical equipment (5 species).
Staphylococcal isolates and methicillin-resistant staphylococci (MRS) strains isolated from 2 veterinary hospitals in South Korea.*
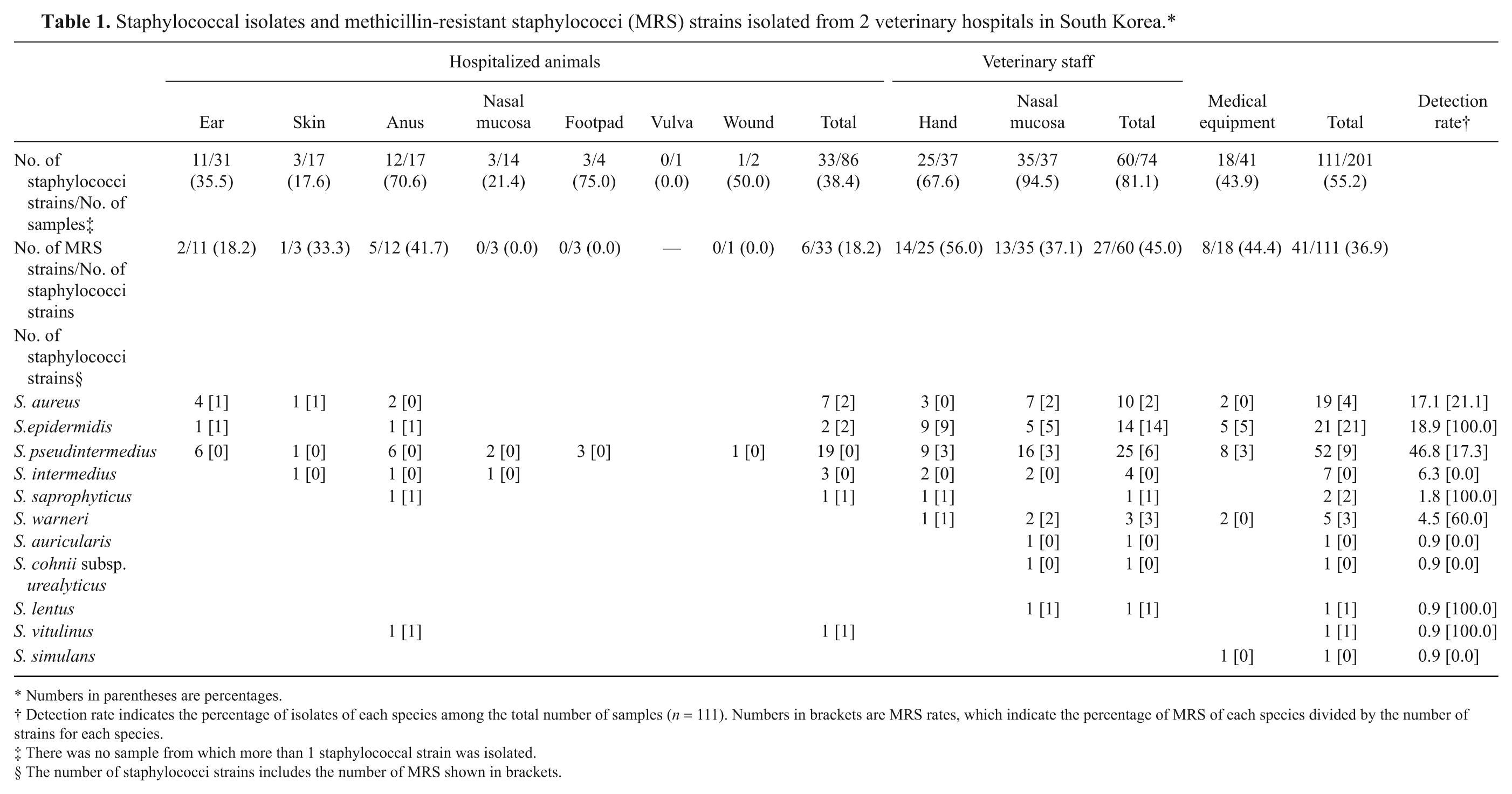
Numbers in parentheses are percentages.
Detection rate indicates the percentage of isolates of each species among the total number of samples (n = 111). Numbers in brackets are MRS rates, which indicate the percentage of MRS of each species divided by the number of strains for each species.
There was no sample from which more than 1 staphylococcal strain was isolated.
The number of staphylococci strains includes the number of MRS shown in brackets.
Methicillin-resistant staphylococci detection in veterinary hospital settings
The MRS detection rates are summarized in Table 1. Among the 111 staphylococcal isolates, 41 strains (36.9%) were identified as MRS. Based on origin, the methicillin-resistance rates of staphylococci isolated from veterinary staff (27/60 strains, 45.0%) and medical equipment (8/18 strains, 44.4%) were significantly higher (P < 0.001) than that from hospitalized animal staphylococci (6/33 strains, 18.2%).
Methicillin-resistant staphylococci were detected in the following 7 species: S. aureus, S. epidermidis, S. pseudintermedius, S. saprophyticus, S. warneri, S. lentus, and S. vitulinus. Higher methicillin-resistance rates were observed among CNS compared with CPS (P < 0.001). For example, S. epidermidis (21/21 strains, 100.0%), S. saprophyticus (2/2 strains, 100.0%), S. warneri (3/5 strains, 60.0%), S. lentus (1/1 strain, 100.0%), and S. vitulinus (1/1 strain, 100.0%) showed higher methicillin-resistance rates than S. pseudintermedius (9/52 strains, 17.3%) and S. aureus (4/19, 21.1%).
There were 25 strains of S. pseudintermedius that had the mecA gene. However, only 9 strains were resistant to oxacillin, whereas the other 16 strains were oxacillin susceptible. The latter have been referred to as oxacillin-susceptible, mecA-positive S. pseudintermedius. 15 These strains were isolated from hospitalized animals (7 strains), veterinary staff (6 strains), and medical equipment (3 strains) in the current study.
Antibiotic resistance
Among the 111 staphylococcal isolates, 49 different antibiotic-resistance patterns and 47 multidrug-resistant isolates (defined as isolates resistant to more than 3 antibiotic subclasses) were observed (Table 2). No vancomycin-resistant staphylococci were identified. Most of the 41 MRS strains were resistant to β-lactam antibiotics, including ceftiofur (97.6%) and amoxicillin–clavulanic acid (85.4%), erythromycin (65.9%), gentamicin (46.5%), and tetracycline (41.5%).
Antibiotic resistance patterns of staphylococci isolated from 2 veterinary hospitals in South Korea.*

Multidrug-resistant (MDR) strains, defined as strains resistant to more than 3 antibiotic subclasses tested in the current study, are indicated by bold text. MSS = methicillin-sensitive staphylococci, including oxacillin-susceptible, mecA-positive Staphylococcus pseudintermedius; MRS = methicillin-resistant staphylococci; AN = amikacin; AmC = amoxicillin–clavulanic acid; C = chloramphenicol; CC = clindamycin; CFT = ceftiofur; CIP = ciprofloxacin; E = erythromycin; GM = gentamicin; OX = oxacillin; SXT = sulfonamide/trimethoprim; TE = tetracycline; SA = Staphylococcus aureus; SE = Staphylococcus epidermidis; SP = Staphylococcus pseudintermedius; SI = Staphylococcus intermedius; SW = Staphylococcus warneri; SS = Staphylococcus saprophyticus; SC = Staphylococcus cohnii subsp. urealyticus; SAC = Staphylococcus auricularis; SL = Staphylococcus lentus; SV = Staphylococcus vitulinus.
Numbers in parentheses are percentages.
There were statistical differences (P < 0.001) between the MDR rates of oxacillin-resistant staphylococci and oxacillin-sensitive staphylococci.
At the species level, S. pseudintermedius showed the highest multidrug resistance rate (30/47 strains, 63.8%), followed by S. epidermidis (15/47 strains, 31.9%). In comparing MRS and methicillin-sensitive staphylococci (MSS), the multidrug resistance rate of MRS was significantly higher (P < 0.002) than that of MSS (Table 2). Although the multidrug-resistant MRS included 3 species—S. epidermidis, S. warneri, and S. pseudintermedius—the multidrug-resistant MSS was composed of only 1 species, S. pseudintermedius (Table 2). The multidrug resistance rates were not significantly different between the isolates from veterinary staff (26/60 strains, 43.3%), medical equipment (8/18 strains, 44.4%), and hospitalized animals (13/33 strains, 39.4%).
Genetic comparison of methicillin-resistant staphylococci isolates
The genotype of 57 mecA-positive staphylococci strains were compared by SCCmec typing, MLST, PFGE analysis, and spa typing. Four MRSA isolates, including 2 from veterinary staff and 2 from hospitalized animals, were identified as ST72-SCCmec IVc-t324. The PFGE pulsotype of these isolates had 94.0–100.0% similarity (Fig. 1A) even though they were isolated from different hospitals. A total of 25 mecA-positive S. pseudintermedius isolates, including 9 MRSP strains and 16 oxacillin-susceptible, mecA-positive S. pseudintermedius strains, were compared by PFGE (Fig. 1B). Thirteen PFGE pulsotypes and 3 SCCmec types, including V (15 strains), IV (1 strain), and nontypeable (9 strains), were observed. The genetic features of the strains isolated from the 2 hospitals differed. The 10 strains isolated from veterinary hospital A contained 9 PFGE patterns, whereas the 15 strains from veterinary hospital B consisted of 3 similar clones according to antibiotic resistance patterns, PFGE patterns, and SCCmec typing. Among these 3 clones, 2 were isolated from both veterinary staff and hospitalized animals. For example, the oxacillin-susceptible, mecA-positive S. pseudintermedius snu 98 strain isolated from veterinary staff and MRSP snu 120 strain isolated from hospitalized animals had indistinguishable PFGE pulsotypes and identical SCCmec type (SCCmec V) and antibiotic resistance pattern (TE-CIP-E-CC). In addition, strains snu 93 and snu 101 isolated from veterinary staff and strains snu 117, snu 119, snu 118, snu 121, and snu 129 isolated from hospitalized animals were all oxacillin susceptible and had identical PFGE pulsotype, SCCmec type (SCCmec V), and antibiotic resistance pattern (TE-CIP-E-CC-SXT). In contrast, other clones, including MRSP strains snu 91, snu 92, snu 97, snu 102, and snu 103, isolated from veterinary staff, possessed nontypeable SCCmec and showed indistinguishable Sma I (Fig. 1B) and Asc I PFGE patterns (data not shown).
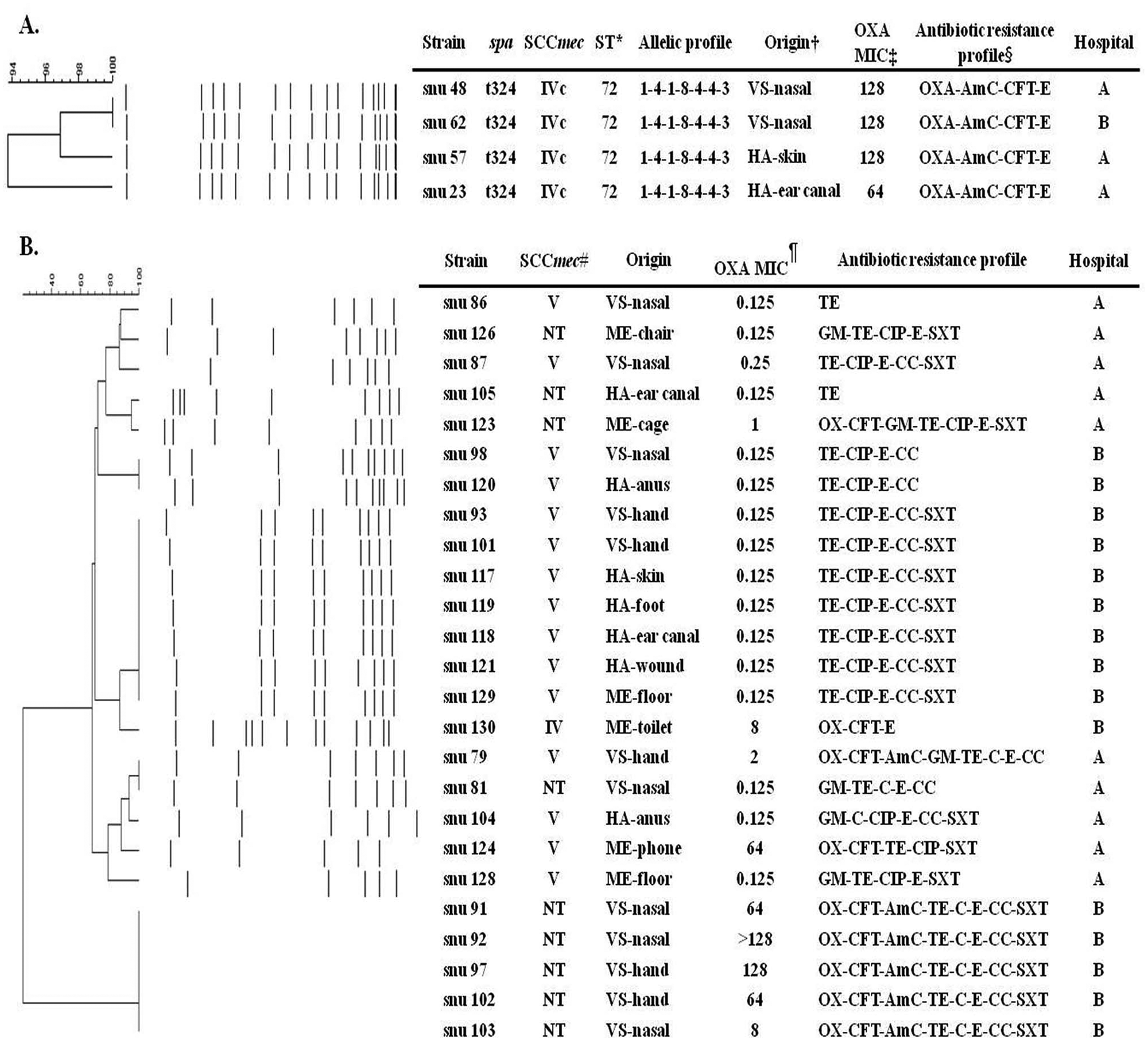
Genetic and phenotypic characteristics of methicillin-resistant, coagulase-positive staphylococci (MR-CPS).
Twenty-one methicillin-resistant S. epidermidis (MRSE) isolates had relatively diverse SCCmec types (6 types), sequence types (12 types), and PFGE pulsotypes (21 types). SCCmec type V and various SCCmec IV subtypes, including IVa, IVc, IVd, IVg, and non-subtypeable IV (IVnt), were detected. SCCmec IVa was the predominant subtype found in MRSE isolated from all sources. Among the 12 sequence types detected, ST258, ST267, ST268, and ST290 were newly identified types. Twenty-one different PFGE pulsotypes having 55–90% similarity were observed. Interestingly, the PFGE pulsotypes of the snu 41 and snu 44 strains were approximately 90% similar, although they were isolated from hospitalized animals and medical equipment, respectively (Fig. 2A).
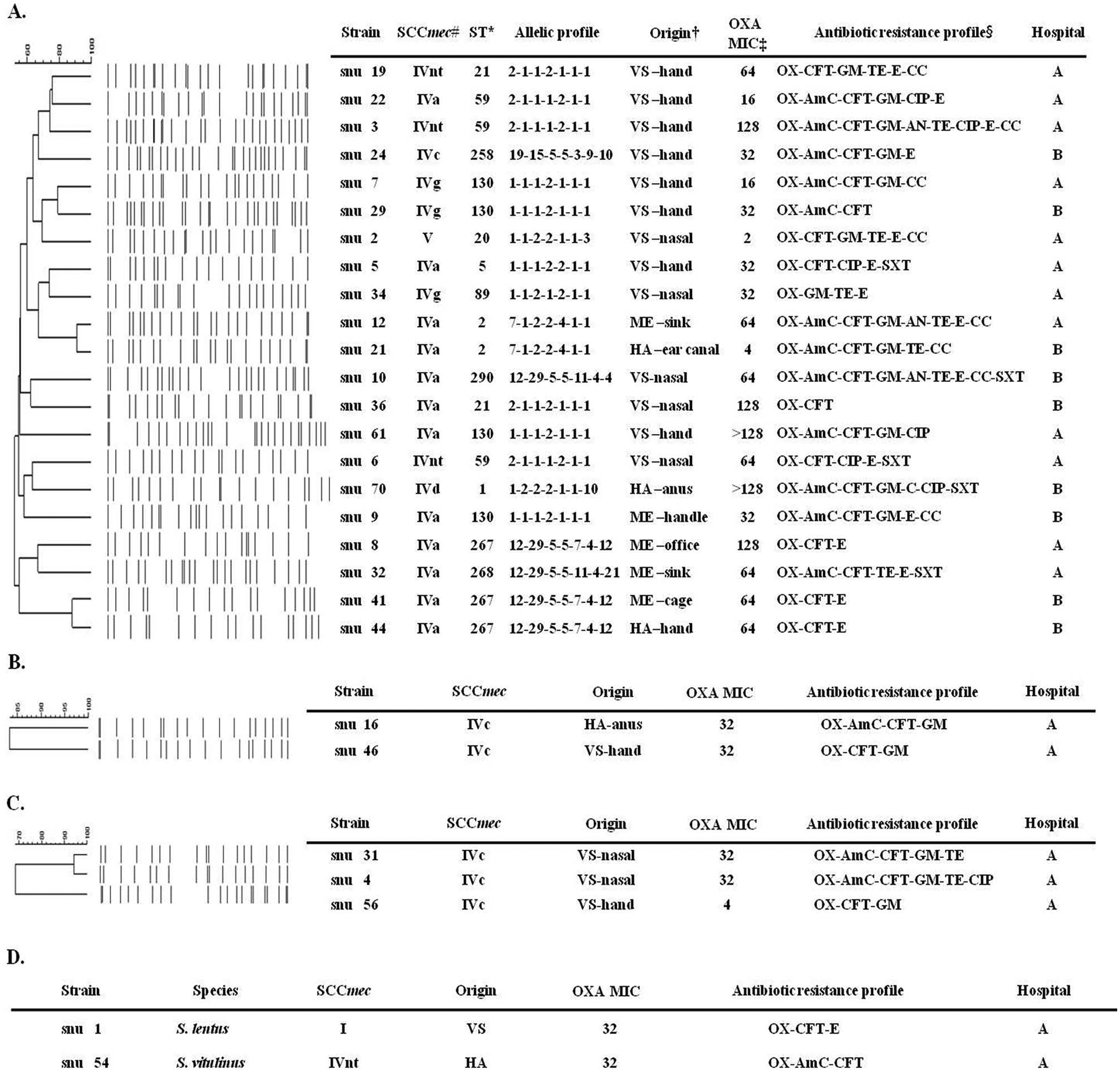
Genetic and phenotypic characteristics of methicillin-resistant, coagulase-negative staphylococci (MR-CNS).
Two strains of methicillin-resistant S. saprophyticus isolated from veterinary staff and hospitalized animals in the same hospital were SCCmec type IVc and showed an 83% PFGE pulsotype similarity (Fig. 2B). The pulsotypes of 3 methicillin-resistant S. warneri strains isolated from veterinary staff were 70–94% similar (Fig. 2C). The methicillin-resistant S. lentus strain recovered from veterinary staff and the methicillin-resistant S. vitulinus strain collected from a hospitalized animal were SCCmec type I and IVnt, respectively (Fig. 2D).
Discussion
In the present study, staphylococci were collected from veterinary staff, hospitalized animals, and medical equipment in 2 tertiary veterinary hospitals. Although only 2 hospitals were investigated, both provide care for large regions and handle many cases requiring long-term antibiotic therapy. Interestingly, the highest staphylococcal detection rate and the most diverse kinds of species were found among veterinary staff and not hospitalized animals or medical equipment. This is presumably due to the different host specificities of the staphylococcal species or the dynamic and frequent exposure of veterinary staff with medical equipment and hospitalized animals. 11 Moreover, methicillin-resistance rates were higher for isolates from veterinary staff and medical equipment than for those from hospitalized animals. These results support the hypothesis that veterinary staff and medical equipment may serve as possible reservoirs or fo-mites for the dissemination of MRS in the veterinary hospital setting.
Among the 11 staphylococcal species detected, S. pseudintermedius and S. epidermidis were the most dominant. In general, S. pseudintermedius is part of the commensal flora in cats and dogs as well as an etiological agent that causes skin infections, such as pyoderma, as well as otitis. 36 In the current study, the anus of hospitalized animals was the primary site colonized by S. pseudintermedius (35.3%, 6 strains from 17 samples), a colonization rate higher than that reported for the rectum in a previous study. 32 Interestingly, the current study found that 43.2% (16 strains from 37 samples) of veterinary staff had S. pseudintermedius in their nasal cavity. This detection rate is higher compared with previous studies in which 5–23% of veterinary staff members were found to have S. pseudintermedius. 18 The higher prevalence of S. epidermidis among veterinary staff and medical equipment compared with hospitalized animals could be explained by the fact that S. epidermidis is not only commensal in human beings but also produces a biofilm that enables attachment to machinery. 30
Methicillin-resistance analysis showed higher resistance rates among CNS compared with CPS. This has been previously reported in human 11 as well as veterinary medicine. 7 In terms of origin, 81.1% of veterinary staff harbored staphylococci, and more than half of these microorganisms (33 strains, 55.5%) were MRS.
In the present study, all MRSA were resistant to every β-lactam tested, OX-CFT-AmC. Among MRSP, 6 strains (63.3%) were resistant to OX-CFT-AmC, and 3 strains (31.7%) were only resistant to oxacillin. Three types were shown in MR-CNS: 16 strains (57.1%) with OX-CFT-AmC resistance, 11 strains (39.3%) with OX-CFT resistance, and 1 strain (3.6%) resistant to oxacillin. These different β-lactam–resistant phenotypes of MRS were also observed in previous studies. 33 The different expression and absence or presence of blaZ or genes regulating mecA might be the influencing factors. 5
In 2004, the Clinical and Laboratory Standards Institute guidelines recommended an MIC breakpoint of 0.5 µg/ml against oxacillin for the MRSP isolates from animals. 29 In 2008, this breakpoint was changed to 4 µg/ml for veterinary MRSP isolates. 9 A previous study, however, has demonstrated that the breakpoint of 4 µg/ml for oxacillin can lead to a failure to detect some mecA-positive S. pseudintermedius isolates. 3 It was also observed that 16 of 25 mecA-positive S. pseudintermedius isolates had an MIC less than 0.5 µg/ml to oxacillin. This might have been caused by reduced expression of the mecA gene, as previously reported for S. aureus. 34
In the current study, multidrug resistance was more common among both methicillin-susceptible and -resistant S. pseudintermedius than other staphylococcal species. All multidrug-resistant MSS were S. pseudintermedius and were resistant to erythromycin, tetracycline, chloramphenicol, clindamycin, trimethoprim–sulfamethoxazole, and fluoroquinolone. These are medicines generally prescribed for treating pyoderma, otitis, and enterocolitis in dogs.27,35 Further studies should be conducted to determine whether there are any advantages for acquiring resistance to various antibiotics in S. pseudintermedius compared with other staphylococci species in veterinary hospitals.
The genetic similarity observed with PFGE patterns among MRS isolated from veterinary staff and hospitalized animals suggested the existence of widely distributed clones with a broad host range or the possibility of movement between veterinary staff and hospitalized animals in agreement with previous studies.1,12 The MRSE strains, on the other hand, showed the most diverse genetic patterns, including 6 SCCmec types, 21 PFGE pulsotypes, and 11 sequence types. The pulsotype patterns differed even among the strains isolated from a single hospital, which had identical SCCmec types and sequence types. For example, both the MRSE snu 41 and snu 44 strains were ST2-SCCmec IVa, but their PFGE pulsotypes had only an 80% similarity. In human medicine, MRSE strains isolated from a single hospital or even a single infection site were found to have distinct genetic characteristics. 17 Consistent with previous reports,17,19 the current study found a genetically heterogeneous population of MRSE.
In the current study, all MRSA strains isolated from the 2 veterinary hospitals were found to have ST72-SCCmec IVc-t324. In South Korea, the predominant clones found in human hospitals and the general community were ST72-SCCmec IVa-t324 and ST72-SCCmec IVA-t324. 20 These sequence types also have been isolated in South Korea from food animals and products, including raw meat and raw milk.24,28 The MRSA strains isolated from companion animals in the present study were different from these sequence types only in terms of the SCCmec type. One possible explanation is that the ST72-SCCmec IVc-t324 clones were established in the veterinary hospital setting, independent from human medical environments, food animals, and MSSA-ST72-t324 strains that had acquired SCCmec IVc from other MRSA or MR-CNS strains. Indeed, it was found that methicillin-resistant S. saprophyticus and methicillin- resistant S. warneri strains harbored SCCmec IVc.
Most of the MRSE strains, unlike MRSA strains, possessed SCCmec IVa. Although MRSE has been intensively studied as a putative SCCmec donor for MRSA, 13 the same SCCmec type from MRSA was more frequently found in methicillin-resistant S. saprophyticus and methicillin-resistant S. warneri. In other countries, S. epidermidis harboring SCCmec IVa is predominantly found in human beings, 19 and it is possible that human MRSE strains are disseminated to companion animals via veterinary staff. More MRSA and MRSE sampling from both veterinary and human medical facilities should be performed to investigate the origin of MRSE harboring SCCmec IVa.
In summary, 11 species of MRS isolated from veterinary staff, hospitalized animals, and medical equipment in 2 South Korean veterinary hospitals were genetically characterized. The study found evidence of possible MRS dissemination between veterinary staff and hospitalized animals. To protect public health, continuous surveillance of MRS strains, more stringent sanitization guidelines for veterinary staff, and the prudent use of antimicrobials in veterinary medicine are needed.
Footnotes
a.
BBL™ CultureSwab™, BD Diagnostic Systems, Sparks, MD.
b.
Promed, Sungnam, Gyeonggi, South Korea.
c.
Vitek II system, bioMérieux Inc., Durham, NC.
d.
BBL™ Sensi-Disc™, BD Diagnostic Systems, Sparks, MD.
e.
Oxacillin, Sigma-Aldrich, St. Louis, MO.
f.
Blood DNeasy kit, Qiagen Inc., Valencia, CA.
g.
Lysostaphin, Sigma-Aldrich, St. Louis, MO.
h.
BioNumerics version 2.0, Applied Maths, Kortrijk, Belgium.
i.
Ridom GmbH, Wurzburg, Germany.
j.
SPSS Inc., Chicago, IL.
The author(s) declared that they had no conflicts of interest with respect to the research, authorship, and/or the publication of this article.
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the National Veterinary Research and Quarantine Service (C-AD13-2006-14-04); National Research Foundation of Korea grants funded by the South Korean government (NRF-2010-013-E00025 and NRF-2010-1-E00057); the Research Institute of Veterinary Science, Department of Veterinary Microbiology, College of Veterinary Medicine; and the BK21 Program for Veterinary Science, Seoul National University, Seoul, South Korea.
