Abstract
Pseudomonas aeruginosa is an opportunistic pathogen that has been recognized as a cause of endometritis in mares. Pulsed field gel electrophoresis was used to characterize and compare isolates of P. aeruginosa from an outbreak of endometritis and unrelated isolates collected at the same time as the outbreak. The restriction endonuclease digestion patterns and antimicrobial resistance profiles of all outbreak isolates were identical. Therefore, a single strain of P. aeruginosa was responsible for the cases of endometritis. The unrelated isolates could be distinguished from the outbreak strain using the techniques outlined in the present study. The results establish that this pathogen was not venereally transmitted between all the horses from which it was isolated, but rather must have been disseminated, at least initially, from a contaminated water source. Once the water used to clean the mares and stallions was replaced, there were no further reports of endometritis caused by this organism on the affected stud. Furthermore, the fertility of the stallions was not affected, in spite of persistent carriage for 1 to 2 months. The current study has shown that the use of pulsed field gel electrophoresis has considerable value in epidemiological investigations of equine urogenital tract infections with P. aeruginosa.
Keywords
Pseudomonas aeruginosa is commonly recognized as a cause of endometritis in mares and is regarded as a venereally transmitted pathogen by some authorities.3,5 However, there has been little characterization of strains involved in outbreaks of endometritis attributed to P. aeruginosa, and thus the common etiology of cases has been based on circumstantial evidence, rather than definitive proof that a single strain of the organism had been transmitted between a group of mares and stallions.
Characterization of strains of P. aeruginosa has previously been based on phage typing, but there has been limited application of this technique to veterinary isolates.2,3 Other methods that may be applied include comparison of antimicrobial resistance patterns. 4 More recently, random amplification of polymorphic DNA and multilocus sequence typing techniques have been employed to genotype equine isolates of P. aeruginosa gathered for epidemiological and retrospective studies.10,15
Medical isolates of P. aeruginosa associated with outbreaks have been compared and typed by pulsed field gel electrophoretic separation of fragments of their genomic DNA generated by digestion with restriction endonucleases with recognition sites that occur infrequently in P. aeruginosa DNA.8,14 This technique, most commonly using the restriction endonucleases SpeI or XbaI for digestion of the genomic DNA, has allowed unequivocal characterization of outbreaks associated with a single strain, and clear distinction of unrelated strains isolated from situations where they may have been thought to be epidemiologically related.6,9,11,13,16 Although this technique has been applied to some veterinary pathogens, including Salmonella sp. outbreaks in equine hospitals, 1 it has not been applied to investigation of outbreaks of disease due to P. aeruginosa. The purpose of the current study was to examine the patterns generated by pulsed field gel electrophoresis (PFGE) of genomic DNA from isolates of P. aeruginosa from an outbreak of endometritis on a Thoroughbred breeding farm to epidemiologically unrelated isolates to assess the potential of PFGE for investigation of venereal transmission of P. aeruginosa.
All isolations of P. aeruginosa from 4 different Thoroughbred breeding farms in New South Wales and Victoria, Australia, during November and December 1998 and January 1999 were made by the Scone Veterinary Diagnostic Laboratory (Scone, New South Wales, Australia; Table 1). Most were obtained from swabs of the male and female urogenital tracts, with 1 isolate made from a case of mastitis. All mares showed signs of endometritis, but the stallions were asymptomatic carriers.
Isolates of Pseudomonas aeruginosa from 4 different Thoroughbred breeding farms in New South Wales and Victoria, Australia, during November and December 1998 and January 1999.*
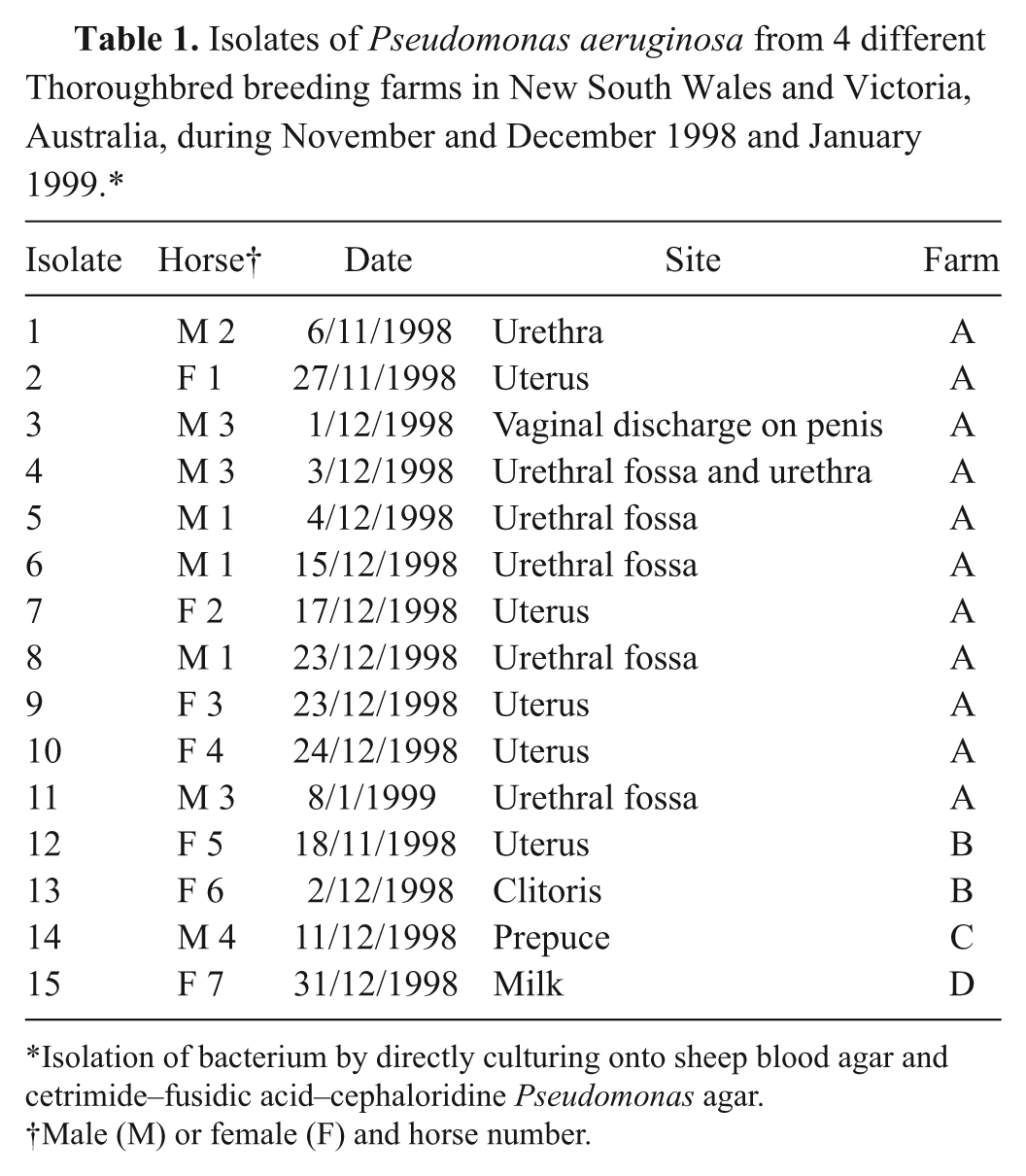
Isolation of bacterium by directly culturing onto sheep blood agar and cetrimide–fusidic acid–cephaloridine Pseudomonas agar.
Male (M) or female (F) and horse number.
The farms applied routine measures to ensure biosecurity during mating. Guarded uterine swabs were collected, using a new disposable speculum and new disposable gloves, from each mare prior to examination and mating. If bacteria were cultured from the swab, the mare was not mated until after she had been successfully treated to eliminate infection. The external genitalia of the stallions were swabbed periodically throughout the breeding season. Mares, but not stallions, were washed with running water prior to mating.
Isolates were cultured on sheep blood agar plates and cetrimide–fusidic acid–cephaloridine (CFC) Pseudomonas agar a and characterized as P. aeruginosa on the basis of colonial appearance, production of typical pyocyanin pigment, and the oxidase reaction. The identification was confirmed biochemically using a commercial kit. b The susceptibility of the isolates to a panel of antimicrobial agents was determined, essentially as described for the Clinical and Laboratory Standards Institute agar disk diffusion method. 12 Isolates were stored as nutrient agar stab cultures at 4°C until transfer to The University of Melbourne (Parkville, Victoria, Australia) for further characterization.
Genomic DNA from each isolate was prepared in agarose blocks as described previously, 1 with the exception that once set, the blocks were added to 0.5 M ethylenediamine tetra-acetic acid [EDTA, pH 8.0], 1% w/v lauryl sarcosine, and 1 mg/ml of proteinase K and incubated at 50°C for 48 hr with 1 change of buffer. The blocks were washed in TE (10 mM Tris base; 1 mM EDTA, pH 8.0) and stored at 4°C in EDTA (0.25 mM, pH 8.0) until digestion. Slices, of approximately 1 mm thickness, were equilibrated in TE and incubated for 30 min in 100 µl of the restriction endonuclease buffer. c The DNA contained within the agarose slices were digested with 10 U of SpeI c then incubated at 4°C for approximately 1 hr and then at 37°C for a further 18 hr. The digested genomic DNA was separated by PFGE. d The slices were loaded into a 1% (w/v) agarose, 0.5× TBE (45 mM Tris–HCl, 45 mM boric acid, 1 mM EDTA, pH 8.0) gel and subjected to an electric field of 6 V/cm, with the pulse time ramped from 1 to 40 sec and an included angle of 120°, for 20 hr at 14°C.
The endonuclease cleavage patterns of the isolates made from the outbreak of endometritis on farm A were identical (Fig. 1), supporting the observation that all of the 11 isolates had identical patterns of susceptibility to the antimicrobial agents tested (Table 2). This cleavage pattern differed markedly from those obtained from epidemiologically unrelated isolates obtained from the reproductive tract of horses from the same region and at the same time as those obtained during the outbreak (Fig. 2). The findings of the present study clearly show that the outbreak of endometritis and stallion genital infection was caused by a single strain of P. aeruginosa, with the same strain isolated from the external genitalia of 3 stallions and the uteri of 4 mares.
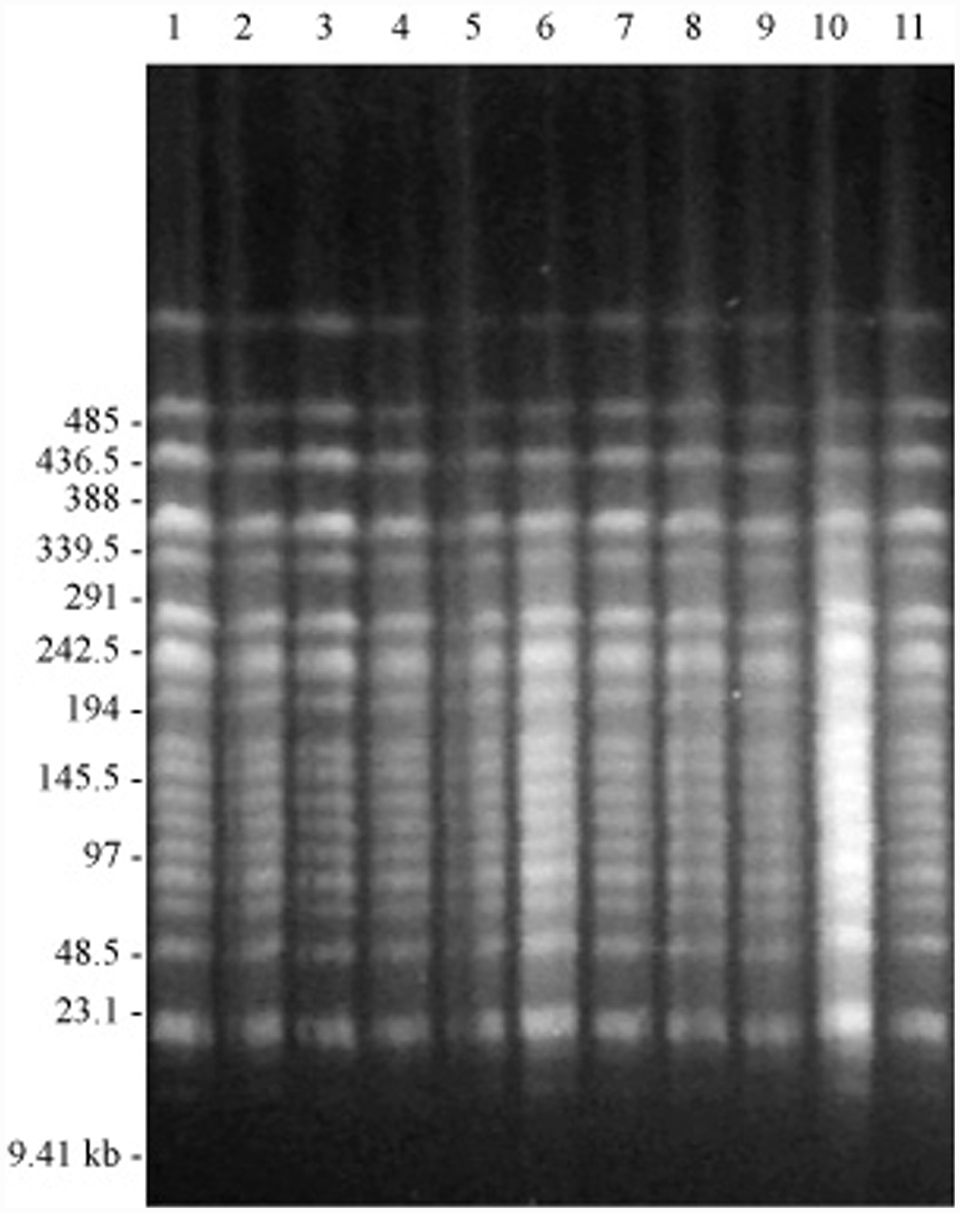
Pulsed field gel electrophoresis patterns of SpeI digested genomic DNA from Pseudomonas aeruginosa isolated from the reproductive tracts of stallions and mares on a single breeding farm experiencing an outbreak of endometritis. Lanes 1–11 show the patterns for isolates from farm A. The molecular weight markers were phage lambda DNA digested with HindIII and a phage lambda ladder. d
Antimicrobial susceptibility of the outbreak strain and other unrelated isolates of Pseudomonas aeruginosa.*
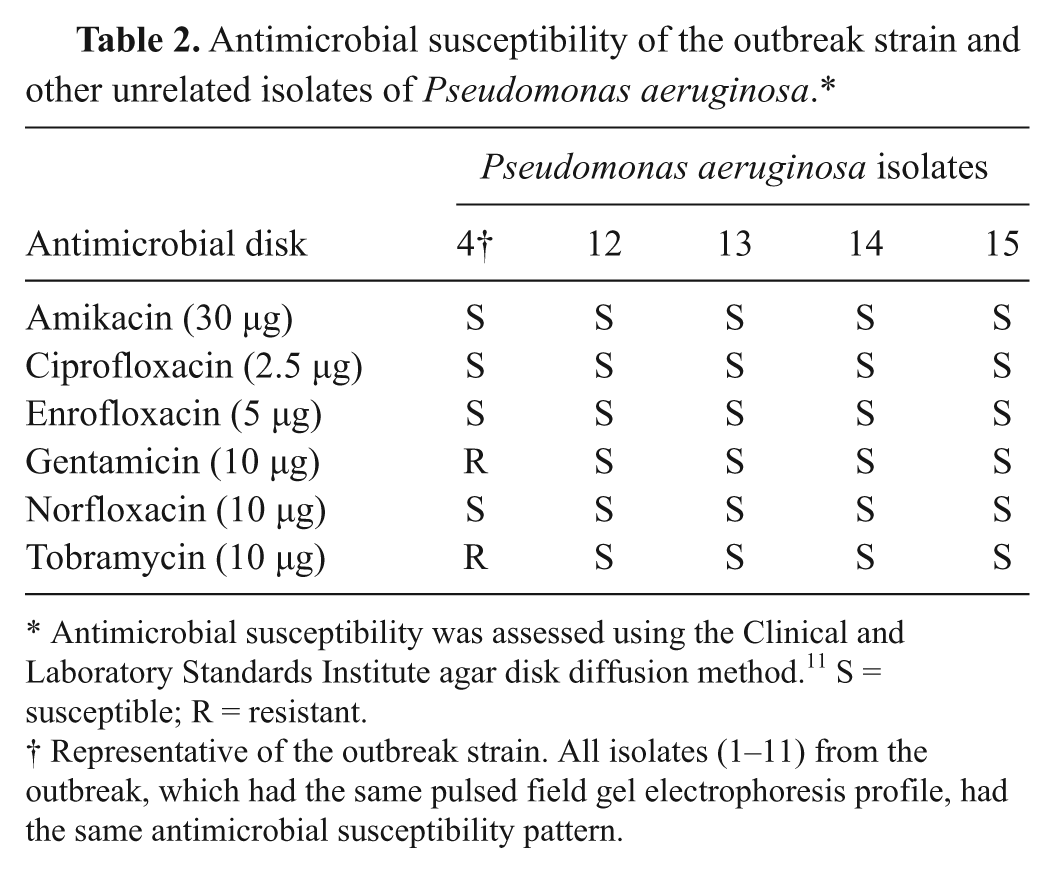
Antimicrobial susceptibility was assessed using the Clinical and Laboratory Standards Institute agar disk diffusion method. 11 S = susceptible; R = resistant.
Representative of the outbreak strain. All isolates (1–11) from the outbreak, which had the same pulsed field gel electrophoresis profile, had the same antimicrobial susceptibility pattern.
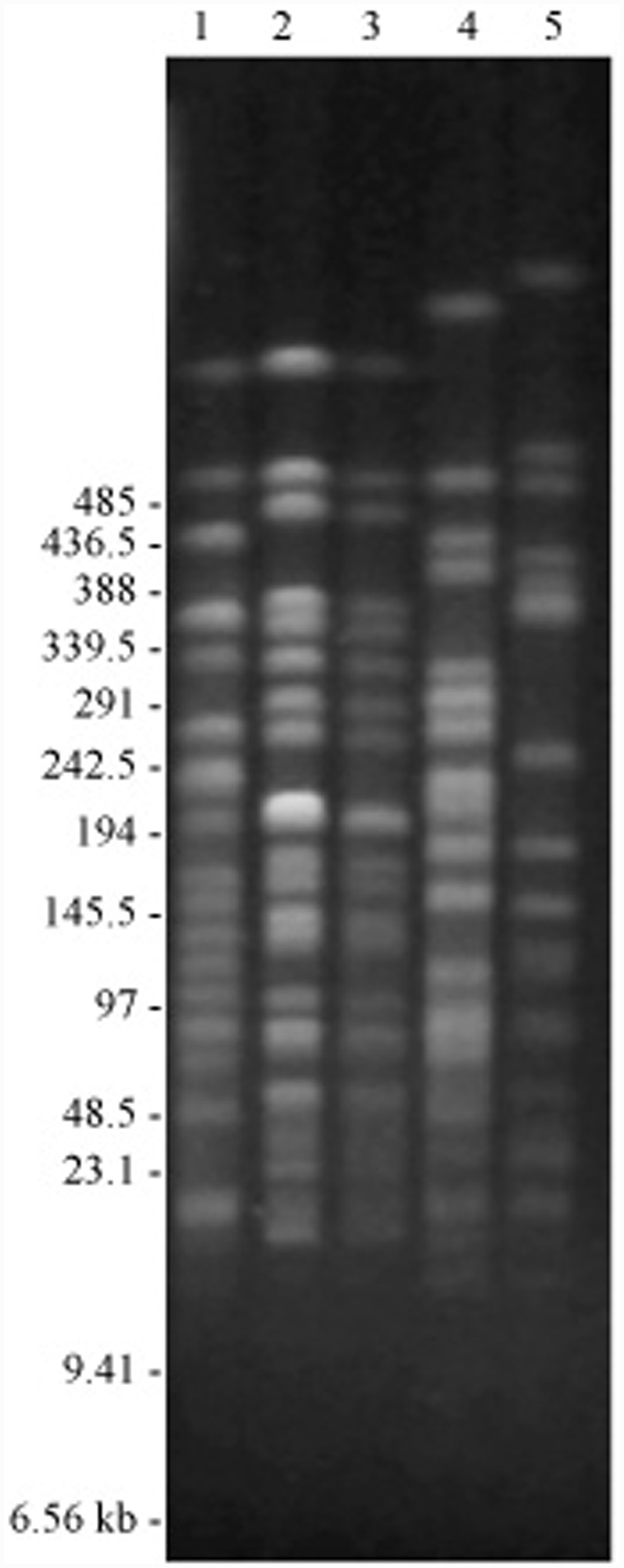
Pulsed field gel electrophoresis patterns of Spe I digested genomic DNA from Pseudomonas aeruginosa isolated from the reproductive tracts of stallions and mares on 4 separate breeding farms. Lane 1 shows isolate 4, a representative of the 11 identical isolates obtained from the outbreak farm. Lanes 2–5 show the isolates obtained from farms B–D. The same molecular weight markers were used as in Figure 1.
In a previous Irish study, in which retrospective genotyping of equine isolates was conducted, little evidence of a clonal relationship was detected between the isolates; however, available information about the origin and links between these isolates was limited. 15 The diversity of the PFGE profiles among the isolates unrelated to the outbreak in the present study would also indicate a non-clonal relationship between epidemiologically unrelated isolates.
A study of P. aeruginosa isolated from genital tracts of horses in a single geographical area of Queensland, Australia, found that a significant percentage of the mares harbored a dominant clonal complex, as determined by multilocus sequence typing. 10 Pseudomonas aeruginosa was not isolated from the stallions sampled. On that basis, the authors of the study 10 concluded that venereal transmission did not play a role in spread of infection and suggested that common environmental sources are involved in the spread of this organism.
Venereal transmission did not account for all of the spread of P. aeruginosa infection during the outbreak described in the current study, as the 3 stallions were serving different mares. As in the Queensland study, 10 this suggests that there was contamination of the external genitalia of the stallions or mares from a common environmental source, possibly the water used to wash the genitalia of the mares prior to breeding. The organism was then transmitted between stallion and mares, inducing endometritis. As P. aeruginosa with the same antimicrobial sensitivity pattern was isolated from 4 mares and 3 stallions, and the mating history of these 7 horses suggested that venereal transmission could not explain the presence of the same strain of P. aeruginosa in all 7 horses, the source of the water used to wash the mares was changed. New infections with this strain of P. aeruginosa did not occur after the water source used to clean mares was changed. Unfortunately a sample of water was not available for analysis.
It is notable that the strain was isolated from stallion 1 on 4 occasions (the first isolate, made on November 3, 1999 was not available for analysis) over nearly 2 months and from stallion 3 on 3 occasions over 1 month, but that during this time the fertility of these stallions was unaffected, and there was not widespread transmission to all mares served by these stallions.
The current study has provided evidence that use of PFGE of fragments of genomic DNA has considerable value in epidemiological investigations of equine urogenital tract infections with P. aeruginosa. Identical restriction endonuclease cleavage patterns can be assumed to indicate a common origin, 7 especially given the great variability seen between epidemiologically unrelated isolates in the present study. Further investigations may allow unequivocal demonstration of the significance of venereal transmission in endometritis caused by this pathogen.
