Abstract
Adherence factors and enterotoxins are major virulence factors of enterotoxigenic Escherichia coli (ETEC). Antibiotics have been used frequently for the treatment and prevention of ETEC infection in piggeries worldwide, including Korea. Therefore, data on both virulence profiles and antibiotic resistance patterns are useful in the epidemiological study of ETEC. A total number of 198 E. coli field isolates were examined. The most prevalent pathotype was F1, followed by a combination of F1 and EAST1. All of the 71 isolates were resistant to more than 2 antibiotics used in a disk diffusion test, and 87.94% of the isolates were found to be resistant to more than 4 antibiotics. Investigations were also conducted to correlate the virulence gene profiles with antibiogram and pulsed-field gel electrophoresis (PFGE). Although a high degree of polymorphism was noted among strains having the same virulence patterns, the highest similarity pattern was observed carrying the same virulence profiles and similar antibiogram. Thus, investigation of both virulence profiles and antibiogram is essential to the epidemiological study of ETEC. Moreover, the PFGE method might be applicable as a tool to reveal genetic relatedness among E. coli strains from piggeries in Korea.
Neonatal diarrhea caused by Escherichia coli infection is responsible for major losses in the pig industry worldwide. Enterotoxigenic E. coli (ETEC) is the main cause of diarrhea during the suckling and weaning periods. 17 Adherence factors (F1 [type 1 fimbriae], F4 [K88], F5 [K99], F6 [987P], F18, F41, adhesin involved in diffuse adherence [AIDA]-I, and porcine attaching and effacing-associated factor [Paa]) and heat-stable enterotoxins (STa and STb), heat-labile enterotoxin (LT), shiga toxin 2e (Stx2e), and enteroaggregative E. coli heat-stable entero-toxin 1 (EAST1) have been known to be virulence factors of ETEC. 2,3,6,13,17,19,24 The ETEC strains attach to intestinal epithelium using adherence factors and produce 1 or more enterotoxins. 16–18,25 Subsequently, ETEC can induce acute and watery diarrhea that may be followed by terminal bacteremia, which remains an important cause of economic loss for swine industries. 6,17,18
Antibiotics have been used frequently for treatment and prevention of ETEC infection in piggeries. However, excessive use of antibiotics has resulted in dissemination of antibiotic resistance in E. coli isolates. 1,9 Multiple antimicrobial resistances may arise from many different genetic determinants, and each may present specific epidemiological features. Therefore, determining antibiotic patterns would be useful in understanding and controlling ETEC infection in swine. 7,11
There are some reports in which virulence profiling and antibiogram of E. coli isolates from pigs were studied, 4,11,12,14,15,25 but only in a few cases have comparative results of both been shown. In the current study, genetic relationships between E. coli strains isolated from piggeries were determined by examining both virulence profiling and antibiotic resistance patterns. Investigations were also conducted to correlate the virulence gene profiles with antibiogram by pulsed-field gel electrophoresis (PFGE), which has been used for differentiation and epidemiological characterization of E. coli isolated from pigs with diarrhea. The PFGE method is a sensitive molecular method to analyze the degree of genetic relatedness or variability among porcine ETEC strains. 21–23 To the authors' knowledge, genetic relatedness compared with virulent and antibiotic patterns of ETEC from pigs have never been reported in Korea. Therefore, the purpose of the current study is to examine virulence genotype, antibiogram, and PFGE profiles of ETEC strains.
Between 2002 and 2007, 198 E. coli strains were isolated from 2- to 8-week-old piglets with diarrhea on different farms in Korea. After incubation at 37°C for 18 hr on MacConkey's and eosin methylene blue agar plates, a single colony was picked up and identified as E. coli by IMViC (indole, methyl red, Voges-Proskauer, and citrate) test and an automated system. a The field isolates were grown overnight in tryptic soy broth (TSB) and stored at −80°C after the addition of 12.5% glycerol until further use.
For polymerase chain reaction (PCR) amplification, 14 sets of oligonucleotide primers b were used to detect the genes encoding for the following 13 virulence factors of ETEC: adherence factors (F1, F4, F5, F6, F18, F41, AIDA-I, and Paa) and enterotoxins (STa, STb, LT, Stx2e, and EAST1). The PCR assay for detection of virulence factors of ETEC was carried out as described previously. 13
Seventy-four E. coli isolates selected by PCR results were tested for antibiotic resistance by the agar disk diffusion method performed on Mueller-Hinton agar c with antibiotic susceptibility disks d , e according to the guidelines of the Clinical and Laboratory Standards Institute (CLSI). 5 The active ingredients of the selected antibiotics were amikacin, ampicillin, amoxicillin, aztreonam, cefoxitin, ceftazidime, ceftriaxone, chloramphenicol, ciprofloxacin, colistin, enrofloxacin, gentamicin, imipenem, nalidixic acid, streptomycin, tetracycline, and trimethoprim/sulfamethoxazole. Escherichia coli ATCC 25922 and Salmonella enterica Typhimurium DT104 (ATCC 2501) were used for quality control. Interpretation was performed according to the criteria recommended by CLSI 5 or the supplier.
Pulsed-field gel electrophoresis was carried out as described by the Centers for Disease Control and Prevention for use in the tracing of E. coli O157:H7 outbreaks with the following modifications. 8 A single colony of isolate to be tested was inoculated in 5 ml of TSB and incubated overnight at 37°C. The cells were washed in SE buffer (75 mmol NaCl, pH 8.0; 25 mmol ethylenediamine tetra-acetic acid [EDTA], pH 8.0) by centrifugation, and the optical density of the cells at a wavelength of 610 nm was adjusted to 1.40. Chromosomalgrade agarose f g was prepared in 10 mmol Tris and 0.1 mmol EDTA to a final concentration of 2.0% and maintained at 55°C. Plugs were formed by mixing 0.5 ml of cell suspension with 0.5 ml of agarose, and this mixture was then added into plug molds. f After solidification, agarose plugs were transferred to lysis buffer (50 mmol Tris, pH 8.0; 50 mmol EDTA, pH 8.0; 1% sarcosine; 0.1 mg of proteinase per milliliter) and incubated for 16–20 hr in a 55°C water bath. The lysis buffer was removed, and the plugs were washed with 5 ml of sterile distilled water for 5 min and then with 3 ml of TE buffer (10 mmol Tris, pH 8.0; 1 mmol EDTA, pH 8.0) for 5 min at room temperature. A final set of 4 washes, 30 min each, was done with 3 ml of TE buffer per wash. After 1 hr adaptation in 300 μl of restriction buffer, the plugs were transferred to the fresh mixture containing 30 U of XbaI or NotI restriction enzyme g and incubated at 37°C for 16–20 hr. The plugs were then briefly soaked in standard 0.5 Tris–borate–EDTA (TBE) prior to electrophoresis. Electrophoresis of prepared samples was performed using a commercial system f by using pulsed field agarose f with 2.2 liters of standard 0.5 TBE running buffer. The gels were then run at pulse ramps from 2.2 to 54.2 sec for 22 hr. After electrophoresis, the gels were stained with ethidium bromide for 30 min, destained with distilled water for 10 min, and photographed under ultraviolet light using a commercial fluorescent imaging system. f
Prevalent pathotypes of virulence genes among 198 Escherichia coli strains isolated from piglets with diarrhea.*
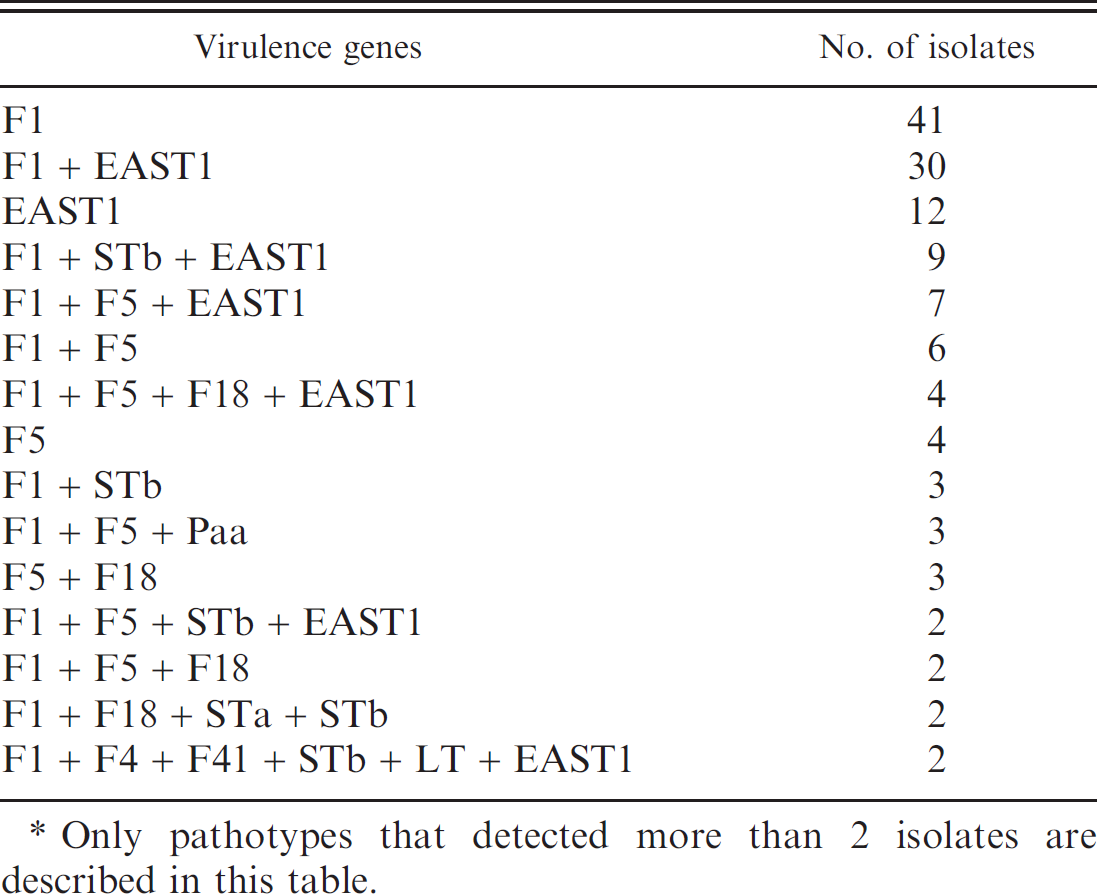
Only pathotypes that detected more than 2 isolates are described in this table.
Antimicrobial resistance of Escherichia coli isolated from piglets with diarrhea.*
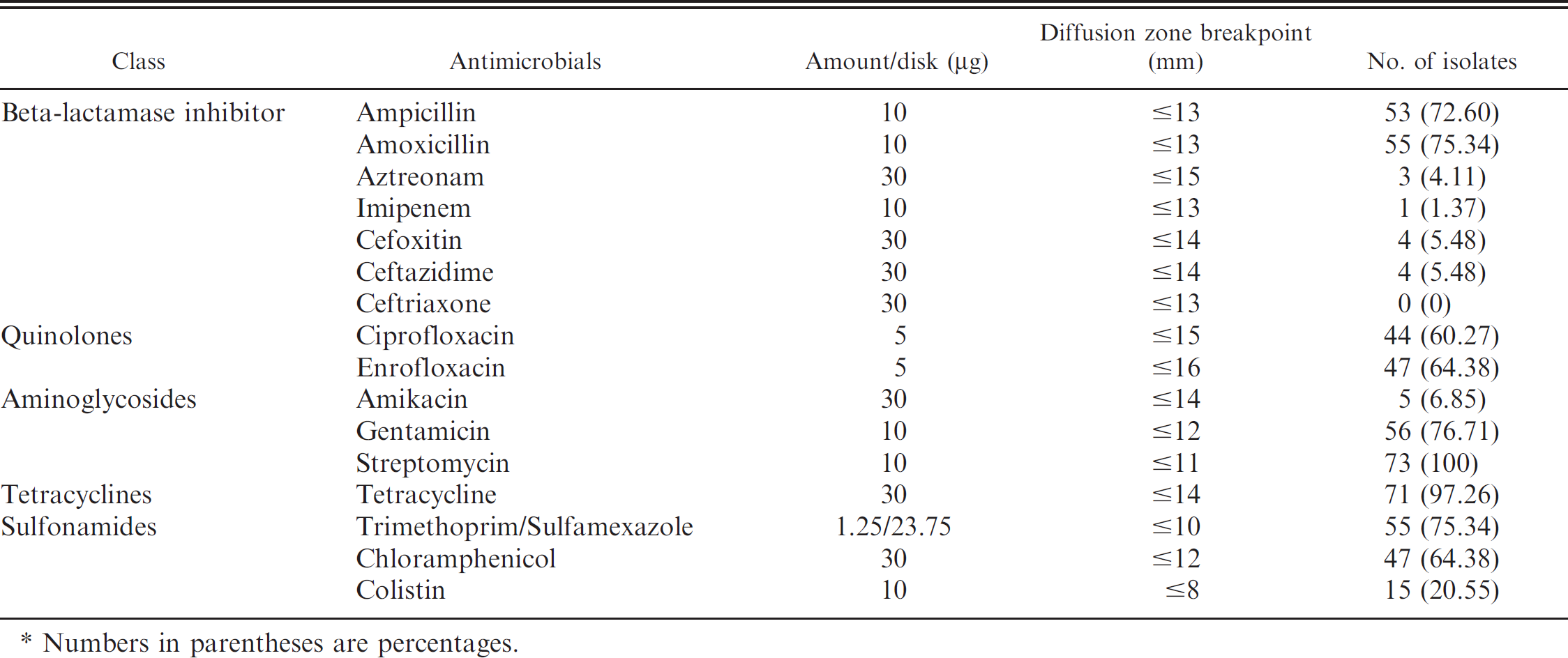
Numbers in parentheses are percentages.
Antimicrobial resistance patterns of Escherichia coli isolated from piglets (n = 74).*
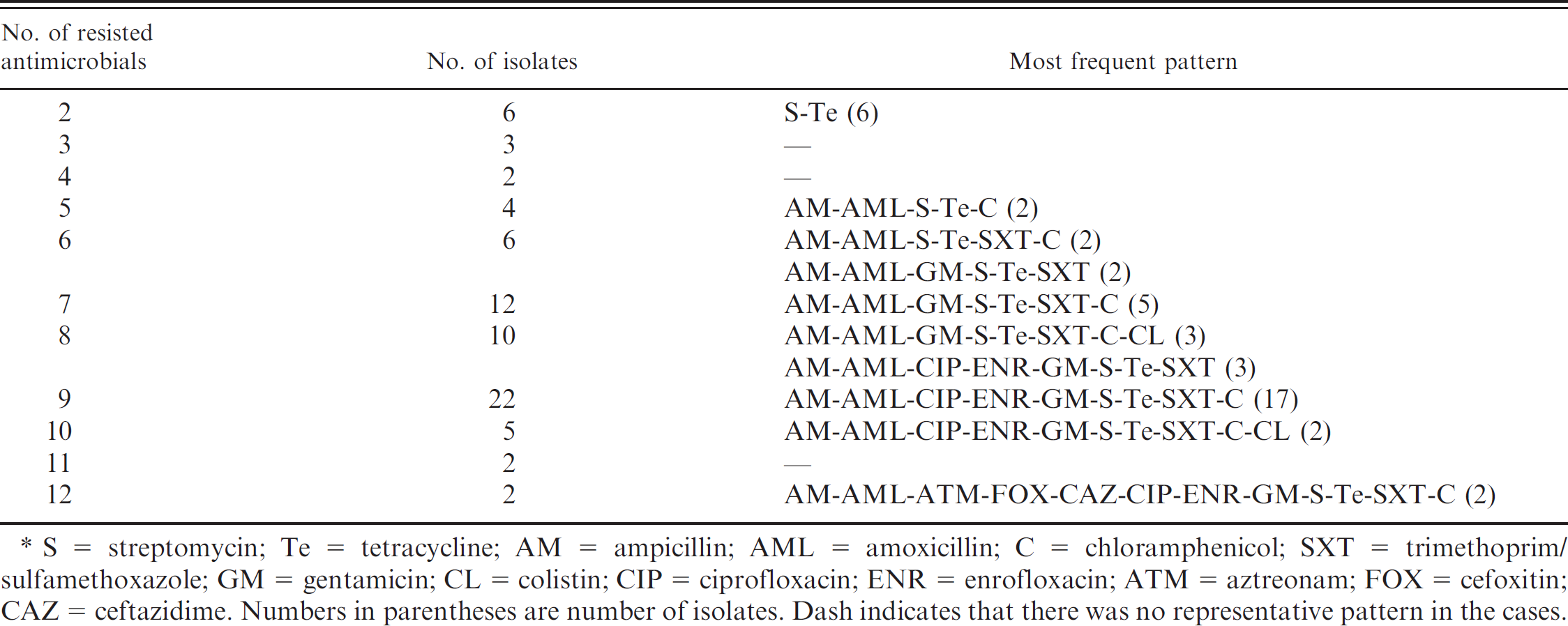
S = streptomycin; Te = tetracycline; AM = ampicillin; AML = amoxicillin; C = chloramphenicol; SXT = trimethoprim/sulfamethoxazole; GM = gentamicin; CL = colistin; CIP = ciprofloxacin; ENR = enrofloxacin; ATM = aztreonam; FOX = cefoxitin; CAZ = ceftazidime. Numbers in parentheses are number of isolates. Dash indicates that there was no representative pattern in the cases.
Gel images were scanned and analyzed using a commercial software. f Dendrograms were created from a matrix of band matching using the Jaccard coefficient and the unweighted pair group method with arithmetic mean analysis.
Of 198 E. coli strains, 180 isolates (90.91%) were shown to be positive for the genes encoding more than 1 of the virulence factors of ETEC. Ninety-five isolates (47.98%) carried both adhesins and enterotoxins. A presence of diverse patterns of genetic profiles in virulence was observed, and the most prevalent pathotype was F1 only, followed by a combination of F1 and EAST1 (Table 1).
Based on the presence of virulence factors, 41 E. coli isolates having only F1 gene and 30 E. coli isolates carrying both F1 and EAST1 were selected for antimicrobial susceptibility testing. In addition, 3 isolates not carrying any virulence factors were used as controls. All isolates were resistant to more than 4 antibiotics used in the disk diffusion test. Isolates showed the highest resistance to streptomycin (100%), followed by tetracycline (97.26%), gentamicin (77.03%), trimethoprim/sulfamexazole (75.68%), amoxicillin (75.68%), ampicillin (72.97%), chloramphenicol (64.86%), enrofloxacin (64.86%), and ciprofloxacin (59.46% Table 2). Multidrug resistance was demonstrated in almost all of the isolates, and 87.94% of the isolates were found to be resistant to 4 or more antibiotics (Table 3).
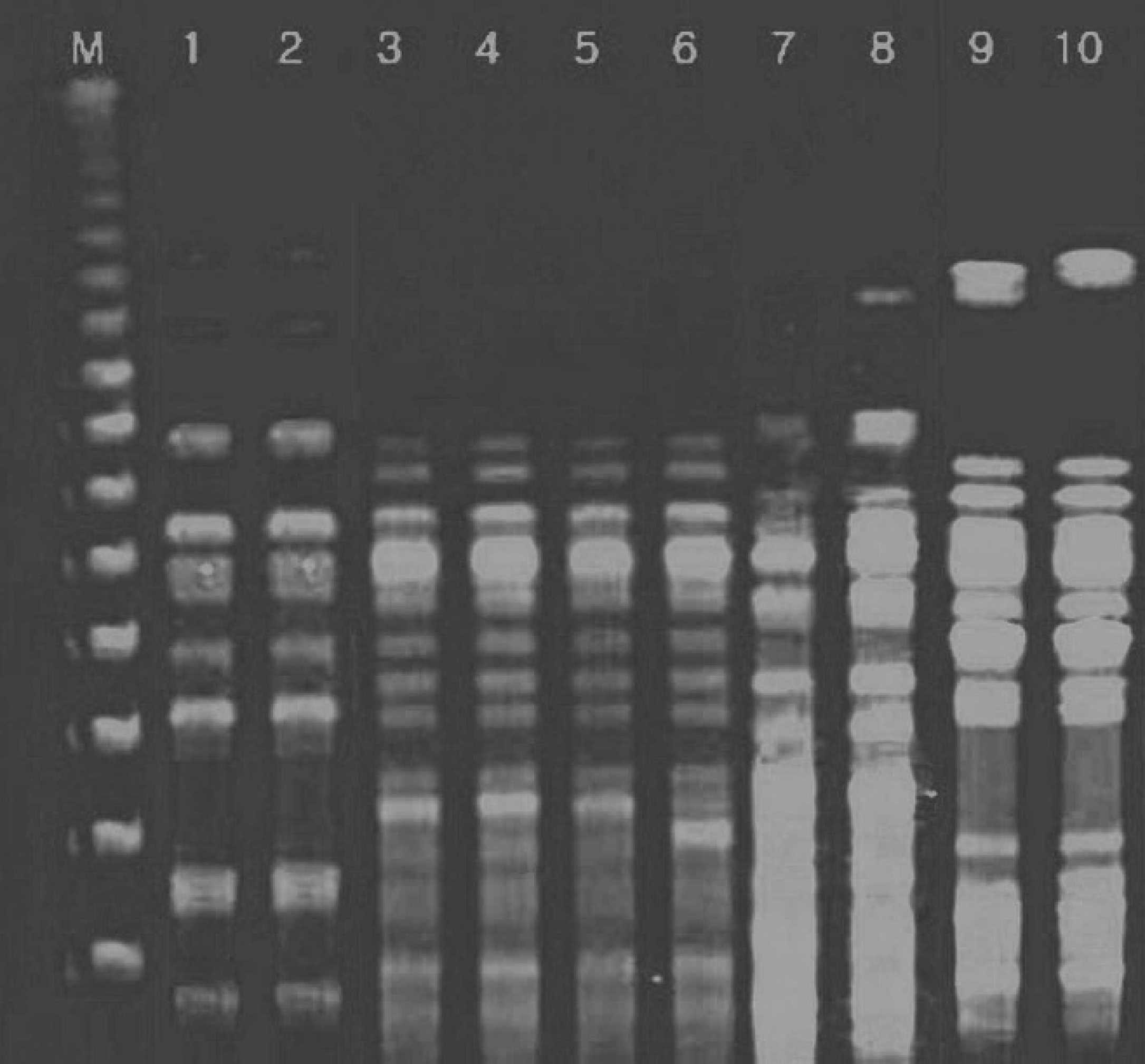
Representative pulsed-field gel electrophoresis patterns of XbaI-digested genomic DNA of Escherichia coli strains from pigs. Lane M: lambda DNA marker; lanes 1, 2: E. coli strains 15, 17, respectively; lanes 3–6: E. coli strains 32–35, respectively; lanes 7, 8: E. coli strains 28, 29, respectively; lanes 9, 10: E. coli strains 48, 49, respectively.
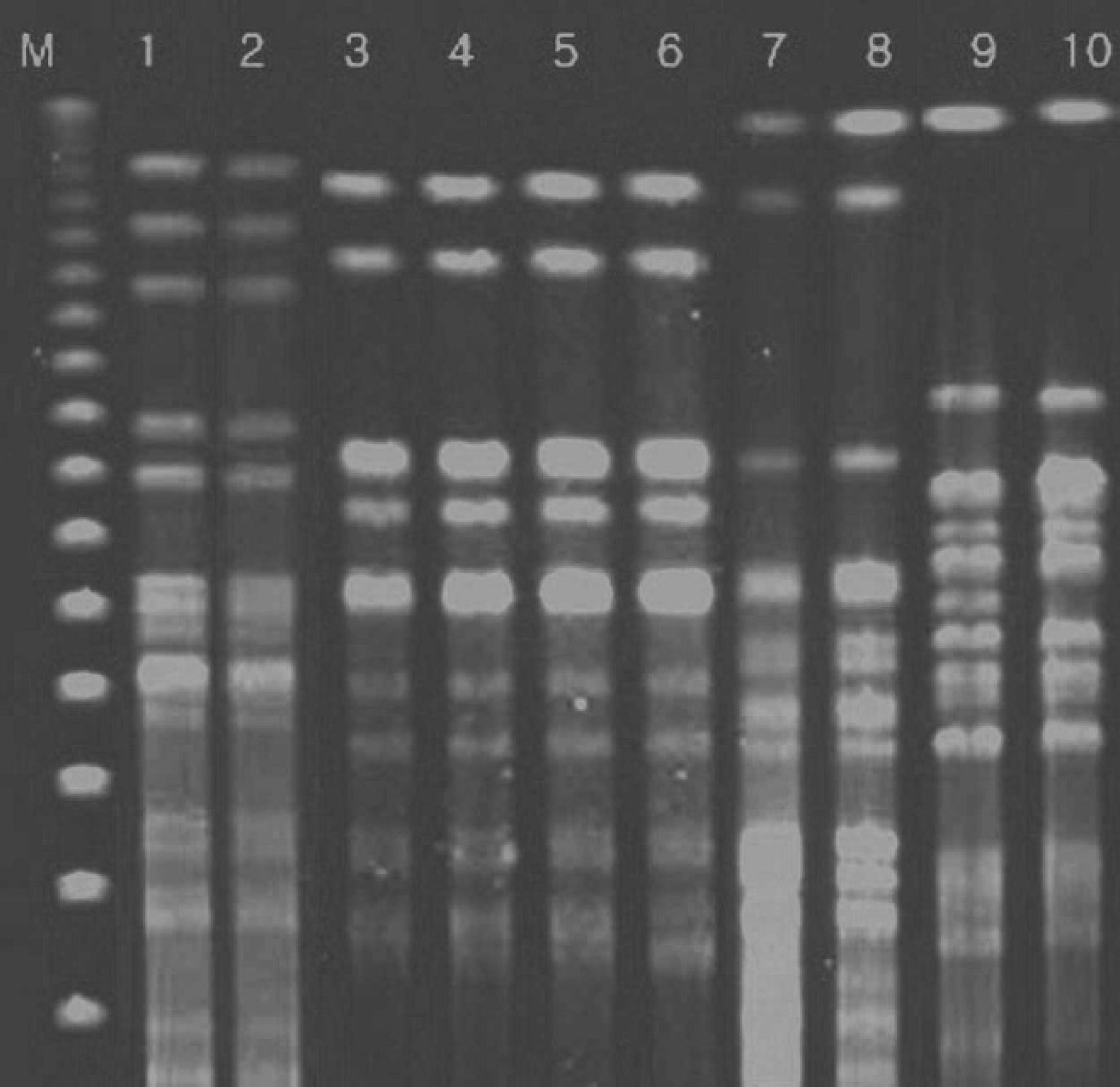
Representative pulsed-field gel electrophoresis patterns of NotI-digested genomic DNA of Escherichia coli strains from pigs. Lane M: lambda DNA marker; lanes 1,2: E. coli strains 16, 18, respectively; lanes 3–6: E. coli strains 32–35, respectively; lanes 7, 8: E. coli strains 28, 29, respectively; lanes 9, 10: E. coli strains 3, 45, respectively.
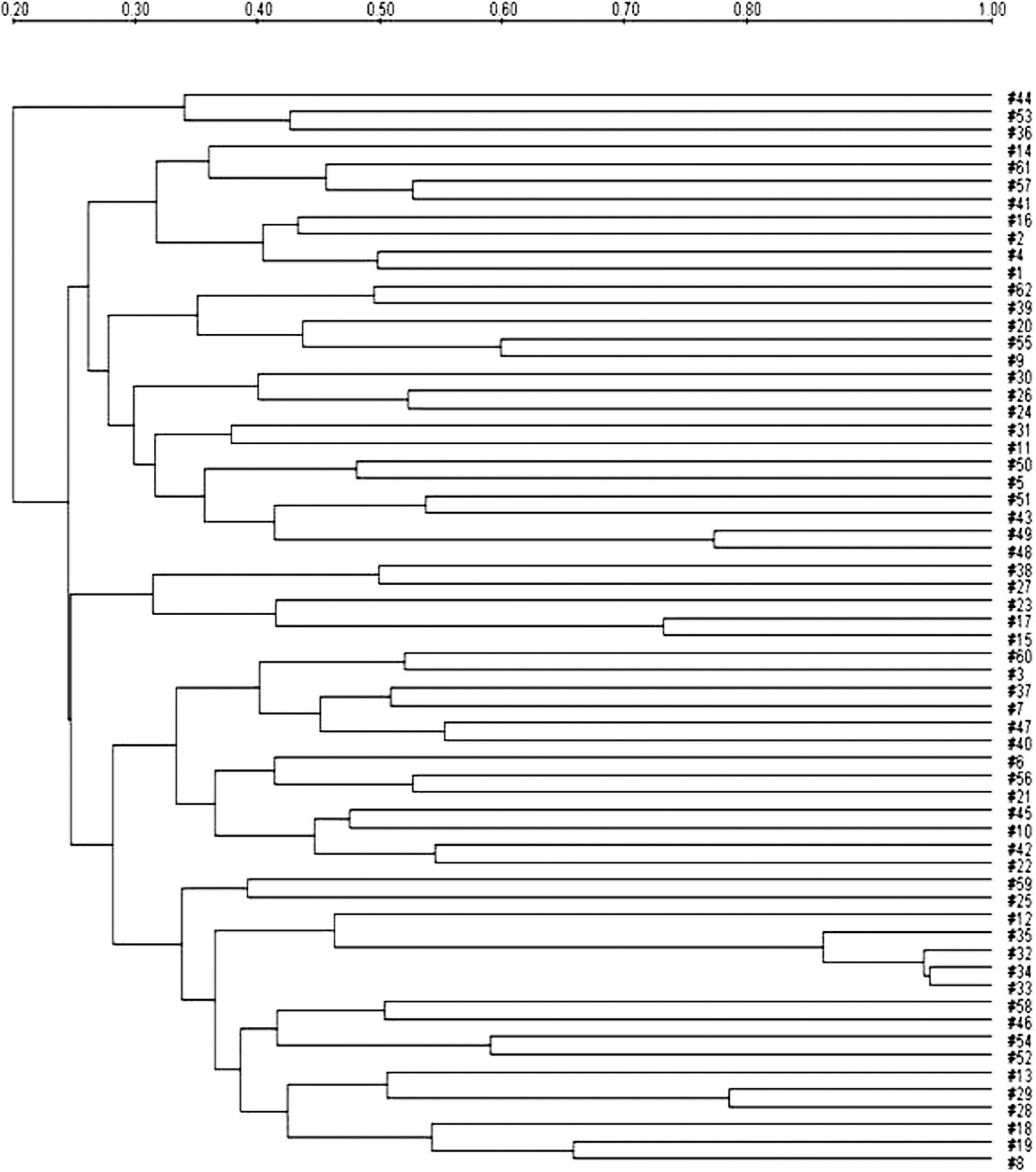
Dendrogram using the Dice coefficient and unweighted pair group method with arithmetic mean analysis showing the relationships between Escherichia coli strains with XbaI restriction. The degree of similarity (%) is shown on the scale.
The genetic relatedness of the E. coli isolates of prevalent pathotypes was examined by the use of PFGE analysis.
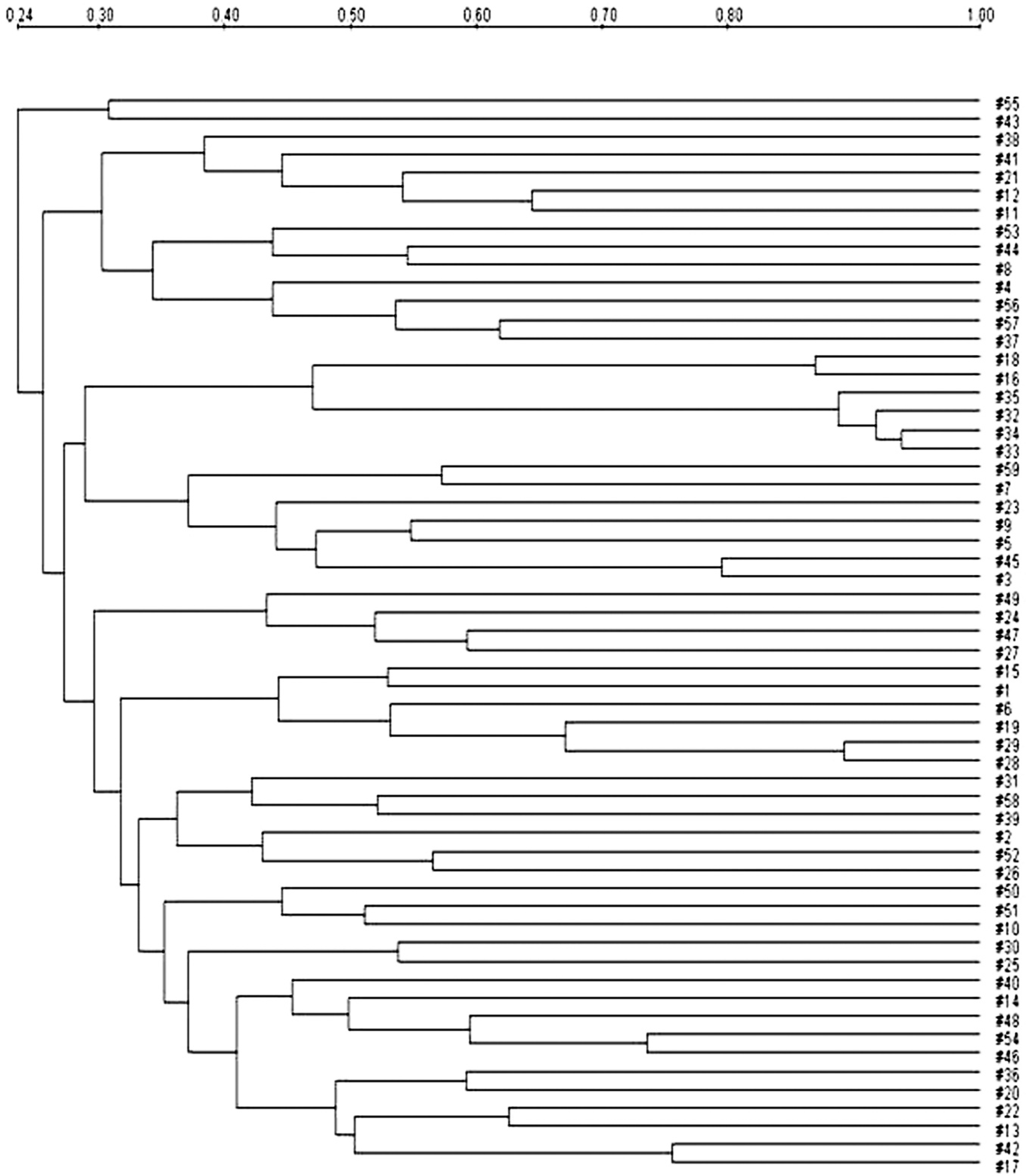

Dendrogram using the Dice coefficient and unweighted pair group method with arithmetic mean analysis showing the relationships between Escherichia coli strains with F1 pili with NotI restriction. The degree of similarity (%) is shown on the scale.
Seventy-four E. coli isolates selected for antimicrobial susceptibility testing were analyzed by PFGE. XbaI or NotI endonuclease was used to produce chromosomal digests with relatively few fragments.
The representative PFGE patterns with XbaI- and NotI-digested genomic DNA are shown in Figures 1 and 2, respectively. The PFGE analysis revealed that each PFGE profile displayed several bands within the 50- to 1,000-kb range. Similarity of PFGE profiles was lower as 20% with XbaI and 24% with NotI as shown by the Dice analysis (Figs. 3, 4). Even though a high degree of polymorphism among strains was observed, there were 4 subclusters having high similarity (Dice coefficient value >0.7). Each of the 4 subclusters consisted of strains carrying the same virulence genes and antibiotic patterns. Of these 4 subclusters, 2 originated from the same piggery during the same time frame.
The isolates showed the diverse patterns of virulence profiles with F1 only as the highest virulence patterns and combination of F1 and EAST1 (Table 1). The isolates of E. coli that harbor genes for adherence factors or enterotoxins are classified as ETEC, 12 and pathogenic ETEC strains that cause colibacillosis must have both of them. 17 However, the purpose of the current study was to find a correlation of virulent pathotypes and antibiograms. Therefore, no virulent pathotype having F1 only was selected for epidemiological examination in this study.
In the antimicrobial resistance test, high resistance to streptomycin, tetracycline, gentamicin, trimethoprim/sulfamexazole, and ampicillin was observed in this study. This result was in agreement with previous studies in Korea. 4,10,14,20 Quinolones, a group of antibiotics, are of special interest in treating Gram-negative infections in the swine industry. The present study showed an increasing percentage of quinolone-resistant E. coli isolates in ciprofloxacin and enrofloxacin. The number of isolates resistant to enroflox-acin increased from 16.9% to 56.7% in previous examinations to 64.4% in the current study. 4,10,20
Although the use of the antimicrobial on food animals was banned in Korea in 1992, resistance to chloramphen-icol was high. Resistance to quinolones was also high. This phenomenon was very similar with the findings of a previous study. 14
A genetic diversity study, carried out using PFGE, has shown high diversity patterns with some regional and genetic relatedness. In PFGE analysis, each of the 4 subclusters consisted of strains carrying the same virulence profiles and antibiotic patterns. Of these 4 subclusters, 2 originated from the same piggery during the same time frame. Therefore, the PFGE method might be applicable for investigating regional source and genetic relatedness.
In conclusion, prevalence of virulence factors of ETEC and antibiotic resistance was observed with significant ratio in piglets with diarrhea. Distribution of ETEC was widespread and antibiotics were used widely in piggeries in Korea. Thus, investigation of both virulence profiles and antibiogram is essential to the epidemiological study of ETEC. In addition, PFGE analysis used in the current study showed high diversity patterns with some regional, genetic, and antibiotic relatedness. Although a high degree of polymorphism among strains having the same virulence patterns was seen, the highest similarity pattern was observed carrying the same virulence profiles and similar antibiogram. This result suggests that PFGE might be applicable as a tool for revealing genetic relatedness among E. coli strains from piggeries in Korea.
Acknowledgements. This study was supported through Biogreen 21 (20070401034009) RDA, BK21 Program for Veterinary Science, KRF (2006-005-J502901), and the Research Institute for Veterinary Science, Seoul National University, Korea.
Footnotes
a.
VITEK®, bioMérieux Inc., Durham, NC.
b.
Bioneer, Daejon, Republic of Korea.
c.
Difco Laboratories Inc., Detroit, MI.
d.
BD, Franklin Lakes, NJ.
e.
Oxoid Ltd., Cambridge, UK.
f.
CHEF-DR® III, Gel Doc 1000, Molecular Analyst® Fingerprinting Plus; Bio-Rad Laboratories, Hercules, CA.
g.
Takara Bio Inc., Otsu, Shiga, Japan.
