Abstract
Staphylococcus aureus is one of the most important etiologic agents of ovine mastitis. To develop effective control measures for mastitis, it is important to type S. aureus strains that have considerable genetic heterogeneity. In the current study, 47 S. aureus strains isolated from ovine mastitis were typed by polymerase chain reaction (PCR) based on coagulase (coa) and protein A (spa) polymorphisms and by pulsed-field gel electrophoresis (PFGE). Eight different coa types and 4 spa types were identified by PCR. While the most prevalent coa type was CG2 (42.56%), the spa types S4 and S1 were the most commonly observed (44.68% and 38.29%, respectively). Nineteen different pulsotypes were identified, and 12 of these were represented by a single isolate. Pulsotypes J and K were predominant and each represented 9 isolates (19.14%). All isolates belonging to J and K pulsotypes were CG2. Although all 9 isolates belonging to the J pulsotype were S4, all isolates in the K pulsotype were S1. While PFGE was found to be the best discriminatory technique for distinguishing strains, coa and spa types were found to be in correlation with PFGE types and can be used for quick, preliminary epidemiologic studies for detecting strains that may cause mastitis.
Mastitis, in clinical or subclinical forms, has a major impact on both economy and animal welfare in sheep production. Although various microorganisms may cause ovine mastitis, Staphylococcus aureus is one of the most important etiologic agents. 15 Accurate and rapid detection and typing of S. aureus is crucial to control the infectious organisms. 10 Molecular typing of S. aureus is a powerful tool that can provide information about genetic characteristics of a specific clone responsible for mastitis. Development of a rational and effective strategy based on genetic variability of S. aureus and directed against clones that commonly cause disease is important for control of mastitis. 12
Coagulase and protein A are important virulence factors and phenotypic determinants of S. aureus. Thus, the coa gene has been a target for polymerase chain reaction (PCR) genotyping, and amplification of this gene has been considered a simple and accurate method for typing S. aureus isolated from distinct sources. 9 Protein A is a membrane-bound exoprotein produced by most of the S. aureus strains. 1 Protein A in S. aureus contains a polymorphic X region characterized by a tandem repeat of 8 amino acid units, and the number of units in the X region is diverse among S. aureus strains.
Pulsed-field gel electrophoresis (PFGE) is accepted as a “gold standard” for typing several Gram-positive and Gram-negative bacteria and some yeast. 8 Pulsed-field gel electrophoresis is a highly discriminatory and sensitive technique for distinguishing strains of S. aureus and has been used for microepidemiologic (local or short-term) and macroepidemiologic (national, continental, or long-term) surveys. 8,13 The purpose of the current study was to type S. aureus isolates obtained from cases of ovine mastitis in Samsun, Turkey, using PCR-based techniques and PFGE. The coa and spa polymorphisms and PFGE were used to identify S. aureus subtypes and to investigate the genotypic relationship between the isolates.
Staphylococcus aureus strains were isolated from sub-clinic mastitic sheep during a 2-year period (2006–2007) in Samsun. The strains were isolated from 47 different folds of sheep. In each fold, different numbers (2–12 strains) of S. aureus strains were isolated. The staphylococcal isolates were identified morphologically and biochemically. 3,17 Bacterial DNAs were extracted according to the manufacturer's instructions a and were kept frozen at −20°C until used.
A multiplex PCR was performed with the oligonucleotide primers b for the detection of 16S ribosomal RNA (rRNA; Staphylococcus spp.-specific) and nuc (S. aureus-specific) genes for identification of S. aureus. 26 Extracted DNA was added to the PCR mixture containing 5 mmol of MgCl2, 0.3 mmol of deoxynucleoside triphosphate (dNTP), 0.8 μmol of 16S rRNA primer, 0.3 μmol of each primer, and 2 units of Taq DNA polymerase. c The amplification was performed as follows: 94°C for 5 min of initial denaturation; 30 cycles of 94°C for 45 sec, 68°C for 45 sec, and 72°C for 90 sec; and a final extension at 72°C for 10 min. The PCR products were loaded onto 1.5% agarose gel containing 1 μg/ml ethidium bromide. The 756-bp (16S rRNA) and 279-bp (nuc) amplified DNA fragments were separated by agarose gel electrophoresis.
For the detection of coa gene polymorphisms, the primers 2 were used to amplify the variable region of the coa gene. For PCR, the reaction mixture contained 2.5 mmol of MgCl2, 50 pmol of each primer, 200 μmol of dNTP, and 1 unit of Taq polymerase. c The amplification was carried out as follows: 95°C for 2 min of initial denaturation; 35 cycles of 95°C for 30 sec, 60°C for 2 min, and 72°C for 4 min; and a final extension at 72°C for 7 min. The PCR products were loaded onto 2% agarose gel containing 1 μg/ml ethidium bromide. The amplified DNA fragments were separated and visualized under ultraviolet (UV) light, and then evaluated with respect to the molecular weight and number of bands. d
For the detection of spa gene polymorphisms, the primers 13 were used to amplify the spa gene. The reaction mixture contained 2.5 mmol of MgCl2, 0.25 mmol of each primer, 250 μmol of dNTP, and 1.5 units of Taq polymerase. c The DNA amplification was carried out as follows: 95°C for 3 min of initial denaturation; 35 cycles of 94°C for 1 min, 60°C for 1 min, and 72°C for 1 min; and a final extension at 72°C for 10 min. The PCR products were loaded onto 3% agarose gel containing 1 μg/ml ethidium bromide. The amplified DNA fragments were separated and visualized under UV light and then evaluated with respect to the molecular weight and number of bands. d
Chromosomal DNA preparation was carried out as described previously. 14 The digestion of DNA was performed with 20 units of the SmaI restriction enzyme. c Digested DNA was electrophoresed in 1.2% agarose gel using a commercial PFGE system. e Electrophoresis was performed as described previously. 22 After electrophoresis, the gel was stained with ethidium bromide for 1 hr, destained in water, and photographed under UV transillumination. The PFGE banding patterns were analyzed by computer software e to define the clusters and pulsotypes, and the patterns were compared by the Dice coefficient method, with a cutoff value of 70%.
The proportions of strains that can be assigned a type by typing methods used in the current study were calculated as described previously. 11,23 The discriminatory power of a typing method was calculated according to a numeric index. 11 The confidence intervals (CIs) of the typing methods were calculated according to the formula described previously. 7 To test PCR (coa and spa typing) reproducibility by interassay analysis, 5 isolates were selected randomly and were tested for 5 consecutive days.
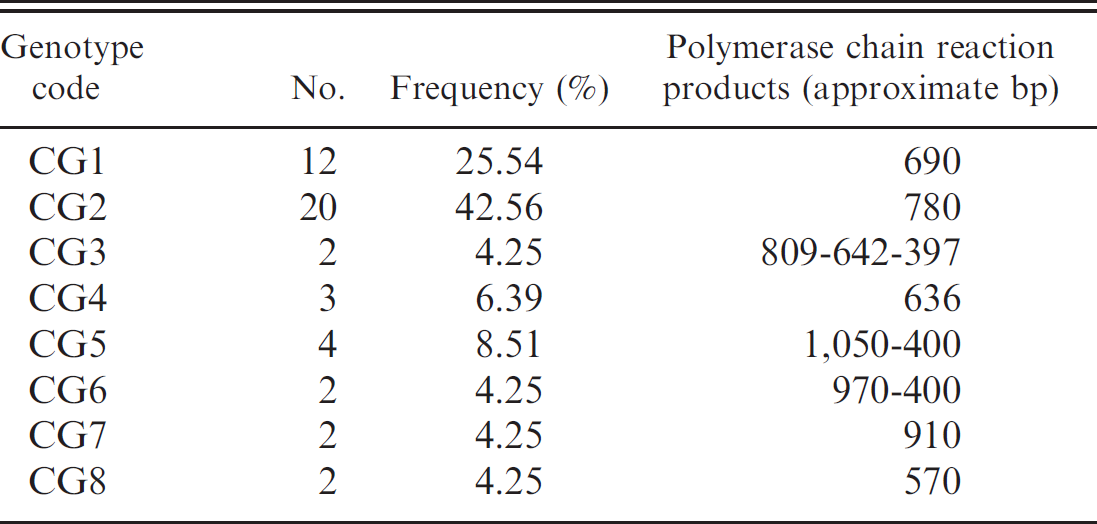
Genotypes of Staphylococcus aureus based on coa polymorphism.
All strains isolated from 47 different folds were identified as S. aureus phenotypically by standard laboratory methods, including catalase, coagulase, and protein A tests. 3,17 All of the strains were catalase, coagulase, and protein A positive. All strains examined were found to be 16S rRNA and nuc positive and were identified as S. aureus strains. All S. aureus strains isolated from each fold were characterized in their PFGE patterns and their coa and spa polymorphisms using the methods described above. The results showed that all strains isolated in the same fold presented identical pulso-types and coa and spa genotypes. Thus, it was considered that 1 fold was represented by 1 strain in the study.
A total of 47 S. aureus isolates from ovine mastitis were evaluated for PCR typing of coagulase gene. The isolates produced PCR products of one or more sizes. Strains were divided into 8 groups (CGs 1–8) based on the PCR product patterns. CG2 was the predominant group and included 20 isolates (42.56%). Strains in 5 groups (CGs 1, 2, 4, 7, and 8) produced a single PCR product, with sizes ranging from 570 to 910 bp. Isolates belonging to CG5 and CG6 exhibited 2 PCR products, and isolates belonging to CG3 exhibited 3 PCR products. Coagulase gene typing results are summarized in Table 1.
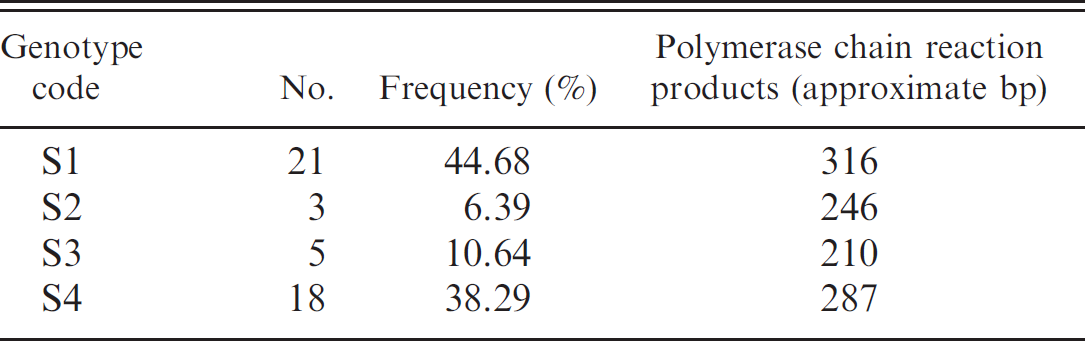
A total of 47 S. aureus isolates were typed based on differences in the X region of the spa gene encoding for protein A by PCR. All isolates produced single PCR products in different sizes ranging from 210 to 316 bp. The results of spa typing by PCR are summarized in Table 2. Isolates were divided into 4 groups according to the spa polymorphisms, where 4 PCR products of different sizes were observed after amplification of the spa gene. Most isolates (44.68%) were assigned to the S1 group with 11 repeats, followed by the S4 group (10 repeats), which included 18 isolates (38.29%). Other types (S2 and S3) had lower numbers of repeats (8 and 7, respectively).
Genotypes of Staphylococcus aureus based on spa polymorphism.
Correlations between the pulsed-field gel electrophoresis (PFGE) pulsotypes and the results of polymerase chain reaction-based typing methods.*
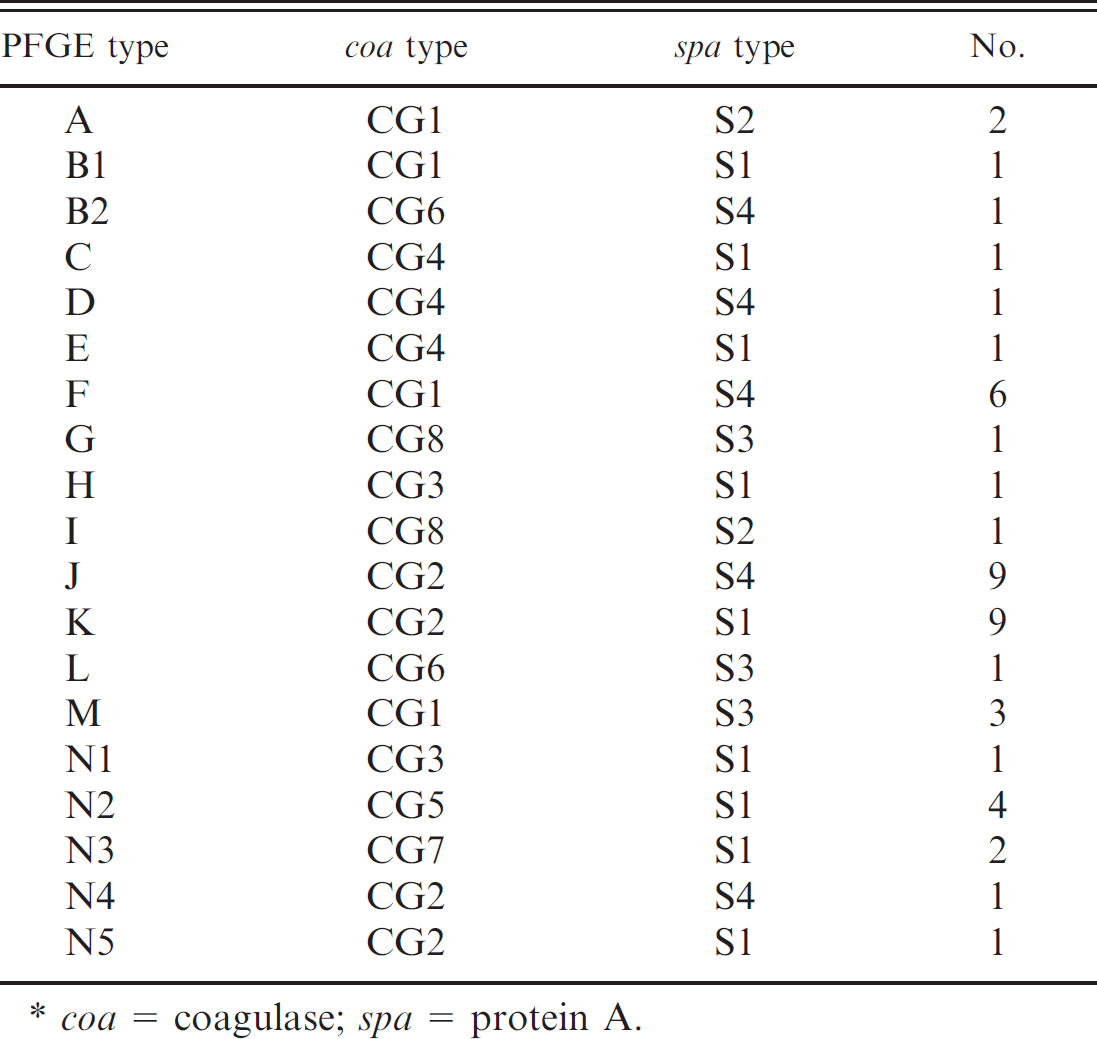
coa = coagulase; spa = protein A.
The PFGE analysis of isolates exhibited 14 clusters and 19 pulsotypes. Pulsotypes J and K were the predominant types, and each included 9 isolates (19.14%). However, 12 of the 19 pulsotypes included only 1 isolate. The PFGE profiles of selected S. aureus strains are shown in Figure 1 and Table 3. Relatedness among the 47 S. aureus strains was evaluated and is depicted in a dendrogram (Fig. 1).
All strains of the pulsotypes J and K belonged to a single coagulase type, CG2. Eleven isolates (23.40%) representing the CG1 type were distributed among A (4.25%), F (12.76%), and M (6.38%) pulsotypes. However, all strains belonging to pulsotype J represented the S4 type, whereas strains belonging to pulsotype K were represented as S1 (Table 3).
All S. aureus isolates produced amplified product(s) in both coa and spa PCR, reflecting that these were typeable. Discriminatory indices for PCR-based coa and spa typing were 0.75 (CI = 65.8–84.1%) and 0.65 (CI = 57.68–72.3%), respectively. The discriminatory index for PFGE was 0.909 (CI = 86.8–94.9%). To determine the reproducibility of PCR for coa and spa typing by interassay analysis, 5 isolates were selected randomly and were tested for 5 consecutive days. The reproducibility of PCR-based typing methods (coa and spa typing) was 100%.
Molecular methods based on DNA amplification by PCR for typing S. aureus strains have been developed during the last decade. In the current study, PCR-based typing methods targeting coa and spa genes, together with PFGE, were evaluated for typing S. aureus isolates from cases of ovine mastitis in different farms. One of the bacterial typing techniques, amplification of a single target gene specific to a pathogen, has been reported as moderate to highly discriminative. 5 The coa typing of S. aureus isolates from various mastitis cases has been carried out, and it has been reported that coagulase gene typing is a useful method for clinical laboratories because of its ease and speed. 21 In the present study, 47 S. aureus isolates could be classified into 8 groups based on the polymorphism of coa gene products by PCR. A 780-bp PCR product was found to be prevalent in 42.56% of isolates. In one study, 19 it has been found that 31 isolates (80.6%) from sheep milk exhibited PCR products 820 bp in size. There are no comparable data available in the literature for S. aureus strains isolated from the milk of small ruminants. It was reported that PCR-amplified coa genes from bovine S. aureus strains mostly showed 580-bp PCR products. 18 In other studies carried out with bovine mastitis isolates, 19,20 different numbers and sizes of PCR products have been reported. Therefore, there is a certain degree of genetic heterogeneity among S. aureus isolates from sheep mastitis. A previous study 24 has reported that predominant genotypes of S. aureus may vary in different countries or geographic locations. In the current study, although the most common coa type, CG2, was found in the north and south regions of Samsun, the second common coa type, CG1, was distributed in all 4 regions of Samsun.
The spa gene is composed of approximately 2,150 bp and harbors a number of functionally distinct regions, including the X region. The X region contains a varying number of 24-bp repeats, and this variability has been used as a discriminative marker with good correlation with PFGE typing. 8,25 The number of repeats in this region may allow strain discrimination. 6 It has also been reported that the number of repeats within the spa X region is related to both the epidemic and the dissemination potential of strains. Strains with higher numbers of repeats in X region of spa have higher epidemic capability. 13 In the same study, it was reported that most (24/33) of the epidemic methicillin-resistant S. aureus strains have more than seven 24-bp repeats in the X region of the spa gene, and that 10 of the 14 nonepidemic strains show 7 or fewer repeats. 6 In the current study, 4 different spa genotypes were found. Among these groups, the S1 and S2 types were responsible for the vast majority of mastitis cases (44.68% and 38.29%, respectively). Strains in the S1 type with 11 repeats and strains in the S4 type with 10 repeats were considered to have more epidemic potential than others. However, all strains may not comply with this rule.
In the current study, the 47 S. aureus isolates from cases of ovine mastitis could be grouped into 19 different types (pulsotypes) by PFGE. The most common pulsotypes were J and K, each consisting of 9 isolates. The similarity between these 2 types was <20%, as derived from the dendrogram. Furthermore, these pulsotypes were unique to a certain geographic region. Although pulsotype J was found in the northern part of Samsun, pulsotype K was found in the southern part of Samsun. Five pulsotypes were observed in low numbers (2–6 isolates), and 12 pulsotypes were represented by a single isolate. These results show that a great number of S. aureus pulsotypes are responsible for mastitis in sheep. In other words, a wide genetic variability was found among the 47 S. aureus isolates investigated. The 5 pulsotypes showing >80% similarity in cluster N were prevalent in all regions. This may be due to movement of sheep from one geographic location to another. In spite of the uncontrolled movement of sheep between regions, predominant pulsotypes J and K were found to be restricted to the northern and southern regions of Samsun, respectively.
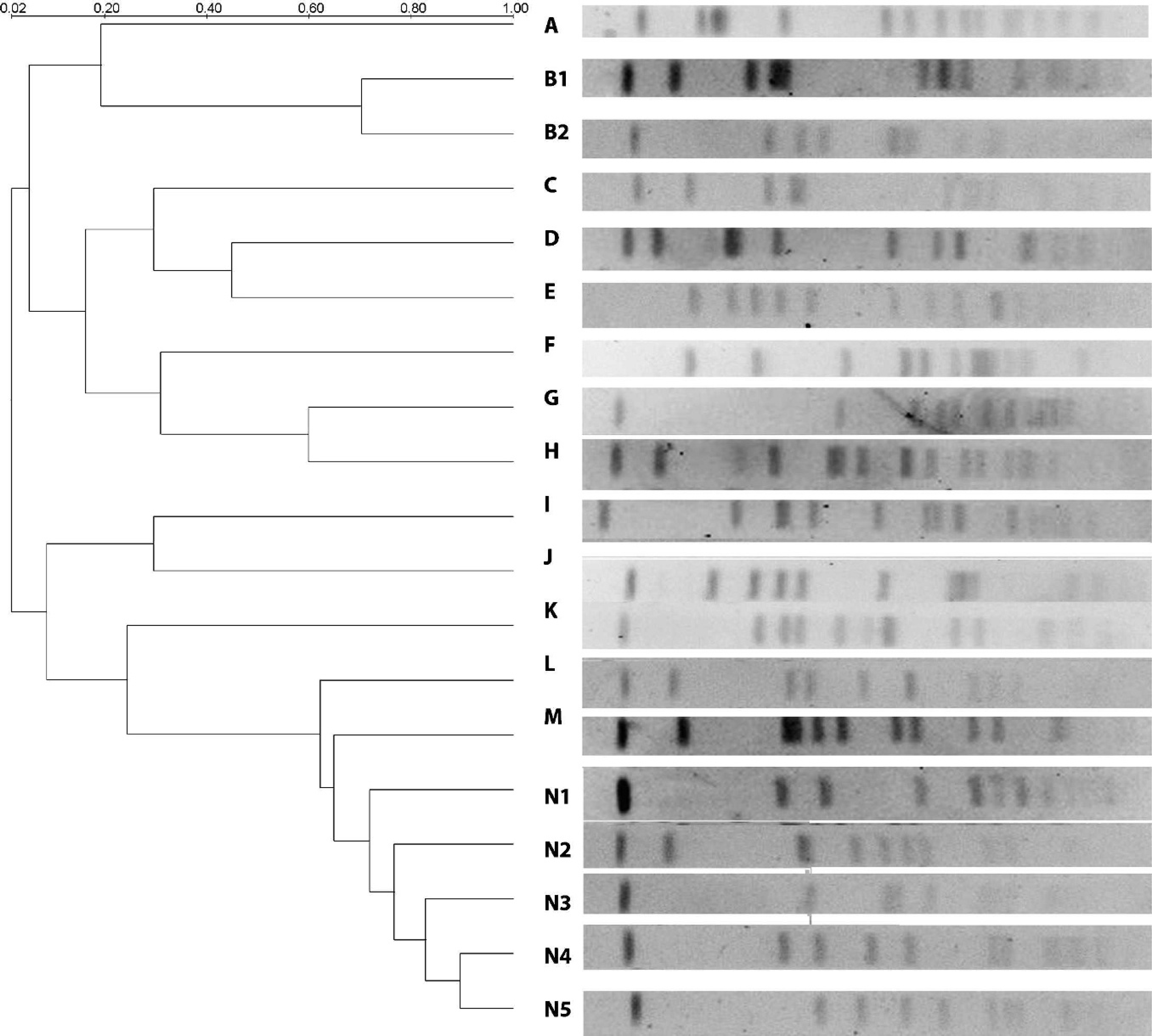

The phylogenetic tree based on the Dice coefficient of pulsed-field gel electrophoresis pattern similarity.
A discriminatory index value of 1.0 would indicate that a typing method was able to distinguish each member of a strain population from all other members of that population. Conversely, a discriminatory index of 0.0 would indicate that all members of a strain population were of an identical type. Pulsed-field gel electrophoresis is currently regarded as the molecular typing gold standard 16 and is highly discriminatory and reliable for both the identification of strains at the subspecies level and the determination of interstrain genetic relatedness. 8,23 In the current study, the discriminatory index values of 3 typing methods—coa PCR typing, spa PCR typing, and PFGE—were 0.75, 0.65, and 0.909, respectively. Similarly, the discriminatory index of PCR-based typing methods targeting the coa gene has been found by a previous study 4 to be 0.75. Also, another study 8 has reported a high discriminatory index (above 0.95) for PFGE typing of S. aureus. Pulsed-field gel electrophoresis was found to be the best discriminatory typing method and gave a wider spectrum of types than the other techniques.
Good correlation between coa types and PFGE types was found in the current study. All pulsotypes correlated with a single coa type. Similarly, each pulsotype was represented by a single spa type. However, isolates grouped in 5 N subtypes (N1–N5) were found to have different coa and spa types.
Results of the present study showed that among the typing methods used, PFGE typing was the best discriminatory method. However, coa and spa PCR typing methods were considered as alternative methods that provide useful information as the first approach in investigating the genotyping of the strains. These PCR-based typing methods may give a preliminary opinion about the genotypic discrepancies of S. aureus involved in mastitis cases of sheep. Furthermore, these typing methods were simple, inexpensive, and quicker than PFGE. Although the results showed that the predominant types were found only in the northern and southern regions of Samsun, the distribution of the types of S. aureus isolates between regions or counties should be evaluated with more isolates from different regions. In conclusion, further studies using a large collection of S. aureus strains from distinct areas should be carried out for determining common characteristics of the predominant strains and developing efficient control measures of mastitis caused by S. aureus.
Footnotes
a.
PureLink™ Genomic DNA Kits, Invitrogen Canada Inc., Burlington, Canada.
b.
Thermo Hybaid, Ulm, Germany.
c.
Buffers and enzymes used in all of the PCR assays were obtained from Fermentas Life Sciences, Vilnius, Lithuania.
d.
Molecular Imaging Software, Eastman Kodak, Rochester, NY.
e.
CHEF-DR® III, Quantity One® Software; Bio-Rad Laboratories, Hercules, CA.
