Abstract
In the current study, the virulence factors in Escherichia coli isolates from bovine mastitis were investigated, and the connection between these factors and infection was evaluated using phenotypic and genotypic analyses. Twenty-seven E. coli isolates were analyzed, and 2 were shown to produce verotoxin. All isolates had the ability to produce biofilms, although at different levels. One isolate was found to be sensitive to the bactericidal activity of bovine serum, 11 were intermediate, and 15 were resistant. Some isolates showed resistance to trimethoprim sulfa (9) and ampicillin (4), intermediate resistance to neomycin (1) and trimethoprim sulfa (5), and simultaneous resistance to ampicillin and trimethoprim sulfa (4). The fimH gene was found in all isolates and was associated with other virulence markers: pap (1), stb (8), cs31a (3), stb and vt2 (2), cs31a and stb (3), east1 and kps (1), stb and east1 (1), cs31a and east1 (1), and cs31a, stb, pap, and iucD (1). Serogroups were determined for 3 isolates: O93:H4, O83:H19, and O15:H11. Phylogenetic analysis showed that 23 isolates belonged to group A and 4 belonged to B1. The findings revealed that these E. coli isolates are opportunistic pathogens with different virulence factors. The results indicate that the pathogenicity route of E. coli in bovine mastitis is not a consequence of 1 specific virulence factor.
Introduction
Bovine mastitis is one of the most complex and costly diseases in the dairy industry due to its high prevalence and economic losses. 39 There is also a potential risk to public health because it may transmit zoonoses and food toxin infections. 9 Mastitis triggered by Escherichia coli is usually sporadic, and clinical signs vary from very severe or even fatal forms to mild mastitis 41 in which cows have only local signs in the udder. While the severity of the disease depends on host immune response and genetic makeup (“cow factors”), 11 virulence of the bacterial strain involved may also play a role.
The E. coli strain that causes mastitis can potentially form a new putative pathotype, mammary pathogenic E. coli, 41 and make use of the same pathogenesis scheme used by other extraintestinal pathogenic E. coli (ExPEC). There are, however, many publications on the absence of known ExPEC virulence-associated genes and intestinal pathogenic E. coli among E. coli mastitis isolates, which indicates that there may be different ways to cause mastitis. 41
Several virulence factors have been detected in pathogenic E. coli. These include toxins, adhesins, invasins, capsule production, the ability to resist serum complement, and iron scavenging. Only isolates with successful combinations of virulence factors will be capable of causing disease. 25 Bovine mastitis resembles a urinary infection in that the infection ascends and is caused by bacteria from the environment. Bacterial virulence factors are required to fight the host’s selection pressure and for the bacteria to colonize, multiply and survive in the udder. 24 The difficulties of treating recurrent infections might be related to the ability of pathogens to form biofilms, although little research exists on biofilms in animals. 28
Considering the losses resulting from this disease, the recognition of E. coli as a highly adaptive organism in different niches, the inefficiency of treatment, and the impact of this infective agent on public health, the present study aimed to investigate the presence of virulence factors in E. coli isolates from bovine mastitis in an attempt to correlate these characteristics and establish a possible connection with mastitis.
Material and methods
Escherichia coli isolates and serology
The current study involved 7 dairy farms located in the Viçosa and Juiz de Fora regions in the State of Minas Gerais, Brazil. Mastitis was defined by classical symptoms. All cows with symptoms of acute clinical environmental mastitis were examined by veterinarians who diagnosed the illness. Animals showed repetitive episodes of illness, without death. One pooled milk sample was aseptically collected once from each selected animal, yielding a total of 27 samples. The collections were performed before treatment with antibiotics. One E. coli isolate was obtained from each sample using a previously published protocol. 22 The samples and controls were kept in brain heart infusion (BHI) a with glycerol (10%) at −80°C until analysis. Each culture was streaked onto BHI agar plates (37°C for 24 hr), and a single colony was inoculated in BHI and incubated at 37°C by shaking until achieve turbidity similar to tube 1 on the McFarland scale, corresponding to approximately 3 × 108 colony forming units per ml (CFU/ml). All isolates were submitted to serological testing using somatic (O1-O172) and flagellar (H1-H50) antisera as previously described. 19
Verotoxin production
The cultures were analytically tested to verify verotoxin production using a previously described method 36 with some modification of the use of essential medium, b with 1.6 mg/l penicillin, 0.4 mg/l streptomycin, and 5% fetal bovine serum. b The morphological changes were evaluated at 24 and 48 hr of incubation. Verotoxin-producing isolates were characterized by initial cell rounding and shrinkage, followed by loss of cell viability. The positive controls used were E. coli O157:H7 c and E. coli J2 d for vt1 and vt2 genes, respectively.
Antimicrobial resistance
The disc diffusion method, 3 with modifications described by the Clinical and Laboratory Standards Institute (CLSI), 15 was performed with antimicrobials most prescribed by veterinarians working in regions where the samples were collected: ampicillin and gentamicin (10 µg each), trimethoprim sulfa (25 µg), neomycin, cephalexin, ceftiofur, and florfenicol (30 µg each), and cefoperazone (75 µg). The zones of inhibition were measured, taken as the average of triplicates, and compared with the break points presented in the Table 1, adapted from CLSI.14,15 The quality control strain used was E. coli American Type Culture Collection (ATCC) 25922.
Antimicrobisals, concentrations, and interpretation of inhibition zones for the evaluation of susceptibility patterns of Escherichia coli.
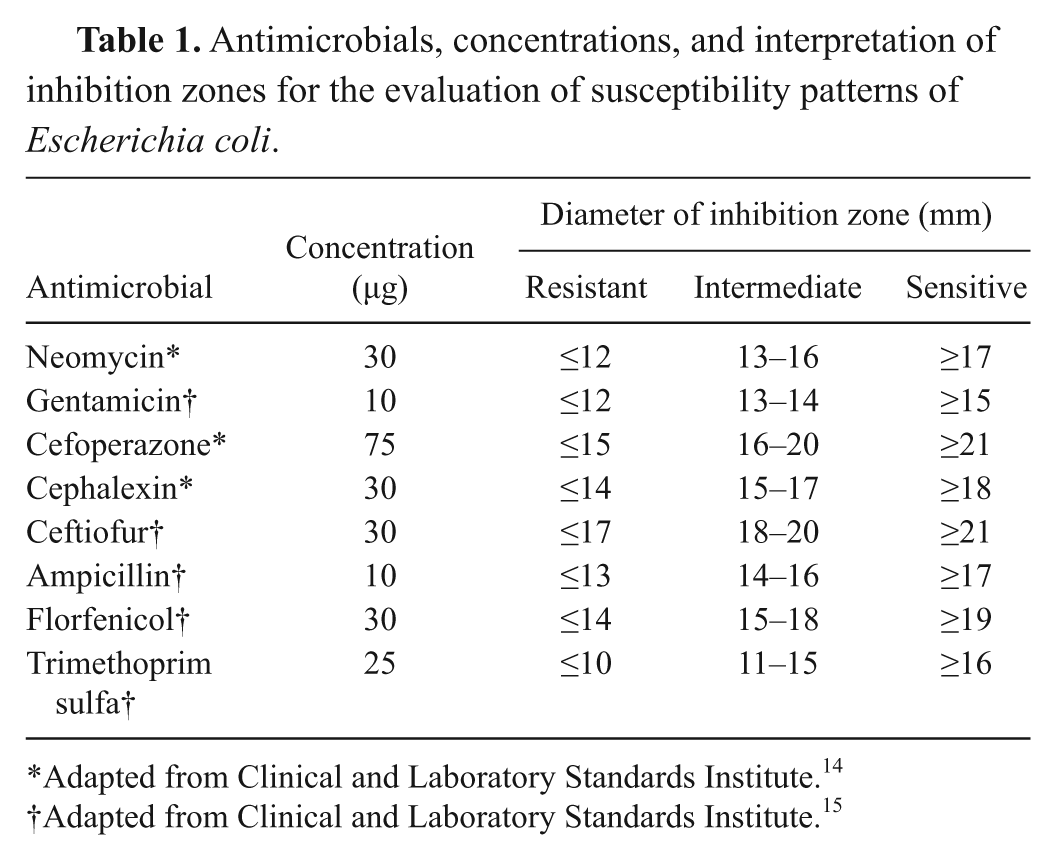
Adapted from Clinical and Laboratory Standards Institute. 14
Adapted from Clinical and Laboratory Standards Institute. 15
Resistance to lytic activity of serum complement
A turbidimetric assay 33 was employed to test the resistance of the cultures to serum bactericidal activity. Escherichia coli strain J-96 was used as the positive control. The isolates that showed optical density at 630 nm (OD630) values below 50% of the initial value were considered sensitive to the bactericidal activity. Isolates with OD630 values above 90% of the initial value were considered resistant, and those that remained between these values were considered intermediate.
Biofilm production
Quantification of biofilm production in plastic microplates was performed as previously described. 43 The OD570 of each well was measured using a microplate reader, e and the means of the triplicates were calculated. The isolates were classified as non-producers, weak, moderate, or strong biofilm producers based on OD of the isolates and the average OD570 of the negative control, according to the methodology used. 43 Briefly, the cut-off OD (ODc) was defined as 3 standard deviations above the mean OD of the negative control. Strains were classified as follows: OD ≤ ODc = no biofilm producer, ODc < OD ≤ 2 ODc = weak biofilm producer, 2 ODc < OD ≤ 4 ODc = moderate biofilm producer, and 4 ODc < OD = strong biofilm producer.
Polymerase chain reaction
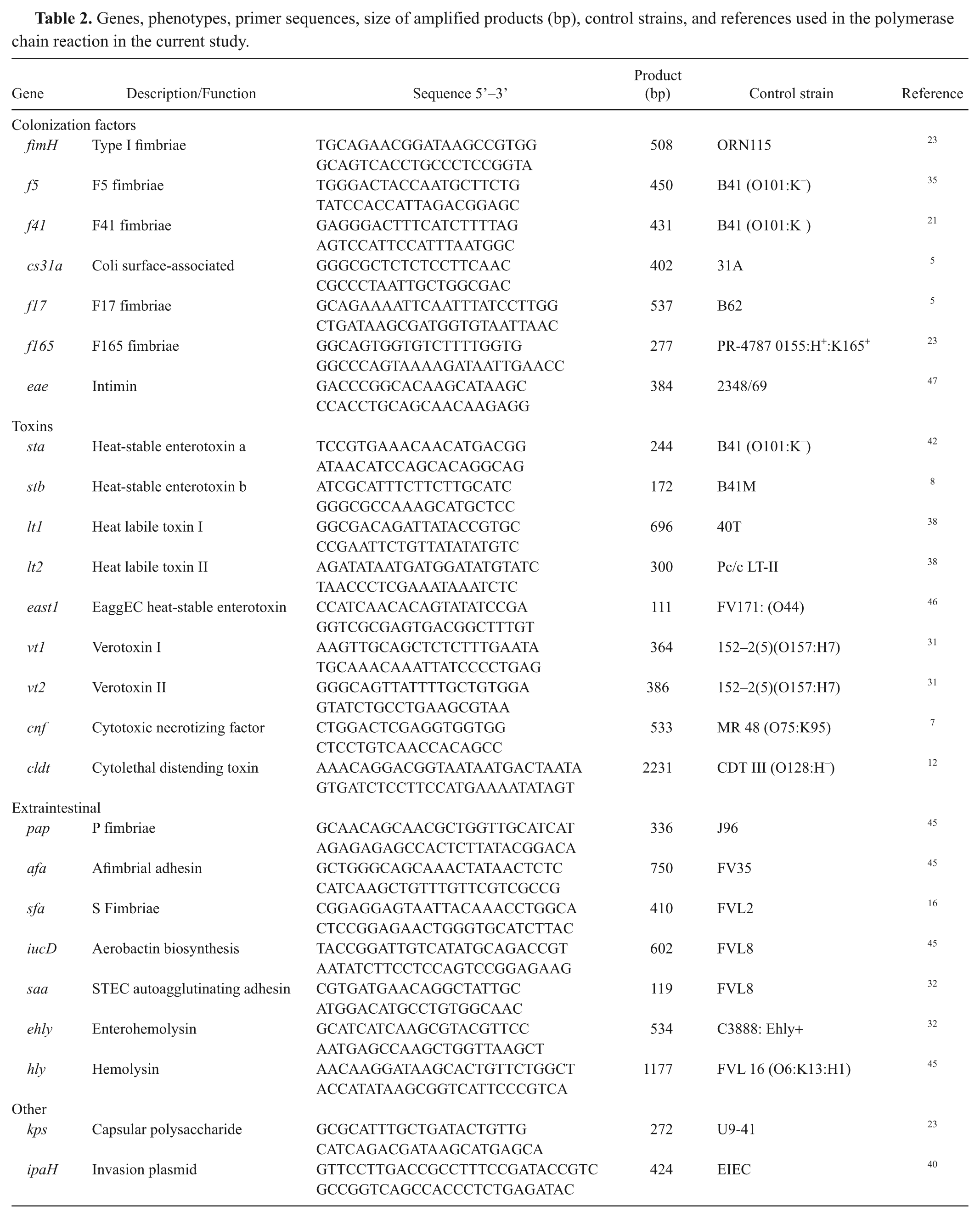
Virulence marker genes were detected by polymerase chain reaction (PCR) at the Laboratory of Bacterial Antigens II, Department of Microbiology and Immunology, Instituto de Biologia, Universidade Estadual de Campinas (Campinas, São Paulo, Brazil; Table 2). The major E. coli phylogenetic groups (A, B1, B2, and D) were determined by triplex PCR as described previously. 13
Genes, phenotypes, primer sequences, size of amplified products (bp), control strains, and references used in the polymerase chain reaction in the current study.
Results
Only 2 cultures, E. coli 25 and 3888, showed an ability to produce verotoxins, which was confirmed by the presence of the vt2 gene. For these cultures, initial cell shrinkage and rounding, followed by loss of cell viability was observed. The most significant cytotoxic effects were observed at the lowest filtered dilution and at 48 hr of incubation.
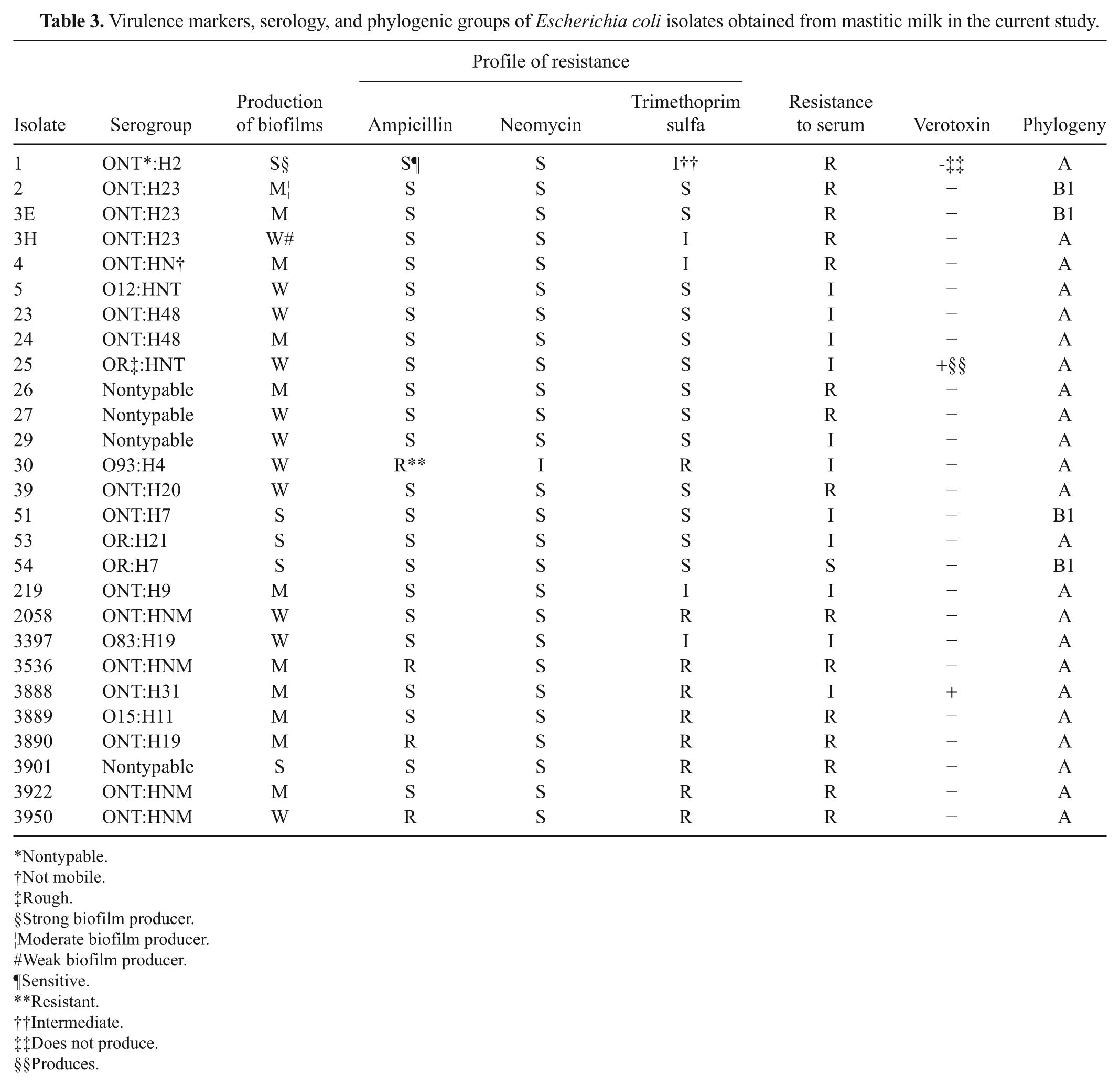
The ability to form biofilms in vitro was detected in all of the isolates, but at different levels of production. Five isolates were considered as strong producers, 11 were moderate producers, and 11 were weak biofilm producers (Table 3). Among the tested antimicrobials, resistance was observed to trimethoprim sulfa (9) and ampicillin (4), and intermediate resistance was observed to neomycin (1) and trimethoprim sulfa (5). Four isolates were resistant to both ampicillin and trimethoprim sulfa (Table 3). According to the growth patterns observed in bovine serum, the isolates were divided into 3 groups depending on their resistance to lytic activity: sensitive, intermediate, and resistant. Only 1 isolate, E. coli 54, was sensitive, 11 were intermediate, and 15 were resistant (Table 3). Serotyping was only possible for 3 cultures, which were identified as O93:H4, O83:H19, and O15:H11 (Table 3). Somatic antigen identification was possible for 4 isolates (O12, O15, O83, and O93), and flagellar identification was possible for 16 isolates (H2, H4, H7, H9, H11, H19, H20, H21, H23, H31, and H48). The flagellar antigen H23 was identified in 3 isolates, followed by H7, H19, and H48 in 2 isolates each, and the other flagellar serogroups were found in 1 isolate.
Virulence markers, serology, and phylogenic groups of Escherichia coli isolates obtained from mastitic milk in the current study.
Nontypable.
Not mobile.
Rough.
Strong biofilm producer.
Moderate biofilm producer.
Weak biofilm producer.
Sensitive.
Resistant.
Intermediate.
Does not produce.
Produces.
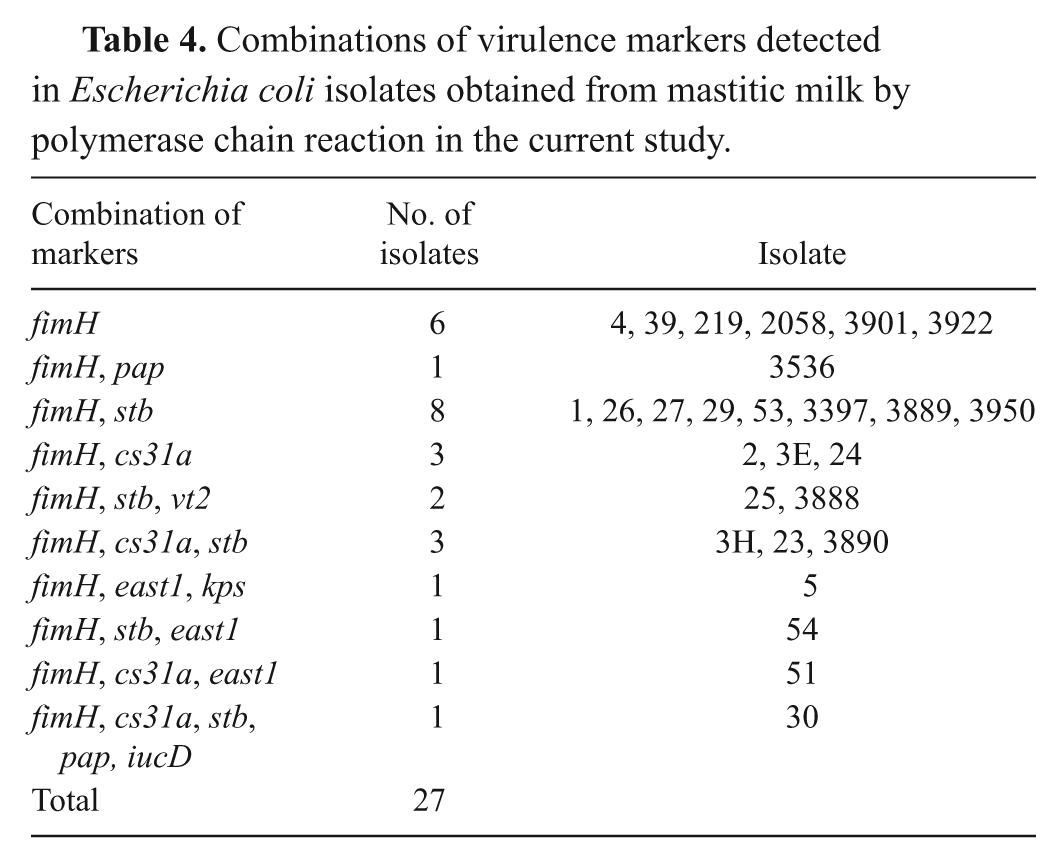
The E. coli isolates were classified in the following phylogenic groups: 23 isolates belonged to group A and 4 to group B1 (Table 3). The fimH gene was the only virulence-associated gene detected in all of the isolates. None of the isolates had the genes for f5, f41, f17, f165, eae, sta, lt1, lt2, cnf, cldt, pap, afa, sfa, saa, ehly, hly, or ipaH. All of the remaining genes were detected in combination with fimH and others. The combination of fimH and sta genes was the most frequent, found in 8 isolates. Full details of the other gene combinations detected are presented in Table 4.
Combinations of virulence markers detected in Escherichia coli isolates obtained from mastitic milk by polymerase chain reaction in the current study.
Discussion
In the current study, 7.5% (2/27) of the isolates were cytotoxic in Vero (African green monkey kidney epithelial) cell culture, confirmed by molecular detection of the vt2 gene (Table 4). Earlier studies4,34 have also detected a low frequency of verotoxin producers obtained from mastitic milk and a greater predominance of the vt2 gene in relation to vt1. Verotoxin-producing strains or Shiga toxin-producing are considered to be pathogenic to human beings if other factors are associated with this virulence. 26
Simultaneous resistance was observed for only 2 antibiotics, ampicillin and trimethoprim sulfa, in 14.8% of the isolates, a common resistance profile in E. coli isolates in other studies.27,37 In contrast, a high level of antimicrobial resistance to many antibiotics and an elevated number of multi-resistant strains among mastitis-associated E. coli strains have been reported in an earlier Brazilian study. 34 Similar reports of high levels of resistance to antimicrobials exist.1,27,37 The findings of the present study suggest that, on the properties sampled, there may not have been selective pressure caused by antibiotic use.
The current study found that 55.5% of the isolates were resistant to serum. However, the isolates with intermediate behavior (40.7%) may be considered as “potentially” serum-resistant, expanding the resistance profile to almost all of the isolates since only 1 was considered as authentically sensitive. An earlier study 30 found serum-resistant isolates (64%) and related the different values found in other studies to possible variations in methodologies and serum concentrations used. There is disagreement in the literature on whether serum resistance is an important virulence factor in mastitis E. coli isolates and if there is a correlation between serum resistance and antimicrobial resistance.2,20,29 In the present study, no relationship between serum and antimicrobial resistance was observed.
Biofilms in animals are believed to be involved in many diseases, such as mastitis. 28 All of the tested isolates were able to produce biofilms in vitro. Recurrent clinical coliform mastitis can also occur as a result of the persistence of the organism within the mammary gland, 10 and the production of biofilms can provide suitable conditions for the persistence of bacteria at this site.
Several studies have attempted to correlate virulence factors with the etiology of bovine mastitis.4,24 The results obtained in the present study showed a combination of different virulence factors in the E. coli isolates (Table 4). The most common adhesins found in both commensal and pathogenic E. coli isolates as well as in other enterobacteria are type 1 fimbriae (fimH gene), and these have influence on biofilm formation. 44 The finding was consistent with the current study, as the fimH gene was detected in all isolates studied.
An earlier study 17 failed to detect virulence factors in E. coli associated with persistent bovine mastitis, among these the stb and pap genes, unlike the current study where both these genes were detected. The cs31a gene (afimbrial adhesin cs31a) has not been commonly detected in earlier studies of mastitis-linked E. coli. 24 In contrast, in the present study, this gene was detected in 8 out of 27 isolates. In the present work, only 3 isolates had the east1 gene, a gene present in enteroaggregative E. coli 8 as well as E. coli isolated from cattle with septicemia and diarrhea. 5
The majority of isolates (24/27) in the present study belong to phylogenetic group A (Table 3), where most commensal strains are grouped.6,23 It was anticipated that the isolates of E. coli obtained from bovine mastitis milk would belong to group B2 or D, the virulent extraintestinal strain groups.6,23 Farm animal E. coli microbiota are characterized by a high proportion of A and B1 strains and a lower proportion of B2 and D strains. 18
The results obtained in the present study indicate that the pathogenic role of E. coli in bovine mastitis is not a consequence of specific virulence factors. Based on the virulence factors examined, it was not possible to group these isolates into pathotypes able to cause mastitis. Further studies using other virulence factors are necessary to elucidate the relationship between the presence of these factors in E. coli isolates from mastitis and their classification in distinct pathotypes and to investigate commensal strains of E. coli as possible agents of mastitis.
Footnotes
Acknowledgements
The authors would like to thank Professor Domingos da Silva Leite (State University of Campinas) for kindly providing the control strains of E. coli and the PCR analysis, Dr Maria Aparecida Vasconcelos Paiva Brito (Empresa Brasileira de Pesquisa Agropecuária) for providing the E. coli isolates from clinical mastitis, Professor Beatriz E. C. Guth (Universidade Federal de São Paulo) for help in serology, and Professor Rosa Maria Silva (Universidade Federal de São Paulo) for kindly providing the E. coli J96.
Notes
The authors would like to thank FAPEMIG (Fundação de Amparo a Pesquisa do Estado de Minas Gerais) for financing the publication of this article. Isabel A. Carvalho is supported by CAPES (Coordenação de Aperfeiçoamento de Pessoal de Nível Superior), and Luis A. Nero is supported by CNPq (Conselho Nacional de Desenvolvimento Científico e Tecnológico) and FAPEMIG.
