Abstract
A multiplex polymerase chain reaction (mPCR) assay was developed for detection and characterization of pathogenic Escherichia coli that cause diarrhea and edema disease in swine. The mPCR assay was designed as a single reaction for detecting 5 different adhesins (K88, K99, 987P, F41, and F18), 3 enterotoxins (LT, STaP, and STb), and the Shiga toxin (Stx2e) associated with porcine pathogenic E. coli. The specificity of the mPCR assay was evaluated by comparison with results from previous analysis of 100 porcine isolates characterized by colony blot hybridization with DNA probes for the 5 adhesins and 4 toxin genes. There was complete agreement between the 2 methods. The mPCR assay for E. coli pathogens isolated from swine was further evaluated by examination of strains containing virulence factors that are known to have different antigenic subtypes or DNA sequence variations. It was found that the mPCR assays targeting genes encoding for K88 and F18 amplified products with the appropriate sizes from strains containing genes for different K88 and F18 antigenic subtypes; mPCR assays targeting the gene encoding for STaP amplified product from only STaP-positive but not STaH-positive isolates; and mPCR assays targeting the gene encoding for the Stx2 amplified products from only Stx2-positive and not Stx1-positive isolates. Similarly, mPCR assays targeting the gene encoding for LTI did not produce the appropriate product from strains containing genes for LTII. The mPCR assays are simple to perform, and they should be useful for diagnosis of porcine colibacillosis, including the genotypic characterization of E. coli isolates from pigs with diarrhea or edema disease.
Keywords
Introduction
Pathogenic Escherichia coli cause several different diseases in swine that are collectively called colibacillosis. 39 Among these, diarrhea in neonatal and recently weaned pigs caused by enterotoxigenic E. coli (ETEC) is frequently responsible for significant morbidity and mortality. 14 Edema disease (ED) can occur following weaning and is also caused by pathogenic E. coli. 3,19 Both ETEC diarrhea and ED are important economic problems for swine producers. There are effective vaccines available that protect neonatal pigs from diarrhea caused by ETEC, but there are no vaccines currently available to protect weaned pigs from diarrhea or ED. 29 Despite the availability of ETEC vaccines, there remains a need for a simple rapid diagnostic assay for the identification and characterization of pathogenic E. coli isolates from swine.
The pathogenesis of ETEC and ED has been studied extensively, and multiple virulence factors have been identified and characterized. 29 The most intensively studied virulence factors include several antigenically and genetically distinct adhesins and toxins. The well-characterized adhesins are filamentous surface structures that facilitate adherence in the small intestine and are also called pili or fimbriae. The majority of ETEC isolates for swine express 1 or more of 5 antigenically distinct adhesins called K88 (F4), K99 (F5), F41, 987P (F6), and F18. 28 Other suspected adhesins have been identified, but their role in pathogenesis is less clear, and they appear to occur less frequently. The heat-labile (LT) and heat-stable (ST) enterotoxins are responsible for the fluid secretion that results in diarrhea in neonatal and weaned pigs. 29 Two types of LT have been described (LTI and LTII), and LTI is commonly associated with porcine ETEC. 39 The ST toxins are subdivided into STa (STI) and STb (STII). The STa enterotoxins are further subdivided into those primarily associated with human ETEC (STaH) and STaP, which was originally identified in porcine ETEC but is also found in human ETEC isolates. 26 The STb enterotoxin is the most frequent toxin found in porcine ETEC from either neonatal or weaned pigs with diarrhea, 4,25 although the role for STb in neonatal diarrhea caused by ETEC is questionable. 6 In ED isolates, the primary known virulence factors are the F18 adhesin and Stx2e toxin, 19 but other adhesins and enterotoxins may also be present in E. coli associated with ED. 27
Diagnosis of neonatal and postweaning colibacillosis is currently and most often based on clinical observations, histologic examination of small intestinal sections for adherent bacteria, bacterial isolation, and characterization of virulence factors by phenotypic or genotypic assays. 13 Methods for genotypic characterization of ETEC and ED isolates for specific virulence factors include the use of specific DNA fragments with colony blot hybridization and, more recently, polymerase chain reaction (PCR), including real-time PCR, has also been used. 29 Many different PCR assays for E. coli virulence genes in porcine isolates have been reported and used in studies examining the prevalence of these genes in porcine E. coli isolates. These include PCR assays formatted as single reactions or multiplex for the detection of multiple virulence genes and product detection by either real-time or gel electrophoresis methods. 4,8,11,16,30,31,38,40 A variety of different known or suspected E. coli virulence genes are targeted in these PCR assays. Some of these targeted genes occur infrequently, and their significance to pathogenesis of porcine diarrhea or edema disease is not known. Although these are useful for studies on the prevalence of known or suspected virulence genes, they may be less useful for routine diagnostic characterization of the most common virulence genes in E. coli pathogens from pigs.
The objective of the current study was to develop a multiplex PCR (mPCR) assay for identification of the major virulence genes of porcine pathogenic E. coli, including genes for 5 different adhesins (K88, K99, F41, 987P, and F18) and 4 different toxins (LTI, STaP, STb, and Stx2e) associated with pathogenic E. coli isolates from swine. This mPCR assay was evaluated using ETEC, ED, and nonpathogenic isolates and by comparing the PCR results with results from previously conducted experiments by DNA colony blot hybridization using specific DNA probes for the 9 virulence genes. The specificity of the mPCR assay was also determined by examining strains expressing virulence genes that are known to vary in DNA sequence, including K88 (K88ab, K88ac, and K88ad) and F18 (F18ab and F18ac) adhesions, and STa (STaP and STaH), LT (LTI and LTII), and Stx (Stx1 and Stx2) toxins.
. Characteristics of porcine Escherichia coli isolates compared by multiplex polymerase chain reaction and colony blot hybridization for 9 virulence genes.

Materials and methods
The E. coli isolates used in the current study are part ofa collection at the National Animal Disease Center and are stored in 20% glycerol at −70°C. A total of 100 porcine isolates were selected to represent a variety of different sources and different combinations of virulence factor genes. Some of the characteristics of these isolates are listed in Table 1. Additional porcine and human isolates carrying genes encoding for K88, F18 antigenic variants, STaP, STaH, Stx1, and Stx2 subtypes were also examined. Bacteria were routinely grown on lysogeny broth agar. 23
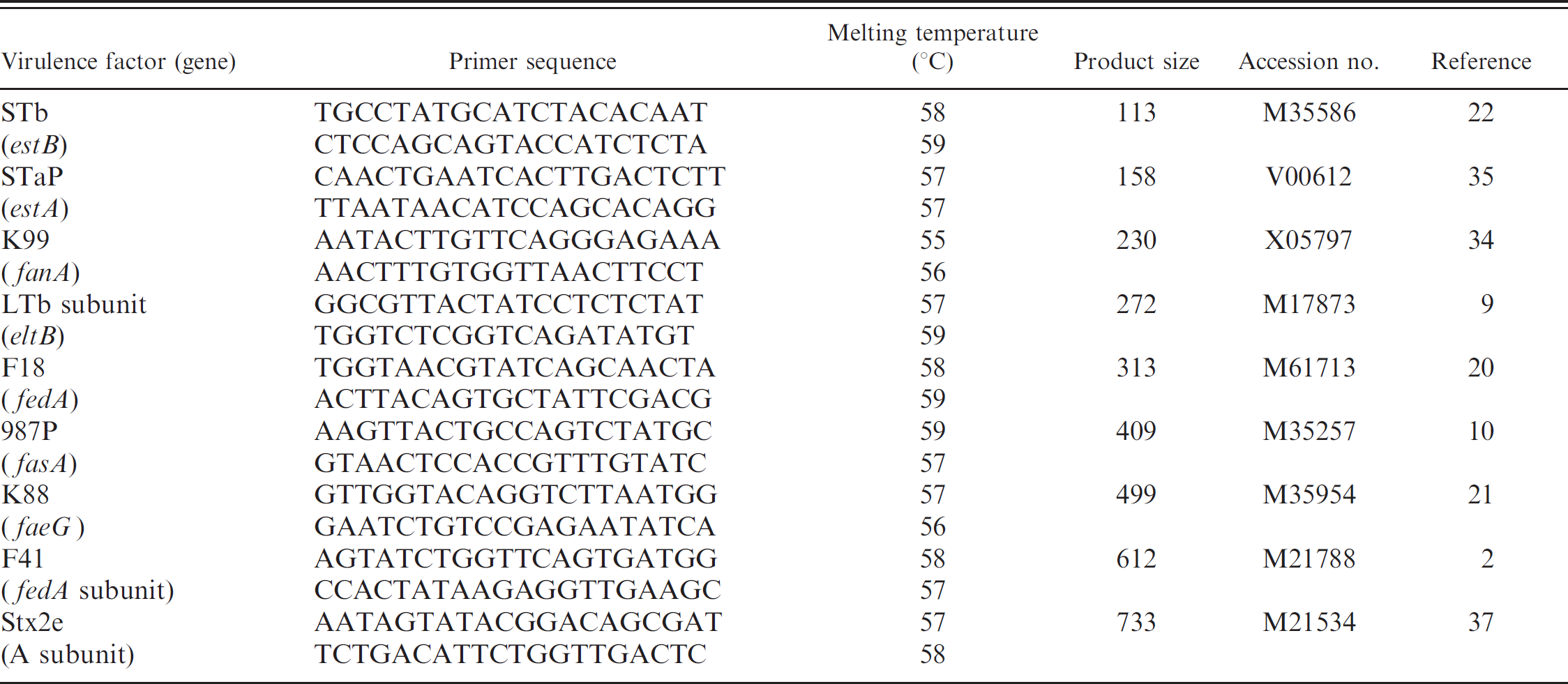
The primers were designed using Primer Designer 5 a software. The sequences of these primers are shown in Table 2, along with their corresponding GenBank accession numbers, the predicted product sizes, and other primer characteristics. The primer pairs were designed to have a similar thermal melting temperature (Tm), to have minimal base pairing interactions with other primers in the reaction, and to yield products that differed from each other by approximately 100 bp, which could be resolved by agarose gel electrophoresis. The Tm estimations for the primers shown in Table 2 were calculated based on nearest-neighbor thermodynamic parameters by the Primer Designer 5 a software. The specificity of each of the 18 primers was initially confirmed using BLASTN (nucleotide database using a nucleotide query; http://www.ncbi.nlm.nih.gov/blast/Blast.cgi) and the GenBank sequence database. 1
. DNA sequence of polymerase chain reaction primers for fimbrial and toxin genes.
The PCR reaction mixtures contained 18 primers at a concentration of 0.5 μmol each, 0.2 mmol deoxyribonucleotide triphosphate mix, b 1× AmpliTaq Gold reaction buffer II, c 5 mmol MgCl2, and 2.5 units of AmpliTaq Gold polymerase c in a final volume of 20 μl. For convenience, master mixes for 50 to 100 tubes containing all the components of the PCR reaction except the Taq polymerase and template DNA were prepared in advance. This master mix was divided into tubes and dehydrated using a Speed Vac concentrator. d Polymerase chain reaction tubes prepared this way were stored at −20°C and were stable for at least 1 year. Crude bacterial DNA template for PCR was prepared by picking a single colony into 50 μl sterile distilled H2O and heating at 95–100°C for 10 min. Template DNA (19.5 μl) was used to reconstitute a previously prepared PCR tube containing dehydrated PCR reaction mix, and 2.5 units of AmpliTaq Gold c was added. Polymerase chain reaction assays were done using a Perkin-Elmer 2400 thermocycler with the following amplification conditions: an initial denaturation at 94°C for 10 min, followed by 30 cycles of denaturation for 30 sec at 94°C, annealing at 55°C for 45 sec, and extension for 1.5 min at 72°C. The extension time was increased 3 sec each cycle, and the final extension was 10 min at 72°C.
The positive control for the mPCR assay consisted of a mixture of crude template DNA, prepared as described above, from strains 431 (K99+, F41+, and STaP+), 1477 (K88+, 987P+, LT+, STaP+, and STb+), and 2456 (F18+, LT+, STaP+, STb+, and Stx2e+). Batches of PCR tubes for positive controls that contained template DNA from strains 431, 1477, and 2456, were prepared in advance, dehydrated, and stored at −20°C as described above. Reaction products from the positive control PCR reactions also served as molecular size standards following separation by agarose gel electrophoresis. Polymerase chain reaction products were separated by electrophoresis using 4% precast agarose gels containing ethidium bromide with Tris-borate buffer (4% NuSieve 3:1 Plus Agar e ). Gels were electrophoresed at 75–100 V for 1.5–2 hr, photographed, and analyzed using an AlphaImager imaging system. f
Results
The mPCR assay for genes encoding K88, K99, F41, 987P, F18, LT, STaP, STb, and Stx2 was evaluated using 100 porcine E. coli isolates (Table 1). These strains had been examined previously in other studies for detecting porcine ETEC virulence factors by colony blot hybridization. 5,25,27 The strains were selected to include a minimum of 10 isolates that hybridized with at least 1 of the 9 virulence gene probes and 10 isolates that did not hybridize with any of these DNA probes. Additional criteria for selecting the 100 strains used to evaluate the specificity of the porcine mPCR included isolates from pigs at different ages, with different diseases, and isolates representing a diversity of pathotypes (i.e., combinations of virulence genes), including pathotypes that occur infrequently. The distribution of these characteristics in the isolates is shown in Table 1. Most of the porcine isolates contained genes for more than 1 virulence factor so that the total number of positive isolates by DNA hybridization and by mPCR for a single virulence factor gene was greater than 10 (Table 1). All of the isolates examined were positive by mPCR for the same virulence genes that were identified in these isolates by DNA hybridization using gene probes.
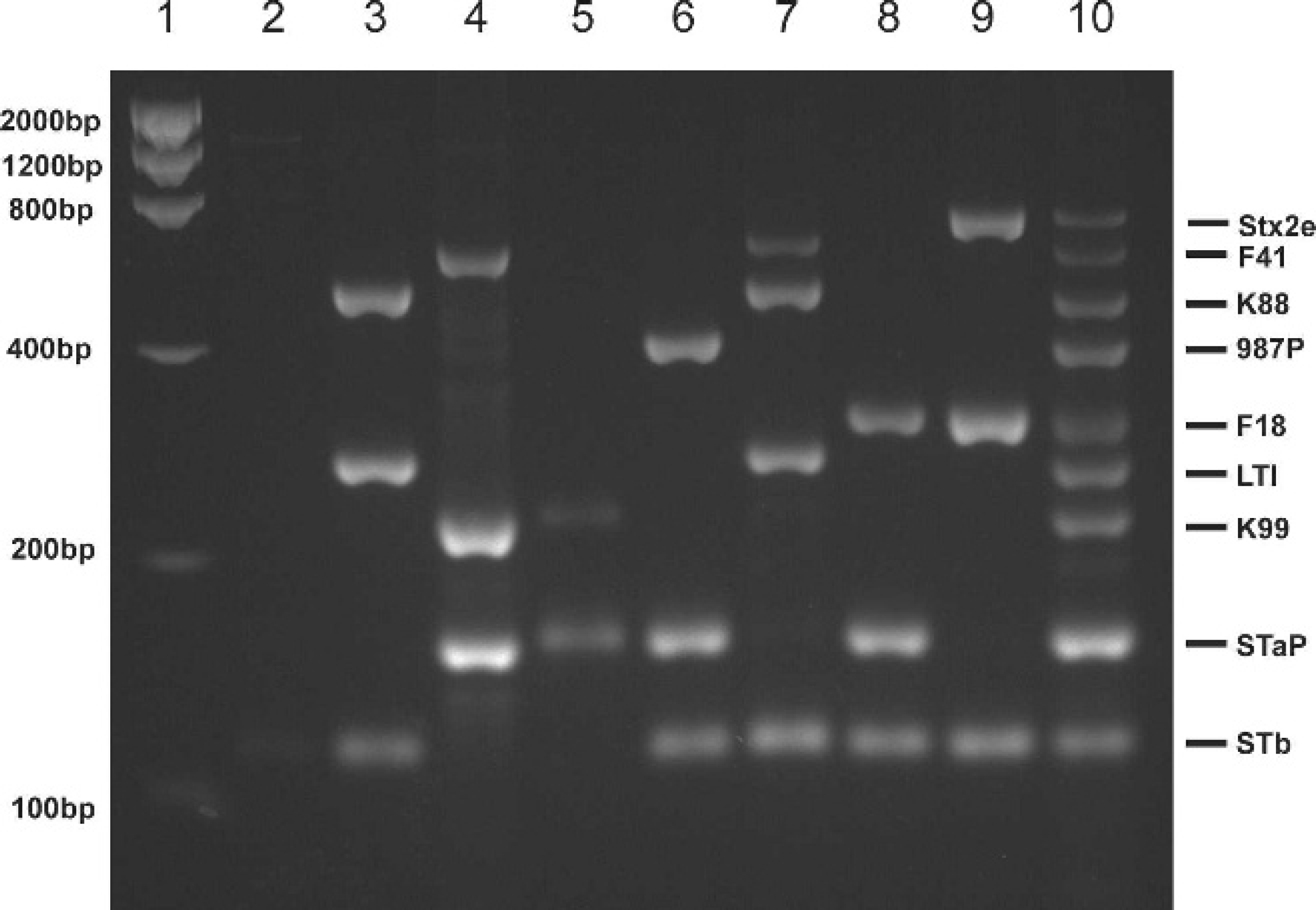
Agarose gel showing multiplex polymerase chain reaction (mPCR) assays for detection of virulence genes. The sizes of the molecular size markers in lane 1 are shown on the left and on the right, the positions of the products for each of the 9 Escherichia coli virulence genes corresponding to the fragments in the positive control mixture in lane 10. Lane 2: nonadherent, nontoxigenic strain 123; lane 3: strain 263, K88ab+, LTI+, STb+; lane 4: strain 431, F41+, K99+, STaP+; lane 5, strain 637, K99+, STaP+; lane 6: strain 987, 987P+, STaP+, STb+; lane 7: strain 2041, F41+, K88ac+, LTI+, STb+; lane 8: strain 2134, F18ac+, STaP+, STb+; lane 9: strain 2228, Stx2e+, F18ab+, STb+; and lane 10: mixture of strain 431 (K99+, F41+, STaP+), strain 1477 (K88+, 987P+, LTI+, STaP+, STb+), and strain 2456 (F18+, LTI+, STaP+, STb+, Stx2e+).
An example of the products of mPCR assays for representative isolates is shown in Figure 1. Each isolate had characteristic PCR products with sizes corresponding to the presence of specific virulence genes previously demonstrated in these isolates by DNA hybridization using gene probes. The products from a positive control mixture of DNA templates from 3 strains (431, 1477, and 2456) are shown in Figure 1, lane 10. All of the PCR products were sufficiently resolved by gel electrophoresis in 4% agarose. The use of this positive control mixture containing products for all 9 virulence genes detectable using the mPCR assay allowed identification of PCR products for each virulence gene in other isolates.
The extent of sequence variations among some of genes encoding for virulence factors targeted by the mPCR assay described here has not been examined in detail. Sequence variations have been reported for genes encoding for K88 and F18 variants. 17,33 Subtypes of LT, STa, and Stx are also known. 18,26,36 The specificity of mPCR targeting genes for K88 and F18 was evaluated using additional strains carrying K88 or F18 antigenic subtypes. Human isolates that contain genes for STaH or Stx1 were also examined by the mPCR assay reported in the present study. Sixteen strains that have been antigenically characterized for their K88 subtype were examined. The 7 K88ab-, 6 K88ac-, and 3 K88ad-positive strains and the 17 K88- positive isolates with undetermined subtypes all carried faeG gene encoding for K88 in the mPCR assay. The primers for the fedA gene encoding for F18 adhesin amplified an appropriately sized fragment in strains that were either F18ab-positive (7 strains), F18ac-positive (10 strains), or F18-positive (16 strains) isolates that have not been characterized for antigenic subtype. The mPCR assay amplified products in all 31 isolates previously determined to be STaP positive by DNA hybridizations but did not amplify a product from the strains that carried the gene for STaH in any of the 8 strains examined. The primers for the stx2e gene, which were designed to amplify a region within the A subunit gene, amplified products of correct size for all 19 isolates previously found to be positive for stx2 by colony blot hybridization and also from stx2+ O157:H7 isolates (3 strains), but not from any of the 5 stx1+ isolates examined. The primers for the gene encoding for LTI did not amplify genes in any of the 8 isolates that hybridized with LTII probes. These results showed that the mPCR assay targeting genes for K88 and F18 was not specific for individual variants and amplified the respective genes from all K88- and F18-positive isolates, regardless of antigenic subtypes. However, primers for estA gene were specific for the gene encoding for STaP subtype; the stx2 primers were specific for stx2; and the eltB primers were specific for the gene encoding LTI. The discrimination of different subtypes of these virulence factor genes may not be necessary for routine diagnostics. However, if additional subclassification is desirable, several PCR and other methods have been reported that can be used for differentiating subtypes of K88 15 and F18. 5
Discussion
The current report describes the design and evaluation of an mPCR assay for genes encoding 5 different adhesins (K88, K99, F41, 987P, and F18) and 4 different toxins (LTI, STaP, STb, and Stx2) for the detection and characterization of E. coli that cause diarrhea and ED in swine. The primers amplified genes encoding for all 3 K88 subtypes, both F18 variants, STaP subtype LTI, and Stx2. These results corroborated earlier findings using colony blot hybridization with specific DNA probes to detect the presence of the virulence genes and confirm the specificity of the PCR primers. The mPCR assay is rapid and takes very little time for sample preparation. Batch preparation and lyophilization of PCR reaction components, including PCR tubes containing DNA templates for positive control strains, further strengthen the usefulness of the assay by reducing the potential for contamination. The usefulness of the multiplex PCR assay with the 9 primer sets reported in the present study, or slightly modified primers, has been demonstrated in reported surveys on the prevalence of E. coli virulence genes in porcine isolates from Iowa, North Carolina, and South Dakota. 24,32,40
There have been numerous other reports of PCR assays for a variety of virulence factor genes in porcine ETEC and ED isolates. 4,8,11,16,30,31,38 These were designed as either single assays or in a multiplex format for various combinations of known or suspected virulence genes associated with pathogenic E. coli. As an example, a comprehensive survey of 58 virulence genes in E. coli isolates from healthy pigs and pigs with diarrhea was recently reported using 18 different uniplex or multiplex PCR assays, with up to 7 primer sets in one of the assays. 8 These authors characterized the distribution of the 58 virulence genes associated with diverse pathotypes of E. coli in human and porcine isolates. The authors found a number of virulence genes in porcine isolates that had been identified previously in human isolates associated with extraintestinal or enterohemorrhagic diseases. The specific role of these additional suspected virulence genes in porcine diarrhea and ED is not known. In addition, many of the virulence genes in the mPCR assays 8 occur infrequently in porcine pathogenic E. coli, and may therefore not be useful for the routine diagnostic characterization of porcine E. coli isolates.
The virulence genes included in the mPCR for porcine pathogenic E. coli reported in the current study were selected because they are well characterized and have been shown to be prevalent in isolates from neonatal and postweaning diarrhea and ED. This mPCR assay rapidly detected all of the genes believed to be important in ETEC and ED pathogenesis. It has been demonstrated that ETEC expression of 1 or more of the adherence factors K88, K99, F41, 987P, or F18 is necessary for adherence to small intestinal epithelial cells. 28,29 Enterotoxigenic E. coli express single or multiple enterotoxins (LTI, STa, and STb) in the small intestine, which cause fluid secretion that results in diarrhea, or ED isolates express Stx2e, which is responsible for the neurological signs of ED. 19,29 Although the role of STb in the pathogenesis of ETEC in neonatal pigs has been questioned, 6 STb is still a good marker for ETEC porcine pathogens as the most prevalent suspected virulence gene. 4,25
Reports of PCR assays for specific genes associated with pathogenic E. coli have rarely confirmed the specificity of the primers used for amplification with more than a few characterized positive control strains. 4,8,11,16,30,31,38 Additional confirmation of PCR primer specificity could include Southern or colony blot hybridization, hybridization of an internal oligonucleotide probe in real-time PCR, restriction endonuclease analysis, or nucleotide sequencing of the product. 12 Each of these methods is valuable, but a choice was made in the present study to confirm the specificity of the PCR primers by examining a large group of strains that had previously been characterized by colony blot hybridization using cloned DNA probes for the virulence genes.
The main criteria for selection of primers were similar Tm values for all 9 primer sets and differences in amplicon size of about 100 bp, which could be resolved easily by agarose gel electrophoresis. The upper limit for the number of primer sets in a single PCR reaction is unclear. A limit on the number of genes detectable in an mPCR assay may be related to the number of products that can all be easily resolved by electrophoresis on a single gel and by the increasing probability of interactions among the primers. The present study included 9 different primer pairs, but it may be possible to include additional primers for other E. coli virulence genes if desirable. Regardless, the mPCR assay reported is capable of identifying and characterizing isolates with genes for virulence factors most frequently associated with pathogenic E. coli from swine. This assay should be useful for veterinary diagnostic laboratories and for others interested in diagnosis of colibacillosis and for rapid characterization of pathogenic E. coli from swine. It may also be useful for identifying novel E. coli pathotypes that contain novel virulence genes not amplified with the mPCR using a rationale similar to that used previously with colony blot hybridization data to identify the 2134P adhesin (F18ac) on E. coli isolated from weaned pigs. 7,27
Footnotes
a.
Science and Educational Software, Cary, NC.
b.
Roche Diagnostics Corp., Indianapolis, IN.
c.
Applied Biosystems, Foster City, CA.
d.
Savant Instruments, Farmingdale, NY.
e.
Reliant Gel System, Cambrex BioScience, Rockland, ME.
f.
Alpha Innotech, San Leandro, CA.
