Abstract
To improve the diagnosis of Staphylococcus aureus mastitis and its potential antibiotic resistance in dairy cattle, a multiplex polymerase chain reaction (PCR) assay was developed for simultaneous species identification and detection of penicillin, erythromycin, and tetracycline resistance genes. The assay was first verified using 85 clinical isolates of S. aureus, and its effectiveness was evaluated by testing 99 mastitic milk samples. The multiplex PCR assay had high detection sensitivity in samples from both groups (100% agreement with single-gene PCR), with detection limits of 100–103 colony-forming units (cfu)/ml. The detection limits of the current assay for nuc, blaZ, erm B, erm C, tet K, and tet M genes were 101, 101, 102, 100, 102, and 103 cfu/ml, respectively. There was a good correlation between genotypic analysis by PCR and phenotypic determination using S. aureus species identification and susceptibility testing methods. High diagnostic sensitivities and specificities were shown for the target resistance genes (blaZ gene, 97.2% and 92.9%; ERM resistance genes, 100% and 95.6%; TET resistance genes, 100% and 96%). Results suggested that the current PCR assay could be used as an alternative diagnostic method in the routine diagnosis for rapid, sensitive, and specific detection of S. aureus and its associated antibiotic resistance genes in mastitic milk samples.
Introduction
Mastitis is an important disease in dairy herds, typically caused by bacterial infection. Among the various pathogens causing mastitis, Staphylococcus aureus is of particular importance, because it is highly infectious. 8 Early diagnosis of S. aureus mastitis provides a valuable means for preventing and controlling this disease. However, the conventional bacterial culture results may sometimes be false-negative due to prior antimicrobial treatment for the cases. 17 Several studies have suggested that polymerase chain reaction (PCR)-based assays can be applied to detect pathogens associated with mastitis and that it has several advantages, including rapid results, high sensitivity, and high throughput of milk samples.17,18,24
Despite these obvious advantages, none of the described PCR assays to date is able to detect antibiotic resistance directly from milk samples. Thus, the ability to choose effective antibiotics for treatment or to make medical projections according to antibiotic resistance patterns, even if S. aureus infection has been confirmed at an early stage, is often hindered. Currently, penicillin (PEN), erythromycin (ERY), and tetracycline (TET) are frequently used for the treatment of mastitis in China, and several reports indicate that S. aureus isolates have a high level of resistance to such antibiotics.12,25 The resistance of S. aureus to PEN is closely associated with the blaZ gene, which encodes β-lactamase.15,28 Resistance to ERY in S. aureus is predominantly mediated by a methylase encoded by the erm genes.5,11,26 Reports indicate that the genes, erm B and erm C, are present in bovine mastitis isolates in China. 25 Resistance to TET in staphylococci is conferred by 2 different mechanisms: active efflux of the drug and protection of the TET target site. Both of these mechanisms are determined by tet genes, 19 such as the tet M and tet K genes, which are the most common in conferring TET resistance in S. aureus.21,22
To detect antibiotic resistance genes of S. aureus in mastitis specimens, a multiplex PCR assay was developed that can simultaneously identify S. aureus and selected genes responsible for PEN, ERY, and TET resistance. In the current study, the applicability and effectiveness of the method was verified and evaluated through isolation of S. aureus from cases of bovine mastitis and through the use of mastitic milk samples, respectively.
Materials and methods
Bacterial strains
Staphylococcus aureus American Type Culture Collection (ATCC) 25923 and 29213 strains were chosen as control strains for species identification by PCR method. Sixteen strains belonging to other genera or species (Salmonella typhi, Bacillus subtilis, Staphylococcus haemolyticus, Staphylococcus epidermidis, Serratia marcescens, Escherichia coli, Klebsiella pneumoniae, Streptococcus agalactiae, Streptococcus dysgalactiae, Streptococcus uberis, Enterobacter cloacae, Pseudomonas aeruginosa, Proteus mirabilis, Candida albicans, Enterococcus faecalis, and Streptococcus pneumoniae) were used to determine the specificity of the primers. Eighty-five S. aureus isolates investigated in the current study were collected from bovine mastitis quarters from 2007 to 2009; all of the isolates were identified through biochemistry testing, and 53 of the isolates were additionally identified by 16S rRNA gene sequencing method. Two of the isolates were chosen according to the results of single-gene PCR as “resistance control strains” for multiplex PCR optimization and detection limit studies. Selected isolates (the strain YLD-14 contained blaZ, erm C, and tet K genes, and the strain SZHE3-1 contained erm B and tet M genes) were chosen based on their distinguishable gel bands.
DNA extraction methods
For pure culture, a DNA extraction kit a was used to extract bacterial genomic DNA with an added step for cell lysis. Cremonesi protocol 4 was applied with some modifications for bacterial DNA isolation from milk samples. In the present protocol, 1.5 ml of milk sample was used for extraction of bacterial DNA, which is 3 times that used in the Cremonesi protocol (0.5 ml). 4 Lysis buffer and binding solutions were increased accordingly to 0.6 ml and 0.5 ml, respectively.
Primer design
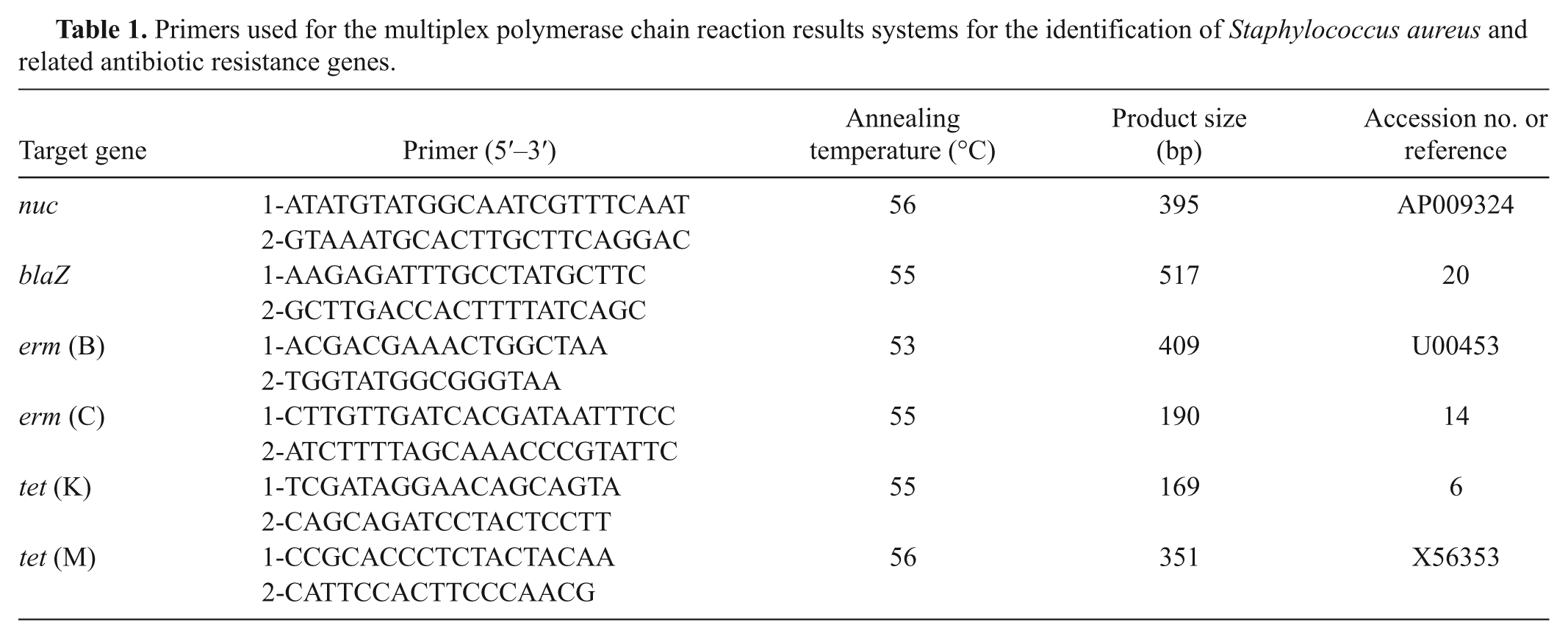
For the PCR assays, all of the primer pair sequences used in the current study, as well as their target genes, annealing temperatures, and predicted amplicon size, are listed in Table 1. For specific identification of S. aureus, the PCR primer pair was designed from the nuc gene. The primers designed for the study were based on sequences available in GenBank using Primer Premier 5.0 software. b All primers used had similar annealing temperatures, which was helpful in increasing the target PCR production.
Primers used for the multiplex polymerase chain reaction results systems for the identification of Staphylococcus aureus and related antibiotic resistance genes.
Polymerase chain reaction amplification conditions
The applicability and specificity of the nuc primers were verified using 2 S. aureus control strains and 16 strains from other species previously listed. A single-gene PCR assay was first applied to detect the nuc gene and the antibiotic resistance genes of 85 bovine mastitis isolates. Of the strains tested, strain YLD-14 (containing blaZ, erm C, and tet K genes) and strain SZHE3-1 (containing erm B and tet M genes) were chosen to optimize the multiplex PCR systems (System I and System II). Using the multiplex PCR assay, 50 isolates were used for detection of all possible resistance gene patterns.
Each single-gene PCR experiment contained 1 µl of template DNA, 1 µl of each primer pair (10 µM), 7 µl of double distilled (dd)H2O, and 10 µl of 2 × Taqmix. a Initial denaturation, which occurred at 95°C for 5 min, was followed by 30 cycles of amplification using the following parameters: 95°C for 30 sec, annealing at 55°C for 30 sec, extension at 72°C for 30 sec, and a final extension step at 72°C for 5 min. The PCR products were analyzed by electrophoresis on a 2% agarose gel. Of the 2 multiplex PCR systems, System I was designed to test for nuc, tet M, and erm C, and contained 1.5 µl of each of the nuc and tet M primer pairs, 0.5 µl of the erm C primer pair, 5 µl of template DNA, 3 µl of ddH2O and 15 µl of 2× Taqmix. a System II contained 1.5 µl of each of the erm B and blaZ primer pairs, and 0.5 µl of the tet K primer pair. Other materials used in System II were similar to those of System I. A pre-PCR step at 95°C for 5 min was applied. A total of 38 cycles were run at the following conditions: denaturation at 95°C for 30 sec, annealing at 50°C for 35 sec, and extension at 65°C for 1.5 min. After the final cycle, the preparation was kept at 65°C for 10 min to complete the reaction. The products were analyzed by electrophoresis on a 3% agarose gel.
Antimicrobial susceptibility testing
The minimal inhibitory concentrations (MICs) of PEN, ERY, and TET on S. aureus isolates were determined using broth microdilution methods, as described by the Clinical and Laboratory Standards Institute (CLSI) guidelines. 3 The S. aureus ATCC 29213 was chosen as the quality control strain.
Detection limits of multiplex polymerase chain reaction systems
The detection limits of the multiplex PCR systems were tested on milk samples for the nuc gene, and all 5 antibiotic resistance genes studied. The YLD-14 and SZHE3-1 broth cultures, at a concentration of 108 colony-forming units (cfu)/ml, were 10-fold serially diluted in sterile bovine milk. Milk samples containing 107–100 cfu/ml were used for DNA extraction. Milk samples without bacteria were used as negative control. The resultant DNA samples were subjected to PCR analysis. The detection limit of PCR for each gene was defined as the lowest concentration (cfu/ml milk) yielding a positive result.
Milk sampling and bacterial culture
The developed multiplex PCR assay for identification of S. aureus and its 5 resistance genes screened a total of 99 bovine mastitic milk samples collected from 4 herds in Beijing, China. Moreover, samples were cultured on mannitol salt agar for detection of S. aureus. Mannitol-positive isolates were subjected to Gram staining, tube coagulase test, and biochemistry test for S. aureus validation. The MICs of S. aureus isolates against PEN, ERY, and TET were determined following CLSI guidelines. 3
Sensitivity and specificity calculations
The sensitivity and specificity of the applied PCR assays were calculated for each target. Hereby, true-positive samples were defined as those positive for resistance genes and resistant to the corresponding antibiotics. False-negative samples were defined as those negative for resistance genes but resistant to the corresponding antibiotics. True-negative samples were defined as those negative for resistance genes and susceptible to the corresponding antibiotics. False-positive samples were defined as those positive for resistance genes but susceptible to the corresponding antibiotics. The following formulas were used, as previously described 13 :
Sensitivity % = [no. of true positive samples/(no. of true positive samples + no. of false negative samples)] × 100; Specificity % = [no. of true negative samples/(no. of true negative samples + no. of false positive samples)] × 100.
Additionally, the correlation between results of nuc/resistance genes detection and conventional culture and/or susceptibility test were analyzed using the SAS 9.1.3 software. c
Results
Detection of genes by single-gene and multiplex polymerase chain reaction assays
Both S. aureus control strains and all of the 85 bovine mastitis isolates were positive for the nuc gene. None of the extracted DNA from the 16 other species’ strains cross-reacted with the nuc primer pair, showing that the designed primer pair was 100% specific for identifying S. aureus isolates (data not shown). After resistance gene detection by single-gene PCR, the strains SZHE3-1 (positive for the erm C and tet M genes) and YLD-14 (positive for the blaZ, erm B, and tet K genes) were chosen to optimize the 2 multiplex PCR systems. After optimization, all target genes could be detected (Fig. 1A, 1B). Such systems were used to detect the 6 target genes from 50 out of the 85 isolates, and the results were in full agreement with those of the single-gene PCR assays, illustrating that the information obtained from the multiplex PCR systems were reliable (data not shown). Partial results are shown in Figure 1C.
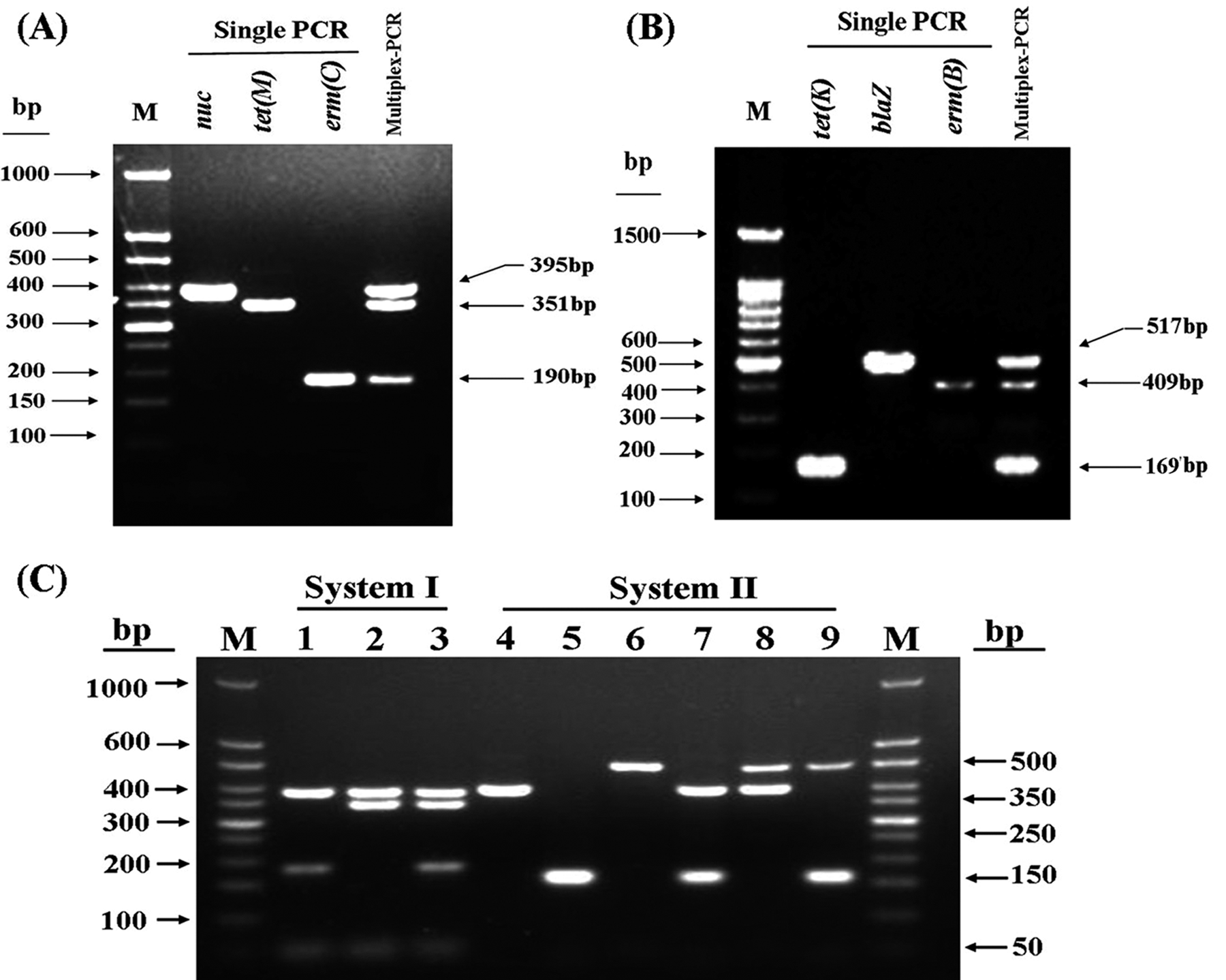
Comparison of single-target versus multiplex polymerase chain reaction (PCR) assays for detection of target genes and application of multiplex PCR for representative mastitis isolates.
Correlation between susceptibility testing and polymerase chain reaction detection
For the 85 S. aureus isolates, the antimicrobial susceptibility results and the resistance gene PCR results were compared. For these isolates, a good correlation between the results of PCR and those of classical resistance testing was found. A total of 71 isolates were resistant to PEN. In 69 of them, the blaZ gene was present. Overall, 14 isolates were susceptible to PEN, 13 of which were blaZ gene negative (Table 2). Forty isolates were resistant to ERY and carried erm B, erm C, or both resistance genes. Of the 21 isolates susceptible to ERY, 20 isolates contained no resistance genes (Table 3). In 55 TET-resistant isolates, tet K, tet M, or both genes were detected. Of the 25 TET-susceptible isolates, 24 were free of TET resistance genes (Table 4). The sensitivity of the PEN, ERM, and TET genes were 97.2% (69/71), 100% (40/40), and 100% (55/55), respectively, and the specificity of the PEN, ERM, and TET genes were 92.9% (13/14), 95.6% (20/21), and 96% (24/25), respectively. Thus, there was a good correlation between the susceptibility testing results and the PCR results.
Comparison of polymerase chain reaction (PCR) results for the blaZ gene and the penicillin minimal inhibitory concentrations (MICs) in 85 Staphylococcus aureus isolates.*
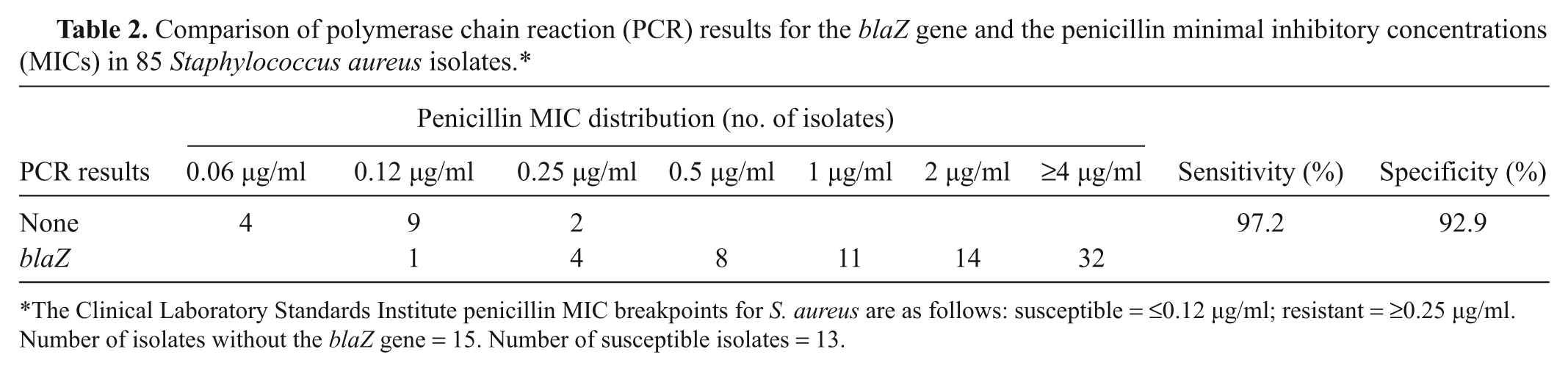

The Clinical Laboratory Standards Institute penicillin MIC breakpoints for S. aureus are as follows: susceptible = ≤0.12 µg/ml; resistant = ≥0.25 µg/ml. Number of isolates without the blaZ gene = 15. Number of susceptible isolates = 13.
Comparison of polymerase chain reaction (PCR) results for erythromycin resistance genes and the erythromycin minimal inhibitory concentrations (MICs) in 85 Staphylococcus aureus isolates.*
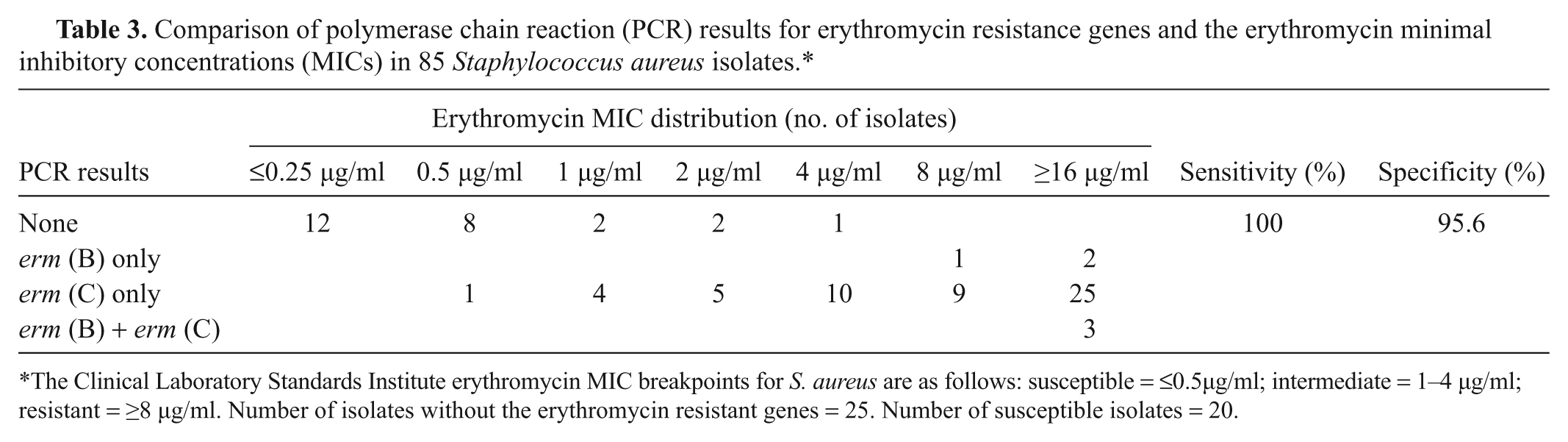

The Clinical Laboratory Standards Institute erythromycin MIC breakpoints for S. aureus are as follows: susceptible = ≤0.5µg/ml; intermediate = 1–4 µg/ml; resistant = ≥8 µg/ml. Number of isolates without the erythromycin resistant genes = 25. Number of susceptible isolates = 20.
Comparison of polymerase chain reaction (PCR) results for tetracycline resistance genes and the tetracycline minimal inhibitory concentrations (MICs) in 85 Staphylococcus aureus isolates.*
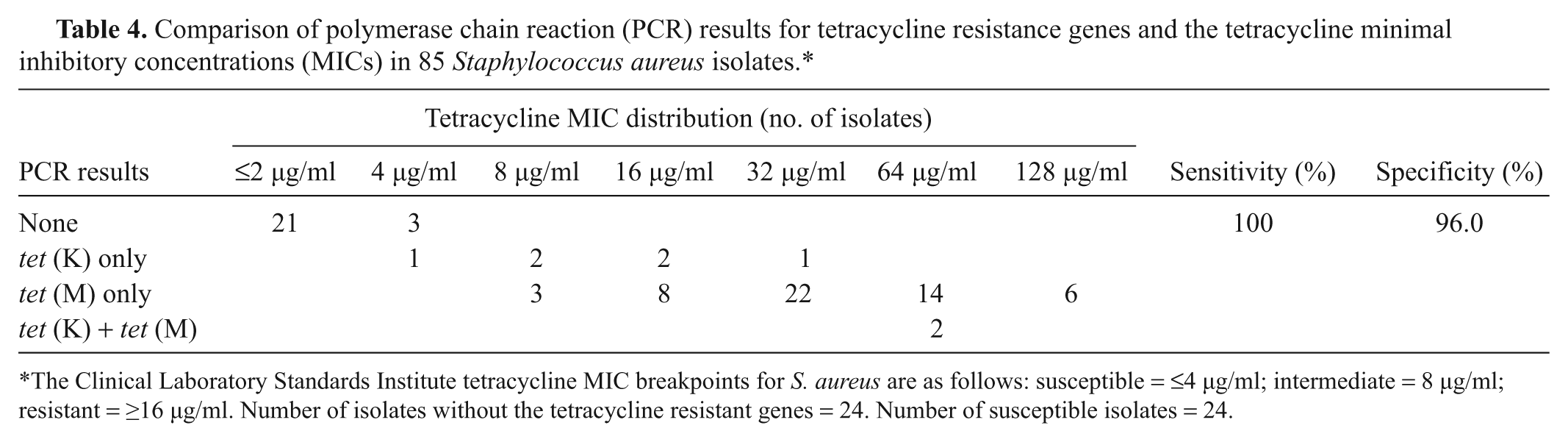

The Clinical Laboratory Standards Institute tetracycline MIC breakpoints for S. aureus are as follows: susceptible = ≤4 µg/ml; intermediate = 8 µg/ml; resistant = ≥16 µg/ml. Number of isolates without the tetracycline resistant genes = 24. Number of susceptible isolates = 24.
Detection limit of multiplex polymerase chain reaction from milk samples
For strain SZHE3-1, the detection limits of nuc, tet M, and erm C genes were 101, 103, and 100 cfu/ml, respectively, using multiplex System I (Fig. 2A). For strain YLD14, multiplex System II was used to detect the limits of blaZ, erm B, and tet K genes, which were 101, 102, and 102 cfu/ml, respectively (Fig. 2B).
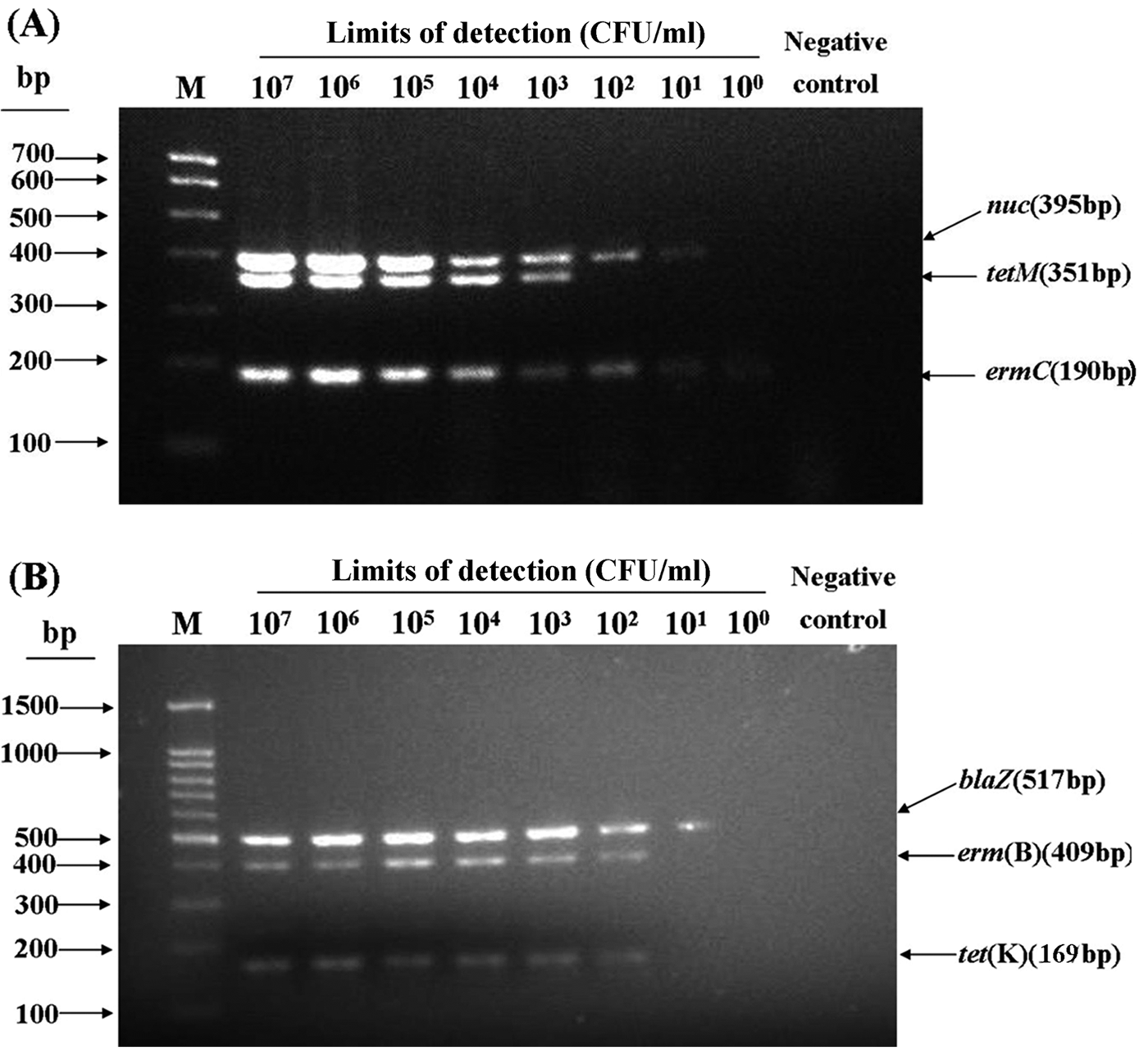
Sensitivity of multiplex polymerase chain reaction (PCR) assay in detecting DNA from milk samples artificially inoculated with “resistant control strain” SZHE3-1 and YLD-14 for nuc, blaZ, erm (B), erm (C), tet (K), and tet (M) genes. A, limits of detection for multiplex PCR System I using strain SZHE3-1. The amount of bacteria used was from 100 to 107 cfu/ml. The limits of detection for nuc, tet (M), and erm (C) genes were 101, 103, and 100 cfu/ml, respectively. Lane M: TransGen a Marker I. B, limits of detection for multiplex PCR System II using strain YLD-14. The amount of bacteria used was from 100 to 107 cfu/ml. The limits of detection for blaZ, erm (B), and tet (K) genes were 101, 102, and 102 cfu/ml, respectively. Lane M: 100-base pair (bp) DNA ladder.
Bovine mastitis sample detection
Of the 99 mastitic milk samples, 21 samples were S. aureus positive, and the remaining 78 samples were S. aureus negative, as determined by the conventional culture method (Table 5). All 21 S. aureus–positive samples were also positive for the nuc gene, and 5 of the 78 S. aureus–negative samples were positive for the nuc gene. Therefore, the sensitivity and specificity for the identification of S. aureus using mastitic milk samples in the multiplex PCR systems compared to the conventional culture method were 100% (21/21) and 93.6% (73/78), respectively. The antibiotic resistance genes detected by PCR in mastitic milk samples are shown in Table 5. Of the 21 S. aureus–positive milk samples, all were positive for the nuc gene, and for 17 samples, the detected genes were in complete agreement with the antibiotic phenotypes of the isolated S. aureus. For the remaining 4 samples, the genes were not in complete agreement with the antibiotic phenotypes: 1 sample was blaZ positive but PEN susceptible; 1 sample was erm C positive but ERY susceptible; and 2 samples were tet M positive but TET susceptible. Thus, the sensitivities and specificities of the assay for detecting genes directly from milk samples relative to isolated S. aureus were 100% (18/18) and 66.7% (2/3) for blaZ; 100% (12/12) and 85.7% (6/7) for erm B and/or erm C; and 100% (15/15) and 66.7% (4/6) for tet K and/or tet M. Following statistical analysis, the Pearson correlation coefficients between the conventional culture of S. aureus and nuc gene detection result, susceptibility test of PEN and the blaZ gene detection result, susceptibility test of ERY and the ERM resistance genes detection results, and susceptibility test of TET and TET resistance genes detection results were 0.8694, 0.7947, 0.8944, and 0.7670, respectively. Further, the correlations between all groups were highly significant (p < 0.0001). Five S. aureus–negative samples contained the nuc gene. Of these, all contained blaZ gene, 4 samples contained erm B and/or erm C genes, and 4 samples contained tet K and/or tet M genes.
Comparison of polymerase chain reaction results for the detection of nuc resistance genes and conventional culture and/or susceptibility testing among 99 mastitic milk samples.*
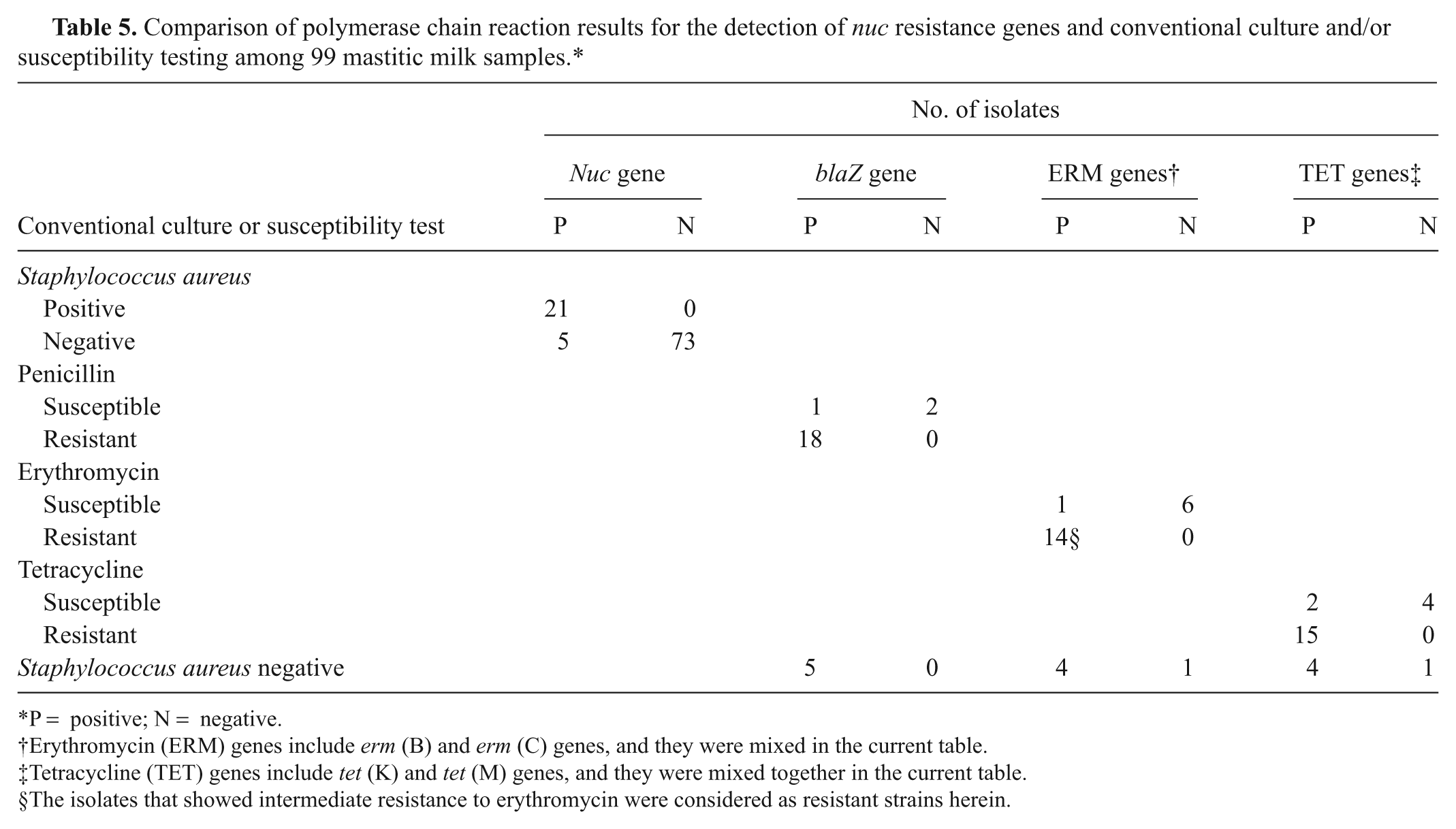
P = positive; N = negative.
Erythromycin (ERM) genes include erm (B) and erm (C) genes, and they were mixed in the current table.
Tetracycline (TET) genes include tet (K) and tet (M) genes, and they were mixed together in the current table.
The isolates that showed intermediate resistance to erythromycin were considered as resistant strains herein.
Discussion
In the present study, 2 multiplex PCR systems were designed to identify S. aureus isolates and determine their antibiotic resistance by detecting the related resistance genes. Detected isolates had a good correlation between PEN resistance phenotypes and blaZ gene detection. Such findings are in agreement with a previous study, 14 but contrary to those from another study. 7 In the present study, by comparing the results from an ERY susceptibility test and the detection of erm B and erm C, a good correlation between the presence of the 2 genes and the resistance phenotype was found, which is in agreement with the first 2 mentioned reports.14,23 The present study showed that there was also a good correlation between the TET resistance phenotype and distribution of the tet K and tet M genes. Thus, the distribution of all 5 target resistance genes, which correlated well with the phenotypes from the 3 antibiotics, is a good theoretical basis to use the developed PCR method to detect antibiotic resistance. As for S. aureus identification, the nuc gene that codes thermonuclease and is considered specific for S. aureus, 2 was chosen as the target gene. In the present study, all S. aureus isolates were nuc positive, while the non–S. aureus isolates were nuc negative, showing that this gene is available and specific.
Although a relative correlation between the 2 methods developed in the current study was shown, some discordance still exists. In the present study, the intermediate results were not considered as resistant, and they were shown separately. Among the intermediate results against ERY and TET, all isolates contained TET resistance genes and most (19/24) contained ERY resistance genes. In another study, the intermediate results were considered as resistant results, 23 which could be applicable herein. In a previous study that compared the resistant results between 2 methods of susceptibility testing, some error was detected for intermediate, susceptible, and resistant results. 16 Though some discordance may exist, the developed multiplex PCR method could be chosen as a fast, practical, and economical technique for the detection of S. aureus and related antibiotic resistance. In practice, antibiotic resistance of mastitis pathogens is only one of the key factors to influence therapy. The explanation for the apparent failure should, therefore, be searched between other factors, which could also influence the outcome of therapy.
Unlike the PCR assay performed using DNA extracted from a pure culture, many polymerase inhibitors have been found in milk samples. 27 Researchers have introduced several protocols to eliminate these PCR inhibitors.4,9,10 In the present study, to increase PCR sensitivity, the multiplex PCR systems and DNA extraction procedure from milk samples were further optimized. In the current PCR protocol, 2 dependent systems were used (System I for nuc, tet M, and erm C gene detection; System II for blaZ, erm B, and tet K gene detection), which afforded 2 advantages. First, the amplicons can be easily discriminated by electrophoresis, since extra time will be needed to separate the 2 amplicons (167 and 190 bases). Secondly, dividing the 6 primer pair reactions into 2 independent 3 primer pair reactions can increase the sensitivity of the multiplex PCR. The high identity (100%) between single and multiplex PCR showed that the optimized PCR systems have a high sensitivity. For isolating bacterial DNA from milk samples, a greater amount of bacterial DNA was isolated through the use of the modified Cremonesi protocol. 4 The current protocol used is more economical, as materials such as lysis buffer solution and binding solution, rather than protease K, were used for lysing cells.
In the current study, the detection limit for the nuc gene was 101 cfu/ml, which is similar to Cremonesi results. 4 The detection limits for the other 5 resistance genes were within the range of 100–103 cfu/ml, also showing high sensitivity of detection. This detection level eliminates the need for bacterial enrichment by culturing of S. aureus and is sensitive enough to be used as a diagnostic tool for identifying S. aureus and its 5 selected resistance genes in mastitic milk samples.
In the mastitic milk samples collected from 4 Chinese dairy herds where the nuc gene was present, there were 5 samples in which S. aureus was not successfully cultured. In clinical laboratories, a substantial proportion of the mastitic milk samples do not yield viable bacteria by conventional culture.1,24 This may be due to low concentration of bacteria in the sample, inappropriate media for some bacteria, and the existence of antibacterial factors in milk samples. 24 In the present study, the S. aureus isolates could grow easily on blood agar, so that the problem of inappropriate media could be excluded, and the limit of detection for bacterial growth in culture using 100 µl of milk is 101 cfu/ml, which is similar to that of the PCR system. The presence of antibiotics that could kill the bacteria may have been a critical factor in the study. Antibiotics were usually used immediately for the quarters suffering mastitis in the detected herds. When those mastitic samples were collected, many also contained antibiotics. Thus, bacteria in milk samples may be killed by antibiotics, which could cause the lack of bacteria growth on media. However, the target of the PCR assay is DNA, and this is not influenced by the survival of bacteria. The sensitivity of the multiplex PCR assay for mastitis samples was generally acceptable compared to conventional culture and MIC determination; however, specificity was low. This is likely due to a particularly low proportion of isolates susceptible to all 3 antibiotics. Nevertheless, the results of the field test should not be overestimated, due to a low number of cases.
The multiplex PCR method developed herein showed high sensitivity and specificity for S. aureus and could detect antibiotic resistance in both clinical S. aureus strains and mastitic milk samples. Furthermore, the results can be obtained directly from clinical samples within 4 hr, which is considerably faster than conventional culture and classic antibiotic susceptibility testing. The method is likely to be helpful for the rapid screening of antibiotic resistance in S. aureus isolates, and should allow the administration of a rapid, more focused and effective treatment and/or control of S. aureus mastitis.
Footnotes
a.
Transgen Ltd., Beijing, China.
b.
Premier Ltd., Palo Alto, CA.
c.
SAS Inc., Cary, NC.
The authors declared that they had no conflicts of interests in their authorship and publication of this contribution.
The Chinese Eleventh “Five-year” National Science and Technology Support Project (No. 2006BAD04A05) and Specialized Research Fund for the Doctoral Program of Higher Education (SRFDP) State Education Ministry (20060019045) supported the current research.
