Abstract
Typhlocolitis and dysentery due to Brachyspira hyodysenteriae infection represent an economically important disease syndrome in growing pigs. Largely disappearing from U.S. swine herds in the late 1990s and early 2000s, Brachyspira-associated disease and bacterial isolation from swine with clinical disease has increased in the last several years, and non–B. hyodysenteriae isolates are commonly identified. Antimicrobial resistance has been demonstrated in Brachyspira spp. isolates from Europe and Asia, and may be the reason for the resurgence in U.S. herds. Seventy-nine clinical isolates identified at the Iowa State University Veterinary Diagnostic Lab were tested with multiple polymerase chain reaction assays to establish species identity, and evaluated for minimum inhibitory concentrations (MICs) using an agar dilution method against lincomycin, gentamicin, valnemulin, tiamulin, salinomycin, and carbadox. Only 38.0% of isolates could be confirmed as the known pathogens B. hyodysenteriae (30.4%) or Brachyspira pilosicoli (7.6%). Twenty of the 79 isolates (25.3%) were identified as Brachyspira murdochii, and 13.9% could not be identified to species. The MIC values were consistently high against lincomycin and moderately high against gentamicin. The remaining antimicrobials had MICs that were at the low end of the test ranges. Brachyspira murdochii and Brachyspira spp. had significantly greater MIC values against several of these drugs than other Brachyspira spp. examined. The increased incidence of these less definitively characterized Brachyspira species with increased MIC values to commonly prescribed antimicrobials may, at least in part, explain the increased prevalence and severity of this disease complex in recent years. Further research is necessary to understand these changes.
Introduction
Brachyspira-associated disease, which includes swine dysentery (SD), characterized by severe mucohemorrhagic diarrhea with high morbidity caused by Brachyspira hyodysenteriae, and porcine intestinal spirochetosis (PIS), associated with mucoid, cement-gray diarrhea with depressed growth caused by Brachyspira pilosicoli, represents a severe threat to the health of growing pigs.1,4,6,7,16,18 Diagnosis can be complicated by the specialized media and growth conditions necessary to identify these species in vitro as well as the presence of other Brachyspira species described as minimally or nonpathogenic.4,12,17,21
While the incidence of Brachyspira-associated disease decreased in the 1990s, possibly due to management changes associated with confinement rearing, detection of this agent from clinically ill swine has increased in recent years. At the Iowa State University Veterinary Diagnostic Laboratory (ISU VDL; Ames, Iowa), Brachyspira isolation increased from 3 isolates in 15 cases tested in 2005 to 466 isolates in 3,465 cases during the first 9 months of 2010. Additionally, confirmatory polymerase chain reaction (PCR) designed to detect B. hyodysenteriae and B. pilosicoli in positive cultures could only identify 36.1% of these clinical isolates to the species level, indicating that the pathogenic potential of other species within this genus may have changed.
Several antimicrobials have been used in the control and prevention of Brachyspira infections, including the lincosamide lincomycin, the aminoglycoside gentamicin, the pleuromutilins tiamulin and valnemulin, the ionophore salinomycin, and the quinoxaline carbadox.8,11,16 It has been proposed that the resurgence of Brachyspira-associated disease in swine may be due to the development of antibiotic resistance to these bacterial agents. Several published reports from Europe have described a high level of resistance in Brachyspira spp. field isolates to commonly prescribed antimicrobials.1,7,14,16,18 Similar evaluations of recent Brachyspira spp. isolates in the United States are lacking. The purpose of the present study was to 1) use a combination of PCR assays to more definitively identify the Brachyspira spp. recovered from pigs with clinical disease; and 2) to perform antimicrobial agar dilution testing on these field isolates to determine the susceptibility patterns to commonly prescribed antibiotics in swine.
Materials and methods
Bacterial isolates
Seventy-nine Brachyspira spp. isolates collected from cases of pigs with clinical disease submitted to the ISU VDL between 2008 and 2010 were identified for testing. Swine farm locations included North Carolina (36), Iowa (23), Minnesota (9), Nebraska (3), Michigan (2), Illinois (2), Missouri (1), North Dakota (1), South Dakota (1), and Ohio (1). Isolates were subcultured onto trypticase soy agar a with 5% sheep blood (TSA) containing 0.008% colistin, b 0.008% vancomycin, c and 0.5% spectinomycin d (CVS agar) under anaerobic conditions using a commercial system e at 42°C and examined for growth at 48 and 96 hr. Isolated colonies identified by hemolysis were subcultured onto CVS agar 3 times to obtain a single genetic clone of Brachyspira from each case. 12 Isolates were stored at −80°C until testing was performed.
Minimum inhibitory concentration testing
Agar antimicrobial susceptibility testing was performed on each isolate against lincomycin f (1.0–64.0 µg/ml), gentamicin g (2.0–64.0 µg.ml), valnemulin h (0.125–8 µg/ml), tiamulin h (0.125–16 µg/ml), salinomycin i (0.126–16.0 µg.ml), and carbadox i (0.004–0.5 µg/ml). Stock solutions of each drug were made by mixing the antimicrobial powders with sterile distilled water (lincomycin, gentamicin, tiamulin, valnemulin), ethanol followed by sterile distilled water (carbadox), or ethanol followed by saline (salinomycin). Solutions were kept refrigerated and made fresh every 2 weeks. The MIC agar plates were prepared by adding the antimicrobial solution of appropriate concentration in 0.5-ml volume to 4.5 ml of molten TSA and poured into culture plates. 20 Plates were used within 24 hr of assembly.
Each isolate was cultured on TSA at 42°C for 48 hr to remove exposure of the bacteria to antimicrobials prior to susceptibility testing. One milliliter of agar was added to a vial containing 3 ml of sterile brain heart infusion broth and sterile glass beads. The mixture was homogenized to liberate bacteria, and 20 µl of the resultant solution containing 8.0 × 10 5 –2.0 × 10 6 cfu/ml was used to inoculate MIC plates. Plates were air dried, incubated anaerobically at 42°C, and examined for the presence of growth associated with strong or weak hemolysis at 48 and 96 hr. The MIC was defined as the lowest concentration in which no bacterial growth was detected. Additionally, each bacterial suspension was cultured for purity to ensure that no aerobic or anaerobic contaminants were present in the sample. American Type Culture Collection (ATTC) strains B. hyodysenteriae B204 (#31212) and B. pilosicoli P43/6/78 (#51139) were used as quality control organisms for agar MIC testing as previously described.5,10,18 Isolates were tested in duplicate; any isolate with MIC values greater than 2 antibiotic dilutions different from one another in replicates were tested a third time to obtain a consensus MIC value.
Polymerase chain reaction testing
The PCR testing was performed using both real-time and gel-based techniques as previously described to identify each isolate to species.19,22 The ATCC isolates B. hyodysenteriae B204 (# 31212), B. pilosicoli P43/6/78 (#51139), Brachyspira intermedia PWS/A (#51140), Brachyspira murdochii 155-20 (#700173), and Brachyspira innocens B256 (#29796) were used as quality control organisms.
Statistical analysis
The MIC results were stratified for each species of bacteria and summarized; median, mode, MIC value to which 90% of isolates tested are susceptible (MIC90), and range were determined for each Brachyspira species against each antimicrobial. Stratified analysis with the nonparametric Kruskal–Wallis test was used to compare MIC values between Brachyspira species for significant differences at P < 0.05. j
Results
An overall summary of the MIC testing performed on these isolates is shown in Figure 1. Most isolates had high MIC values against lincomycin (mode = 64 µg/ml) while 70.9% (56/79) of isolates demonstrated MIC values in the middle of the tested range against gentamicin (mode = 8 µg/ml). The MIC values were at the low end of concentrations for all isolates against the remaining tested antimicrobials: 81.0% (64/79) of isolates had MIC ≤ 0.5 µg/ml against valnemulin, 79.7% (63/79) of isolates had MIC ≤ 0.5 µg/ml against tiamulin; 98.7% (78/79) of isolates had MIC ≤ 0.5 µg/ml against salinomycin; and 97.5% (77/79) of isolates had MIC ≤ 0.03 µg/ml against carbadox.
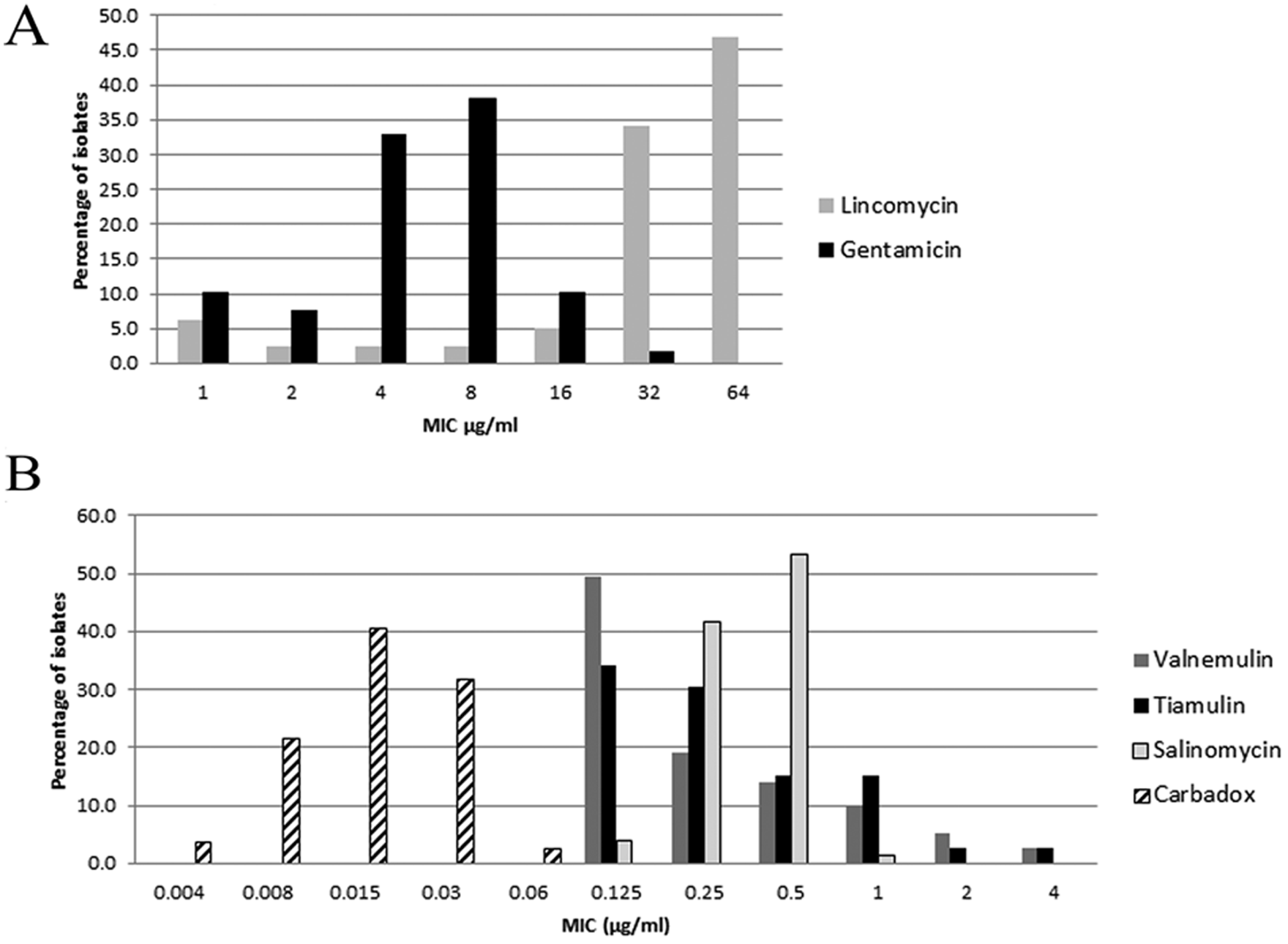
Percentage of isolates with listed minimum inhibitory concentration (MIC) values in µg/ml of (
Figure 2 shows the distribution of isolates by species identified by PCR testing. Seventy-four of the 79 (93.7%) isolates were classified as a single Brachyspira species. Only 30 of these 74 isolates (40.5%) were identified as B. hyodysenteriae or B. pilosicoli, and nearly 15% (11/74) could not be speciated as 1 of the 5 swine-associated Brachyspira species using previously published assays. Twenty-seven percent (20/74) of isolates from swine with clinical disease were identified as B. murdochii.
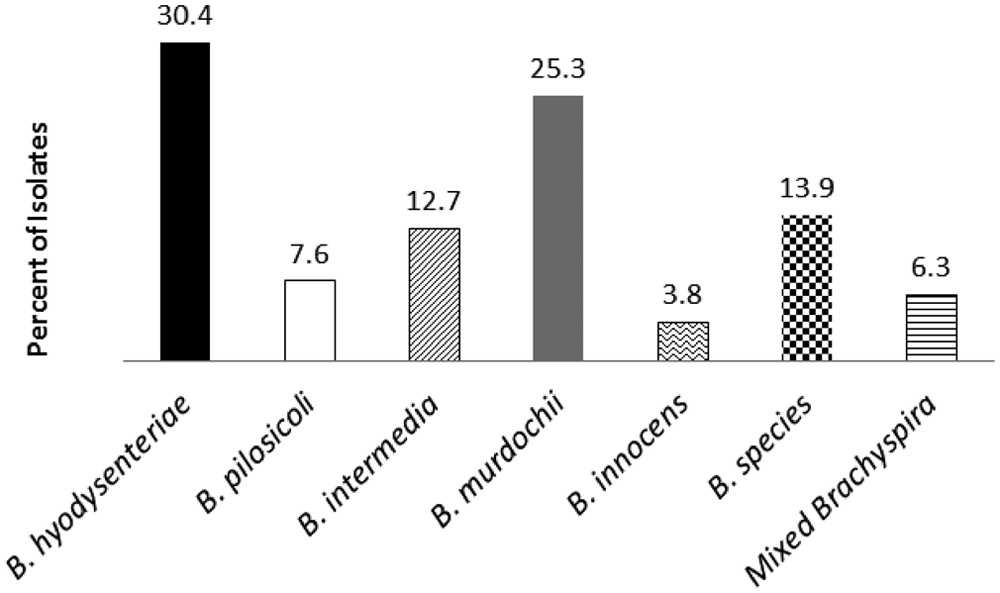
Percentages of each species of Brachyspira isolates recovered from swine with clinical disease identified by real-time and gel-based polymerase chain reaction.
The MIC values by Brachyspira species are listed in Table 1. Brachyspira murdochii had significantly greater MIC values (P < 0.05) than several other Brachyspira species against valnemulin, tiamulin, and carbadox. Brachyspira spp. had significantly greater MIC values at P < 0.05 against salinomycin and valnemulin but significantly lower MIC values against gentamicin when compared to the other Brachyspira species recovered from these clinical samples.
Median, mode, minimum inhibitory concentration value to which 90% of isolates tested are susceptible (MIC90), and range of MIC values for the listed antimicrobials by Brachyspira species.*
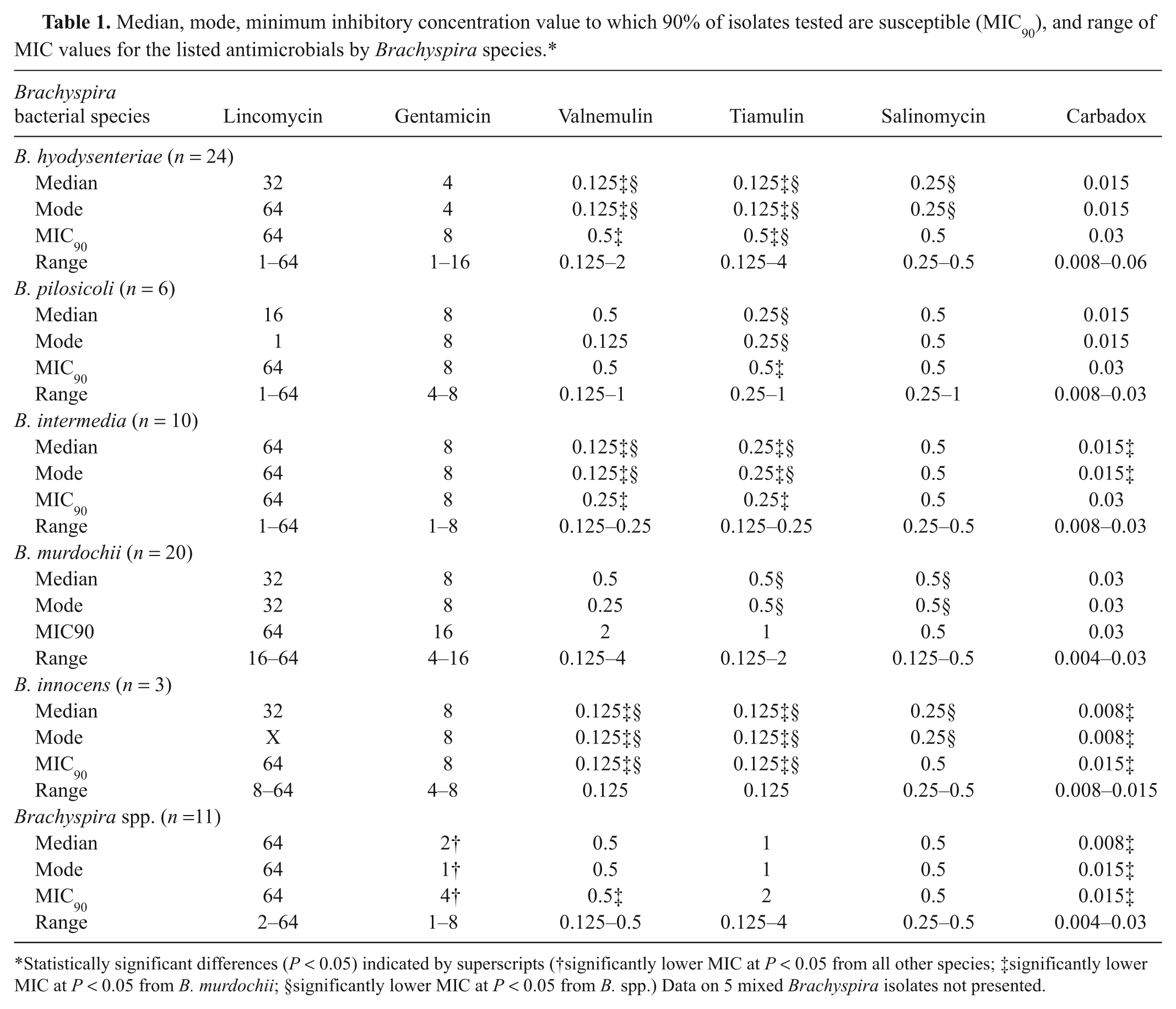
Statistically significant differences (P < 0.05) indicated by superscripts (†significantly lower MIC at P < 0.05 from all other species; ‡significantly lower MIC at P < 0.05 from B. murdochii; §significantly lower MIC at P < 0.05 from B. spp.) Data on 5 mixed Brachyspira isolates not presented.
Discussion
Even after triple cloning of individual colonies to select a single population for species identification and MIC testing, 5 of the 79 isolates (6.3%) contained a mixed Brachyspira population. This may contribute to some of the difficulties observed with species identification or it may indicate that mixed-species Brachyspira infections are more common than previously thought. The inability to categorize nearly 15% of isolates to species may indicate significant genetic alterations in this genus or possibly the emergence of a novel species.
While no Clinical Laboratory Standards Institute antimicrobial breakpoints for Brachyspira have been established, previous studies have defined guidelines for antimicrobial resistance to tiamulin and valnemulin at MIC ≥ 2 µg/ml. 11 Application of these criteria to the present study revealed that only 4.7% (6/79) and 3.2% (4/79) of isolates would be classified as “resistant” to valnemulin and tiamulin, respectively, compared with 7.4% and 17.6% in isolates from Spain. 7 MIC90 gives further information on the susceptibility of the population and can be useful in evaluating the development of resistance over time. Evaluations of isolates in Europe have identified significant increases in MIC90 of B. hyodysenteriae against tiamulin over time.7,9,18 In the Czech Republic, MIC90 values increased from 0.25 µg/ml in 1998 to 4 µg/ml in 2001, while in Germany, MIC90 increased from 0.125 µg/ml in 1989–1993 to 8 µg/ml in 2000.14,18 Based on the current study, an increased MIC against tiamulin does not appear to be widespread in U.S. isolates. A previous study 4 identified MIC90 of U.S. Brachyspira (Serpulina) isolates at 0.5 µg/ml; the MIC90 in these isolates from 2008–2010 was 1 µg/ml, which is not significantly different from that seen in 1998. Results of MIC testing using agar methods are generally 1–2 dilutions greater than those identified for broth methods. 10 The low MIC values seen in the present study, particularly against the pleuromutilin drugs, compared to those seen in studies utilizing broth methods indicate that antimicrobial resistance in U.S. isolates is not developing as rapidly as is seen in isolates from other countries.
The MIC values against the other antimicrobials tested in the current study demonstrate similar patterns to those seen elsewhere.4,11,16,18,21 The MIC values against lincomycin were consistently high. For gentamicin, values were in the mid-range of levels tested and similar to those seen in 1998 (MIC90 = 10.0 µg/ml); and for salinomycin and carbadox, values were in the low end of the range tested.
In vitro phenotypic descriptions including degree of hemolysis has been used to characterize Brachyspira, with B. hyodysenteriae showing strong beta hemolysis while B. pilosicoli, B. intermedia, B. murdochii, and B. innocens are classified as weakly hemolytic.13,15 All (10/10) B. intermedia isolates, 91% (9/11) of Brachyspira spp. isolates, and 20% (2/6) of B. pilosicoli isolates in the current study, however, demonstrated strong beta hemolysis, and 8.7% (2/23) of the B. hyodysenteriae isolates showed weak hemolysis. These altered phenotypic traits may provide evidence of changes in Brachyspira ecology and pathogenic potential.
Isolates used in the present study were limited to those recovered from pigs with clinical enteritis. Brachyspira species thought to be minimally pathogenic were frequently identified as the only Brachyspira species recovered from these cases, including B. murdochii (27% incidence) and B. intermedia (13.5% incidence). In a recent survey of swine with Brachyspira-associated disease in Japan, weakly hemolytic strains were identified as the sole Brachyspira isolate in 17 of 21 (81.0%) cases in swine with signs of dysentery, colitis, or diarrhea. 1 Similarly, in the present study, weakly hemolytic Brachyspira were identified as the sole species in 39.2% (29/74) of cases from clinical swine, and 33.8% (25/74) were species other than B. pilosicoli. Future challenge studies with these species would be needed to fulfill Koch’s postulates and prove causality; utilization of field isolates in addition to well-characterized lab strains in challenge models may give a more complete picture of the pathogenic potential of weakly hemolytic Brachyspira.
The 2009 publication of the genome of B. hyodysenteriae has provided insight into virulence mechanisms used by this pathogen. 2 At least 6 genes that encode for hemolysins, 15 genes that encode for proteases, and 4 genes that encode for DNA recombinases have been identified. Additionally, a prophage-like gene transfer agent (VSH-1) capable of moving 7,500 base-pair segments between strains of this species has been described.2,5 Similar sequences that may encode for mobile genetic elements have been identified in other Brachyspira species. 2 Changes in phenotypic behavior in vitro and the association with clinical disease in vivo may indicate that DNA transfer between Brachyspira species is also possible.
While most Brachyspira spp. isolates had low MIC values to the tested antimicrobials, significant increases in MIC values were most evident in the B. murdochii and unclassifiable Brachyspira spp. in the current study. This is of particular concern since these 2 groups may be emerging swine pathogens that could be less susceptible to antimicrobials than previously established Brachyspira pathogens. 22 Moreover, Brachyspira spp. have been associated with clinical disease in other species, including birds, dogs, and human beings; zoonotic potential could be an important consideration in Brachyspira risk assessments.3,5 Further research into virulence gene prevalence and monitoring of clinical Brachyspira spp. isolates will be useful in more definitive characterization of this disease complex.
Footnotes
Acknowledgements
The authors thank Dorothy Murphy for help with the Brachyspira isolates, and Novartis Animal Health and Phibro Animal Health for providing antimicrobial compounds.
Notes
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding was provided by Elanco Animal Health, Novartis Animal Health, Phibro Animal Health, and the National Pork Board.
