Abstract
Classical swine fever (CSF) is an economically devastating disease of pigs. Instrumental to the control of CSF is a well-characterized assay that can deliver a rapid, accurate diagnosis prior to the onset of clinical signs. A real-time fluorogenic-probe hydrolysis (TaqMan) reverse transcription polymerase chain reaction (RT-PCR) for CSF was developed by the United States Department of Agriculture (USDA) at the Plum Island Animal Disease Center (CSF PIADC assay) and evaluated for analytical and diagnostic sensitivity and specificity. A well-characterized panel including Classical swine fever virus (CSFV), Bovine viral diarrhea virus (BVDV), and Border disease virus (BDV) isolates was utilized in initial feasibility and optimization studies. The assay was initially designed and validated for use on the ABI 7900HT using the Qiagen QuantiTect® Probe RT-PCR chemistry. However, demonstrating equivalency with multiple one-step RT-PCR chemistries and PCR platforms increased the versatility of the assay. Limit of detection experiments indicated that the Qiagen QuantiTect® Multiplex (NoROX) and the Invitrogen SuperScript® III RT-PCR kits were consistently the most sensitive one-step chemistries for use with the CSF PIADC primer/probe set. Analytical sensitivity of the CSF PIADC assay ranged from <1–2.95 log10 TCID50/ml on both the ABI 7900HT and ABI 7500 platforms. The CSF PIADC assay had 100% diagnostic sensitivity and specificity when tested on a panel of 152 clinical samples from the Dominican Republic and Colombia. The ability to perform this newly developed assay in 96-well formats provides an increased level of versatility for use in CSF surveillance programs.
Keywords
Classical swine fever (CSF) is a highly contagious viral disease of domestic and wild pigs. The etiological agent is the Classical swine fever virus (CSFV), an enveloped virus belonging to the genus Pestivirus in the family Flaviviridae. Other animal pathogens within this genus include Bovine viral diarrhea virus 1 and 2 (BVDV) and Border disease virus (BDV), both of which can infect pigs. 4 Although BVDV and BDV do not cause clinical disease in swine, antibodies produced in pigs infected with these agents are cross-reactive with CSFV antigen, reducing the specificity of serological-based assays. 9
Some of the current diagnostic methods for CSF include the detection of viral antigens in tonsils using fluorescent conjugated antibody, antigen capture enzyme-linked immunosorbent assays (ELISAs), 2 or detection of genomic RNA by reverse transcription polymerase chain reaction (RT-PCR).5,6,10 The disease may present in a peracute, acute, subacute, chronic, or persistent form. Subacute and chronic forms of CSF are often associated with previously vaccinated herds or low virulence viruses and may remain clinically undetected. 9 In this regard, infection with low virulent viruses often do not present with clinical signs consistent with CSF. As CSF is classified as a notifiable disease to the World Organization for Animal Health (OIE) and the economic consequences of an outbreak are significant, rapid and accurate diagnosis or rule-out of CSF is essential. This is particularly the case, as serological and antigen tests incur a potentially high false-positive rate due to cross-reactivity occurring with BVDV and BDV.2,4
The objective of the current study was to develop a reliable, sensitive, and specific real-time reverse transcription polymerase chain reaction (real-time RT-PCR) test for CSF and validate it for fitness for use with nasal swabs from pigs. The new assay, named Classical Swine Fever Plum Island Animal Disease Center (PIADC CSF) real-time RT-PCR, was compared with the previously developed Risatti CSF real-time RT-PCR 8 assay deployed for use in the U.S. CSF Surveillance Program. Experimental observations including assay equivalency, repeatability, and agreement were evaluated and determined using statistical analyses to compare the PIADC CSF assay to the previously developed Risatti CSF assay. 8 Beyond this study is the additional validation of the PIADC CSF assay on other clinical sample types, such as whole blood from swine. 3
Viral RNA was purified from cell culture propagated virus and clinical swab samples using a commercial kit. a A total of 160 sequences spanning all major genotypic groups and subgroups of CSFV, including 105 CSFV, 28 BVDV, and 27 BDV strains, were aligned to genotype 1.1 reference CSFV Alfort 187 using BioEdit software. b This alignment was used to design a TaqMan probe, c forward and reverse primers d for the PIADC CSF assay (Table 1). All primers were synthesized and purified by column purification. d Probe sequences were synthesized and labeled with 6-carboxyfluorescein (FAM) d at the 5′ end and with minor groove-binding nonfluorescent quencher dye (MGBNFQ) e at the 3′ end. Three, one-step RT-PCR kits were tested to optimize the new assay: the QuantiTect® Probe RT-PCR Kit, f QuantiTect® Multiplex-NR RT-PCR NoROX Kit, g and SuperScript® III RT-PCR Kit. h For the PIADC CSF assay, the final reaction mixture consisted of 22.5 µl of master mix including optimized concentrations of 0.2 µM forward primer, 0.4 µM reverse primer, and 0.2 µM probe (Table 1), and 2.5 µl of viral RNA. Additionally, ROX i (2.5 µM) was added to a final concentration of 50 nM to the Multiplex-NR g and SuperScript III h master mixes for use with the PIADC CSF assay on the ABI 7500 Fast j platform. Alternately, ROX (25 µM) was added to a final concentration of 500 nM to the Multiplex-NR and SuperScript III master mixes for use with the PIADC CSF assay on the ABI 7900HT k platform.
Development, optimization, and validation of a Classical swine fever virus real-time reverse transcription polymerase chain reaction assay: primer and probe sequences designed for the detection of viral RNA.*

PIADC = Plum Island Animal Disease Center; Tm = melting temperature of nucleotide sequence.
Probes were labeled with 6-carboxyfluorescein (FAM) at the 5′ end and with minor groove-binding nonfluorescent quencher dye (MGBNFQ) at the 3′ end.
Cycling conditions on the ABI 7900HT and ABI 7500 were as follows: reverse transcription at 50°C for 30 min, enzyme activation at 95°C for 15 min, followed by 45 cycles at 94°C for 15 sec and 56°C for 60 sec. Fluorescence data for all real-time RT-PCR assays were acquired during the annealing and extension phase in the reaction using the FAM detection channel on each instrument. The results were designated positive for threshold cycle (Ct) values <40, and negative for a Ct values ≥40 or negative for samples with which threshold had not been attained before cycling completed at 45 cycles. Positive and negative amplification and extraction controls were included in each evaluation. The CSF PIADC assay was performed on the ABI 7900HT and ABI 7500 (in standard mode), and the Risatti CSF assay was performed on the SmartCycler II l and ABI 7900HT for direct comparison of the assays without differences due to the platform.
Analytical sensitivity and specificity of the CSF PIADC assay were determined using a panel of 157 CSFV, 6 BVDV, and 3 BDV isolates, and pestivirus-negative samples. The analytical panel of virus isolates was evaluated using the optimized assay conditions (above) for the CSF PIADC assay on the ABI 7900HT and ABI 7500 platforms and the Risatti CSF assay on the SmartCycler II platform (raw data not shown). Raw data from an additional smaller representative panel of 21 CSFV, 6 BDV, and 5 BVDV isolates is presented (Table 2).
Development, optimization, and validation of a Classical swine fever virus real-time reverse transcription polymerase chain reaction assay: comparison of threshold cycle values between assays and platforms used.*
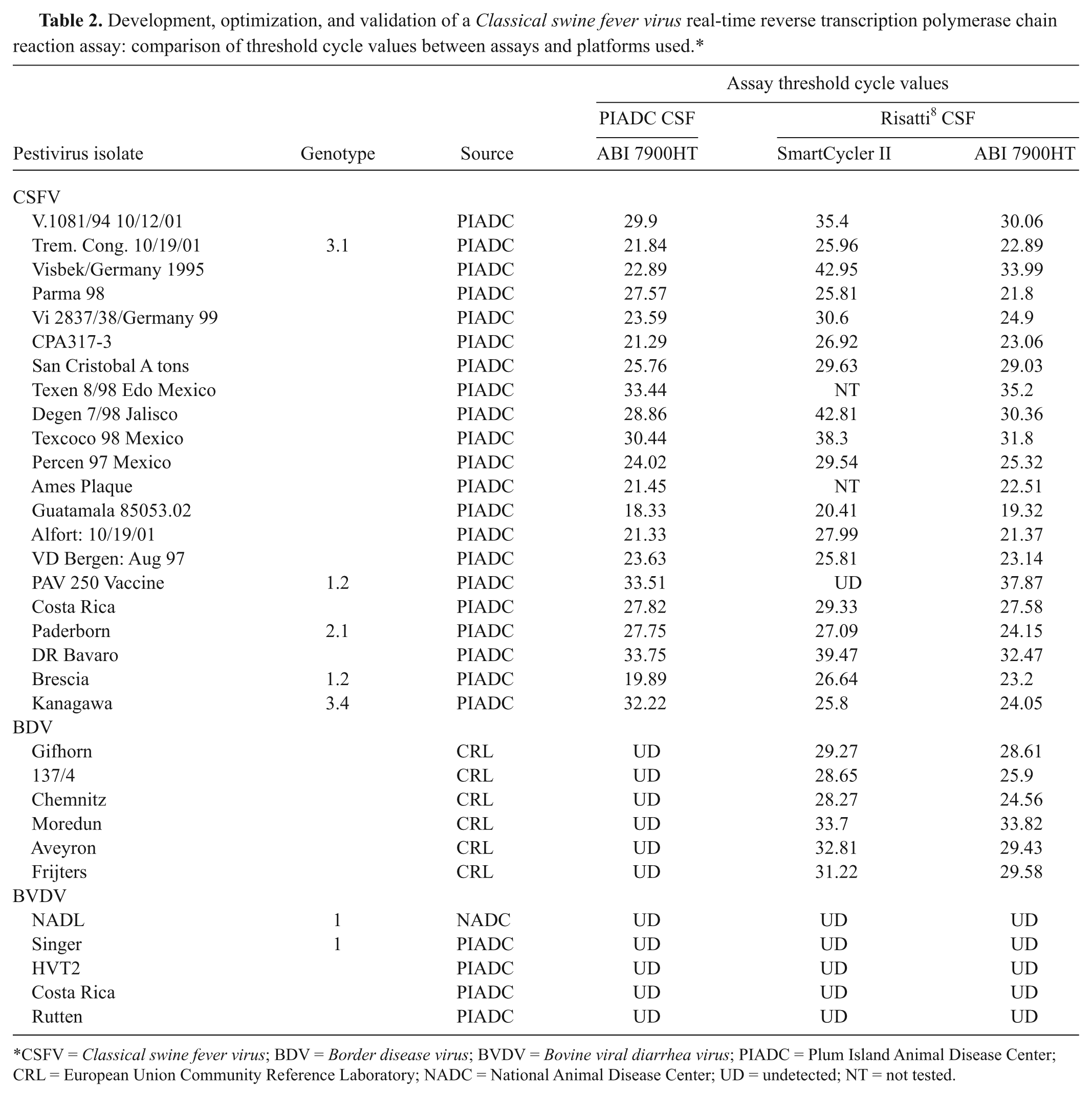
CSFV = Classical swine fever virus; BDV = Border disease virus; BVDV = Bovine viral diarrhea virus; PIADC = Plum Island Animal Disease Center; CRL = European Union Community Reference Laboratory; NADC = National Animal Disease Center; UD = undetected; NT = not tested.
For sensitivity and limit of detection (LOD) studies, infectious titers of 9 different CSF isolates were determined by calculating end-point titrations using median tissue culture infectious dose (TCID50; Table 3). Analytical performances and sensitivities of the CSF PIADC assay were determined by analyzing 10-fold dilutions of purified CSFV RNA from titered isolates of CSFV using master mixes prepared from the Probe, Multiplex-NR, and SuperScript III chemistries and amplification on the ABI 7900HT or ABI 7500 platforms (Table 3). Diagnostic sensitivity and specificity were determined by testing 58 known positive nasal swab samples from the Dominican Republic 7 and 93 known negative nasal swab samples from Colombia 7 using the optimized CSF PIADC and Risatti CSF assays.
Development, optimization, and validation of a Classical swine fever virus (CSFV) real-time reverse transcription polymerase chain reaction assay: analytical sensitivity presented as the limit of detection for diverse CSFV isolates using various assays and platforms.*
CSFV = Classical swine fever virus; PIADC = Plum Island Animal Disease Center; ND = not determined. Titer and LODs expressed as log10 TCID50/ml.
Qiagen QuantiTect® Multiplex RT-PCR NoROX Kit supplemented with ROX.
Using a large panel of 157 CSFV, 6 BVDV, and 3 BDV isolates, analytical sensitivity was found to be greater than 0.99 (95% confidence interval [CI]: 0.96, 1.00) for both the CSF PIADC and the Risatti CSF assays. Analytical specificities were 0.92 (95% CI: 0.58, 0.99) and 0.68 (95% CI: 0.40, 0.87) for the CSF PIADC and Risatti CSF assays, respectively. Both assays resulted in negative reactions for all BVDV isolates tested; however, the Risatti CSF assay consistently resulted in the positive detection of BDV. Additional testing was conducted using a somewhat smaller representative panel of 21 CSFV, 6 BDV, and 5 BVDV isolates, and pestivirus-negative samples resulted in similar sensitivities of 99% between the tests; however, the resulting specificities were 92% for the CSF PIADC assay and 68% for the Risatti CSF assay (Table 2). This indicates that the CSF PIADC assay is a more specific test and better suited as a surveillance tool because of the risks and consequences resulting from false-positive samples.
Sensitivities and LODs for the CSF PIADC and Risatti CSF assays were determined and compared using dilutions of RNA obtained from selected titered CSFV isolates. Only results from the final optimized assays comparing the CSF PIADC assay using ROX-supplemented Multiplex-NR chemistry on the ABI 7900HT and ABI 7500 platforms as compared to the Risatti CSF assay on the SmartCycler II platform are shown (Table 3).
Performance of the CSF PIADC assay during diagnostic sensitivity and specificity testing on nasal swab samples from the Dominican Republic resulted in 59 positive and 93 negative reactions, while the Risatti CSF assay resulted in the detection of 54 positive and 98 negative reactions. This indicates a greater sensitivity (100%) for the CSF PIADC assay on diagnostic samples. The agreement in results comparing the CSF PIADC and Risatti CSF assays were evaluated with Cohen kappa statistic and estimated at κ = 0.93 (95% CI: 0.87, 0.99).
To further compare the CSF PIADC and Risatti CSF assays, a total of 22 pigs were experimentally infected intranasally with 1 ml of CSFV Brescia (16 animals) or BDV (6 animals). Six uninfected pigs served as negative controls. Animals were evaluated daily for clinical signs. Following onset of clinical signs, animals were sampled by sterile nasal swabs m that were then placed into 1.5-ml sterile tubes containing 1 ml of Dulbecco minimal essential medium containing antibiotics and antimycotics. n Both assays detected CSFV equally from all 16 pigs experimentally infected with CSFV; however, the Risatti CSF assay also gave positive signals for animals experimentally infected with BDV revealing a lower specificity.
Overall, the CSF PIADC assay resulted in lower Ct values as compared to the Risatti CSF assay, regardless of amplification platform. When comparing Ct values obtained for the CSF PIADC assay on the ABI 7900HT to the Risatti CSF assay on the SmartCycler II, there did not appear to be a strong linear relationship or strong linear agreement between the 2 platforms (Table 2). The 2 assays showed stronger linear relationship and linear agreement when both amplifications were conducted using the ABI 7900HT.
The estimated LODs ranged from <1 to 3.33 using the CSF PIADC assay and from <1 to 2.16 using the Risatti CSF assay (Table 3). Amplification efficiencies were 87% for the PIADC CSF assay on the ABI 7900HT and 63% and 86% for the Risatti CSF assay run on the SmartCycler II and ABI 7900HT, respectively.
Intra-assay variability was evaluated for the CSF PIADC assay using the ABI 7900HT and the Risatti CSF assay using the SmartCycler II and ABI 7900HT. Estimated intra-assay variability for the PIADC CSF assay was 0.13 (95% CI: 0.10, 0.16) as compared with 1.00 (95% CI: 0.81, 1.30) and 0.16 (95% CI: 0.13, 0.21) for the Risatti CSF assay on the SmartCycler II and ABI 7900HT, respectively. Results suggest similar well-to-well variability between the Risatti CSF and the PIADC CSF when amplified on the ABI 7900 HT, and also indicate greater tube-to-tube variation when the Risatti CSF assay is amplified using the SmartCycler II.
With all PIADC CSF assay testing of culture isolates and clinical material, the BDV, BVDV, and known negative samples resulted in negative reactions on both platforms and with all chemistries evaluated. In tests conducted with diagnostic samples using the ABI 7900HT, resulting Ct values were lowest using the Multiplex-NR kit followed by the SuperScript III kit, and finally the Probe kit. In tests conducted with diagnostic samples using the ABI 7500, Ct values were lowest using the SuperScript III as compared to the Multiplex-NR kit.
The cross-reactivity observed with the CSF Risatti assay and BDV RNA is likely due in part to the high level of sequence conservation shared among pestivirus 5′ untranslated regions, making specific primer and probe designs challenging. 1 Indeed, in the original development and optimization of the Risatti CSF assay, 8 several BVDV (n = 6) and BDV (n = 3) isolates were tested. While no fluorogenic signal was detected using these near neighbor samples, no FAM threshold levels were indicated. Furthermore, agarose gel electrophoresis revealed BVDV- and BDV-specific PCR amplicons of expected sizes indicating the potential for cross-reactive forward and reverse primers but a lack of probe affinity for BVDV and BDV in the Risatti CSF assay. In a follow-up diagnostic evaluation of the Risatti CSF assay, 7 sensitivity and specificity were also evaluated. In this testing, sensitivity was reported as 100% while specificity was reported as 89.9%. The specificity samples were negative nasal swabs (n = 384) and did not include known pestivirus near neighbors such as BVDV or BDV to challenge the assay. 7
During a disease outbreak or in ongoing surveillance, high-throughput capabilities will be necessary to handle the high volumes of samples needing to be tested. The Risatti CSF assay was originally developed and validated only for single-tube extraction and amplification using a SmartCycler system. 8 In contrast, the CSF PIADC assay described herein was developed to function on high-throughput 96-well platforms including the ABI 7900HT and ABI 7500 systems. The greater specificity and versatility of the CSF PIADC assay, having equivalency on different chemistries and high-throughput 96-well diagnostic formats, will be a benefit to meeting the needs of a national surveillance program or diagnostic surge in the event of a CSF outbreak.
Footnotes
Acknowledgements
The authors thank the European Reference Laboratory for Classical Swine Fever (the Institute of Virology, School of Veterinary Medicine, Hannover) for supplying numerous CSFV, BDV, and BVDV isolates used in this testing. The authors also thank Dr. Luis Rodriguez (USDA, ARS, PIADC, Greenport, NY) for supplying diagnostic nasal swabs used in the current testing and Dr. Amaresh Das (USDA, APHIS, VS, NVSL, FADDL) for critical reading. Disclaimer: Mention of trade names or commercial products in the current article is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the U.S. Department of Agriculture.
a.
RNeasy® Mini Kit, Qiagen Inc., Valencia, CA.
b.
BioEdit software, Ibis Therapeutics, Carlsbad, CA.
c.
TaqMan probe, Applied Biosystems, Foster City, CA.
d.
Primer synthesis, Invitrogen Corp., Carlsbad, CA.
e.
MGBNFQ, Applied Biosystems, Foster City, CA.
f.
QuantiTect® Probe RT-PCR Kit, Qiagen Inc., Valencia, CA.
g.
QuantiTect® Multiplex RT-PCR NoROX Kit, Qiagen Inc., Valencia, CA.
h.
SuperScript® III RT-PCR Kit, Invitrogen Corp., Carlsbad, CA.
i.
ROX, Invitrogen Corp., Carlsbad, CA.
j.
ABI 7500 Fast platform, Applied Biosystems, Foster City, CA.
k.
ABI 7900HT platform, Applied Biosystems, Foster City, CA.
l.
SmartCycler® II, Cepheid Inc., Sunnyvale, CA.
m.
Dacron nasal swabs, Fisher Scientific, Pittsburg, PA.
n.
Dulbecco’s minimal essential medium, Gibco, Grand Island, NY.
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
The USDA APHIS and Department of Homeland Security Office of Science and Technology provided cooperative support for the current work.
