Abstract
Because pigs are considered intermediate hosts for the emergence of novel influenza virus reassortants with associated zoonotic potential, monitoring and characterization of circulating influenza viruses in pigs are important for adequate control of infection. For this, rapid molecular diagnostic methods other than immunoassays are needed. Three novel single-step multiplex reverse transcription polymerase chain reaction (RT-PCR) assays were developed in the current study for simultaneous detection and subtype differentiation of Influenza A virus in pigs. A conventional single-step pentaplex RT-PCR was designed for concomitant detection of the generic matrix (M) gene, hemagglutinin H1 and H3, and neuraminidase N1 and N2 genes of Swine influenza virus (SIV). In the other 2 single-step tetraplex real-time RT-PCR assays, the primers and fluorescent probes were targeted for the simultaneous detection of common M, H1, H3, and N2 SIV genes (first assay), and for M, H1, and H3 SIV genes and the H5 gene of highly pathogenic avian influenza virus of Eurasian lineage (second assay). The real-time RT-PCR assays had detection sensitivity limits ranging from 101 to 103 copies of respective in vitro RNA transcripts of M, H1, H3, H5, and N2 genes. The multiplex assays were evaluated by using SIV isolates, clinical specimens, and the appropriate synthetic template. The recent H1N1 pandemic strain isolated from pigs also was tested in simplex RT-PCR and realtime RT-PCR assays with the H1 primers and probes. The efficacy of the multiplex RT-PCR and real-time RT-PCR shows the suitability of multiplex RT-PCR and real-time RT-PCR for rapid subtype identification and monitoring in North American pigs of Influenza A virus.
Keywords
Swine influenza (SI), an acute, infectious, and highly contagious febrile respiratory disease of swine, with significant economic impact on the swine industry worldwide, 14 is caused by a single-stranded RNA virus that belongs to the genus Influenzavirus A in the family Orthomyxoviridae. Monitoring and early identification of circulating Influenza A virus (FLUAV) in pigs is required for adequate control of widespread dissemination of infection. Influenza outbreaks in pigs are mainly caused by 3 predominant subtypes: H1N1, H1N2, and H3N2. 2,27 Other sporadic subtypes, such as H3N1, 9 H4N6, H3N3, 7 and H2N3, 11 have also been reported, although their potential for transmission and persistence in pigs remains unknown. Further, sporadic occurrences of zoonotic Swine influenza virus (SIV) in humans have been reported. 13,19,23,25 The presence of less-stringent species barrier in pigs, 6 which serve as a global reservoir for FLUAV, raises important concerns for the emergence of novel and highly communicable influenza viruses with zoonotic potential. 24 Increased and continuous vigilance is required in monitoring FLUAV in pigs.
The application of rapid molecular diagnostic tests would allow detailed information on circulating FLUAV subtypes in swine and on characterizing changing genotypes with a potential for transmission across species barrier. Conventional serologic methods for subtype differentiation of FLUAV and surveillance based on hemagglutination inhibition testing and enzyme-linked immunosorbent assay have become more complex and challenging in recent years. 1,10 Increased difficulties are identified in developing adequate vaccination strategies and in implementing effective control measures by using some of these tests because of the emergence of new strains and the presence of heterogeneous anti-SIV antibody populations in field samples. 12
Conventional reverse transcription polymerase chain reaction (RT-PCR) assays have been described to identify either the hemagglutinin H1 and H3, 3,4 or the neuraminidase N1 and N2 subtypes simultaneously. 3 A multiplex RT-PCR that uses a dual priming oligonucleotide system has also been reported for screening FLUAV subtypes in Korean pigs. 8 The use of real-time RT-PCR for FLUAV subtyping rather than conventional RT-PCR could offer other additional advantages, such as short turnaround time, easy real-time monitoring, automated throughput detection, increased specificity because of the use of an additional specific probe, and more asserted confirmation of specific target amplification. To date, there is only a single report for subtype differentiation of FLUAV in pigs when using real-time RT-PCR, 18 and the report is based on individual real-time RT-PCR assays to identify the different SIV genes and is limited in terms of rapidity, convenience, cost efficiency, and broad-range detection of circulating FLUAV subtypes.
The purpose of the present study was to develop and evaluate rapid and convenient multiplex molecular diagnostic assays for simultaneous detection and subtype differentiation of circulating FLUAV in North American pigs. Consequently, 3 novel single-step multiplex RT-PCR assays were developed for screening FLUAV and concurrently identifying their subtypes. The assays consisted of a conventional single-step pentaplex RT-PCR for concomitant detection of the generic matrix (M) gene common to FLUAV and H1, H3, N1, and N2 SIV subtypes. The other 2 assays were single-step multiplex real-time RT-PCR assays in which the primers and probes in the first assay (tetraplex real-time RT-PCR-1) were targeted for the simultaneous amplification of the generic M gene and H1, H3, and N2 SIV subtypes and in the second (tetraplex real-time RT-PCR-2) for concurrent detection of M gene, H1 and H3 SIV, and H5 gene of highly pathogenic Avian influenza virus (HPAIV) of an Eurasian lineage in pigs.
In the current study, SIV standard reference strains of each representative of H1N1 (A/swine/Iowa/15/30), H3N2 (A/swine/Texas/4199−2/98), and H1N2 (A/swine/Ontario/52156/03) subtypes, 20 North American field isolates grown in eggs, and 15 lung specimens from clinically infected pigs (kindly provided by Canadian diagnostic and research laboratories: Dr. S. Carman, University of Guelph, Ontario; Dr. C. Gagnon, University of Montreal, Quebec; Dr. O. Labrecque, MAPAQ, Quebec) were used. The samples were positive for FLUAV based on simplex realtime RT-PCR performed with the matrix (M) primers and probe (Table 1). Seventeen negative lung specimens from swine were included as well. Extraction of RNA from SIV reference strains and field isolates was performed by using a viral RNA kit a and from clinical specimens by using RNeasy kit b according to the manufacturer's instructions. Oligonucleotide primers and probes for RT-PCR and realtime RT-PCR were selected from available nucleotide sequences in GenBank and by using BLAST tools (http://www.ncbi.nlm.nih.gov/) for specific amplification of the H1, H3, N1, and N2 subtypes of SIV and the H5 subtype HPAIV H5N1 virus of Eurasian origin. Primers were also selected to amplify relatively shorter PCR products for their use in multiplex assay formats, thus ensuring maximal PCR efficiency of detection.
A one-step RT-PCR kit c was used for amplification of RNA in a conventional single-step pentaplex RT-PCR. The 25-μl reaction mixture contained 4 sets of primers, each at 0.4 μM; and M-specific primers at 0.2 μM final concentration; 5 μl of 5x buffer; 1 μl of enzyme mix; and 0.3 μl of RNase inhibitor (20 U/μl). The RT was carried out at 50°C for 30 min, followed by denaturation at 95°C for 15 min and subsequent 35 cycles of amplification, each consisting of 30 sec at 94°C, 30 sec at 57°C, and 30 sec at 72°C. The RT-PCR products were examined on 3% agarose gel after electrophoresis. Among the 5 pairs of primers used in the reaction, 2 were targeted for H1 and H3, and the other 2 for N1 and N2 SIV genes (Table 2). A generic M gene-specific primer pair was also included in this multiplex reaction for detecting the M gene conserved in subtypes of FLUAV. For examining SIV isolates and clinical specimens, the reference H1N1, H3N2, and H1N2 strains were used as positive controls in the reaction. The results of RT-PCR showed bands on agarose gel that corresponded to expected sizes specific to different subtypes present (Fig. 1). They were easily discriminated by identifying a 129-bp product for H1, 211 bp for H3, 267 bp for N1, and 503 bp for N2. As expected, a common band that corresponds to 100 bp was detected from M gene amplification from different SIV subtypes (Fig. 1).
Selected primers and probes for tetraplex real-time reverse transcription polymerase chain reaction 1 and 2.

Primer and probe sequences 1, 2, 3, and 4 follow GenBank accession numbers U11857, AF153233, AF153237, and EF535027, respectively.
Selected primers and probes for pentaplex reverse transcription polymerase chain reaction (RT-PCR).

Primer sequences 1, 2, and 3 follow GenBank accession numbers U11857, AF153233, and AF250364, respectively.
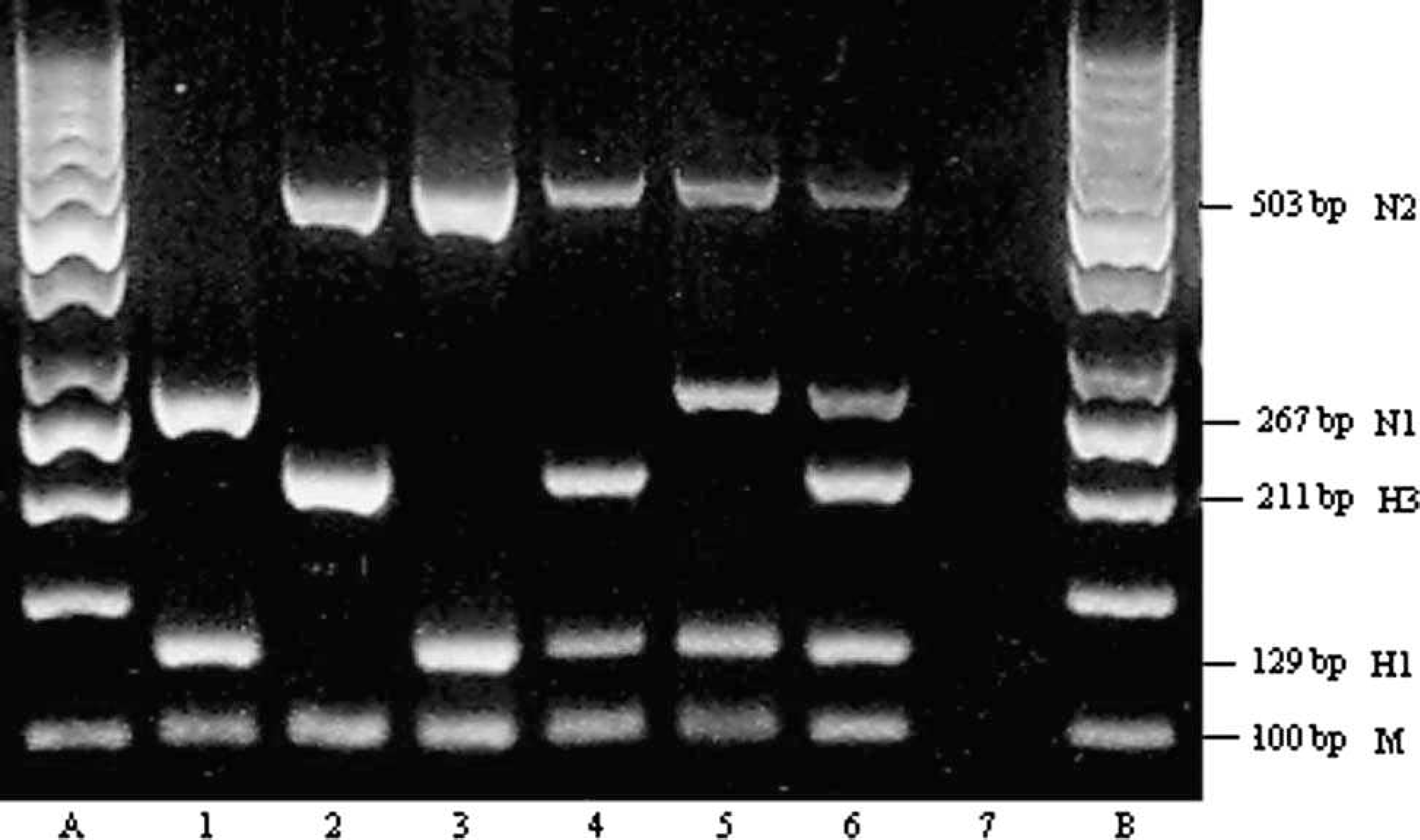
Band pattern on 3% agarose gel of the products of single-step pentaplex reverse transcription polymerase chain reaction (RT-PCR). Lanes A, B: 50-bp molecular size marker; lanes 1–3: products from Swine influenza virus reference H1N1 (A/swine/Iowa/15/30), H3N2 (A/swine/Texas/4199−2/98), and H1N2 (A/swine/Ontario/52156/03), respectively; lane 4: H3N2 and H1N2 mix; lane 5: H1N1 and H1N2 mix; lane 6: lung sample coinfected with H1N1 and H3N2, showing 5 distinct bands, corresponding to M, H1, H3, N1, and N2, respectively; lane 7: Madin-Darby canine kidney control cells.
To overcome certain limitations because of a lack of high-level physical-containment facilities for procuring and propagating the HPAIV H5N1 field sample and extracting viral RNA, an in vitro RNA transcript was prepared from H5 target template synthesized by the annealing of complementary oligonucleotides. 17 This in vitro H5 transcript, along with other H1 and H3 in vitro RNA transcripts, 21 prepared from the reference H1N1 and H3N2 SIV, were used for determining the sensitivity of single-step tetraplex real-time RT-PCR. The specificity for H5 subtype detection by real-time RT-PCR-2 was also verified by identifying the sequence homology of H5 primers and fluorescent probe used in this reaction toward other genetically diverse HPAIV H5 subtypes reported. 15,16,26
For carrying out single-step tetraplex real-time RT-PCR assays, a multiplex real-time RT-PCR kit d was used according to the manufacturer's instructions. The final 50 μl of each reaction mixture contained 4 sets of primers (Table 1), each at a final concentration of 0.2 μM, and the fluorescent probes at 0.16 μM, except the M gene-specific primers and probe, each at a final concentration of 0.12 μM and 0.03 μM final concentrations, respectively. Real-time detection was performed in thermocycler e with 30 min RT at 50°C and a 15 min denaturing/polymerase activation step at 95°C followed by 40 cycles of amplification, each consisting of 30 sec at 94°C, 30 sec at 56°C, and 20 sec at 72°C. The threshold cycle was calculated automatically by the associated thermocycler software when the specific fluorescent signal was above the background. Fluorescent values were acquired in each round during annealing and extension steps in FAM, HEX/VIC, CY5, and ROX channels.
In the tetraplex real-time RT-PCR designed for the simultaneous detection of M, H1, H3, and H5 (real-time RT-PCR-2), the reaction mixture contained the same 3 sets of primers and probes (M, H1, and H3) as were used in realtime RT-PCR-1 (Table 1). In addition, a primer pair and probe for amplifying HPAIV H5 subtype were also included (Table 1). The ROX fluorescent signal for the common M gene was detected with all the SIV reference strains and the subtypes tested, and not with negative samples. In real-time RT-PCR-1, HEX/VIC and CY5 fluorescent amplification signals for H1N2 subtype, FAM and CY5 signals for H3N2 subtype, and only HEX/VIC signal for H1N1 subtype were obtained. In real-time RT-PCR-2, HEX/VIC signal for H1 subtype, the FAM signal for H3 subtype, and CY5 signal for H5 subtype were detected. Typical amplification plots were produced across serially diluted in vitro RNA transcripts of target genes (108−101 copies/μl) in tetraplex real-time RT-PCR-1 and −2. For the target HPAIV H5, the RNA transcript prepared from synthetic template was used in real-time RT-PCR-2. Based on cumulative data from 3 independent assays, sensitivity limits that ranged from 101 to 103 copies were obtained, depending upon different selected target genes detected. The representative example of the amplification graphs obtained in single-step tetraplex realtime RT-PCR-1 and −2 are shown in Figures 2 and 3, respectively. The standard curves for these assays displayed a linear dynamic range for at least 6 orders of magnitude, with a correlation coefficient r 2 > 0.99 and overall amplification efficiencies greater than 93%.
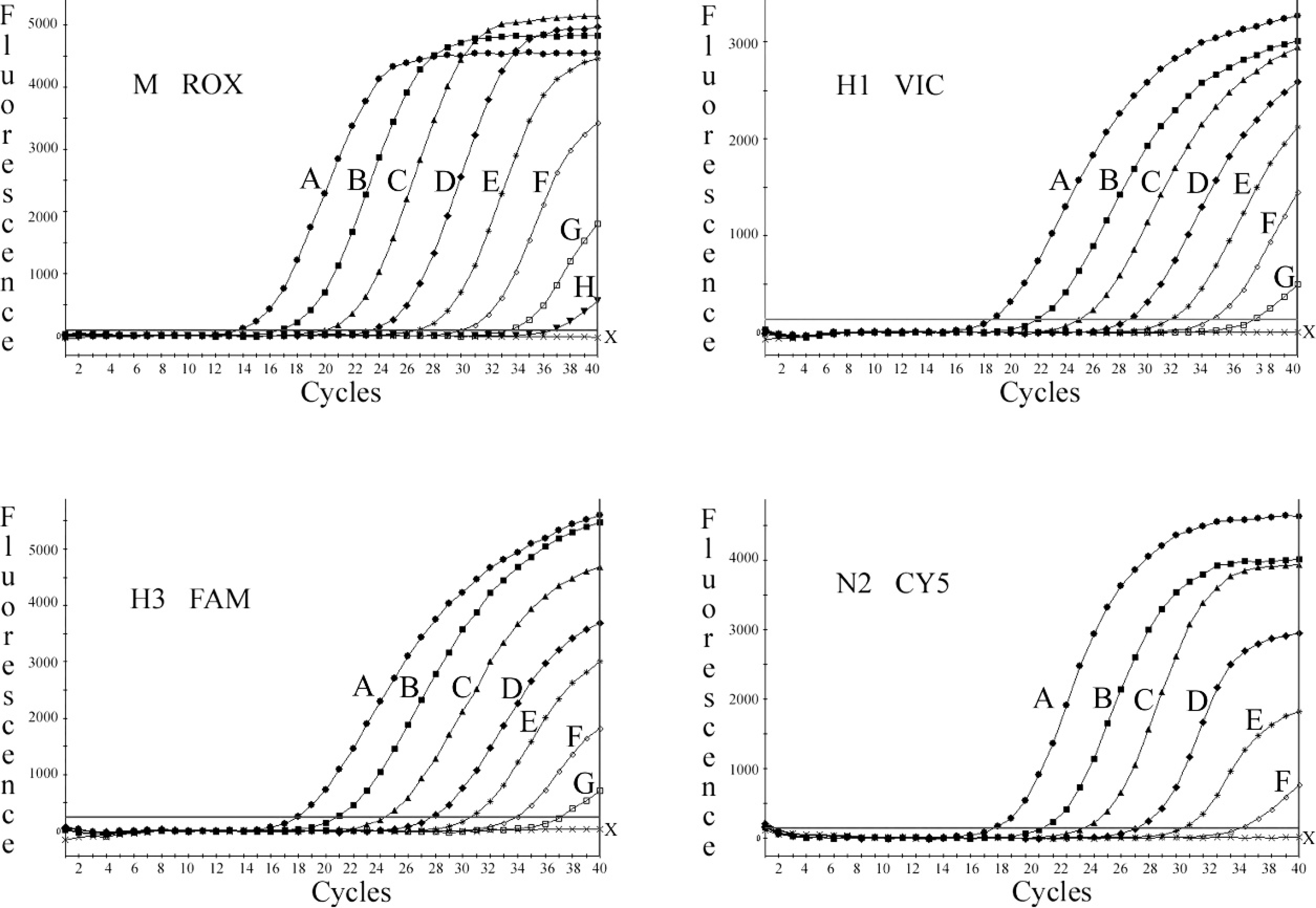
Amplification signal acquisitions in tetraplex real-time reverse transcription polymerase chain reaction-1 for the fluorophores FAM, HEX/VIC, CY5, and ROX, corresponding to respective in vitro RNA transcripts of H3, H1, N2, and M serially diluted, representing 108 (A); 107 (B); 106 (C); 105 (D); 104 (E); 103 (F); 102 (G); and 101 (H) copies/μl; negative control (X).
The single-step multiplex RT-PCR and real-time RT-PCR assays were evaluated for simultaneous detection and subtyping of FLUAV in pigs by using the North American SIV isolates and infected lung specimens. It was possible to determine the SIV subtypes of all the samples examined. Overall, with the multiplex RT-PCR, 11 field isolates and 5 lungs showed the H1N1 subtype, whereas 9 isolates and 6 lungs were of the H3N2 subtype. Interestingly, in 2 other lung samples, SIV profiles that represented both H1N1 and H3N2 were observed. A representative sample is shown in Figure 1 (lane 6). For 2 clinical lung samples, the M and H3 genes were amplified only. The subtyping results from conventional single-step multiplex RT-PCR were in concordance with that of single-step multiplex real-time RT-PCR for M, H1, H3, and N2. A single-step H1 RT-PCR (Fig. 4) performed with the primers listed in Table 2 for the target A/H1 and a simplex H1 real-time RT-PCR carried out (data not shown), with the same primers and a probe listed in Table 1 (A/H1), detected the pandemic H1N1 also while tested with a swine isolate (A/swine/Manitoba/MAFRI-17/2009; kindly provided by Dr. S. Alexandersen and Dr. J. Pasick, Canadian Food Inspection Agency, Winnipeg, Canada). The RT-PCR and real-time RT-PCR assays described in the current study were evaluated with the negative lung samples and RNA extracted from some other porcine viruses, such as Porcine reproductive and respiratory syndrome virus, Porcine respiratory coronavirus, Transmissible gastroenteritis virus, and Porcine rotavirus. No amplification was obtained for any of those tested samples.
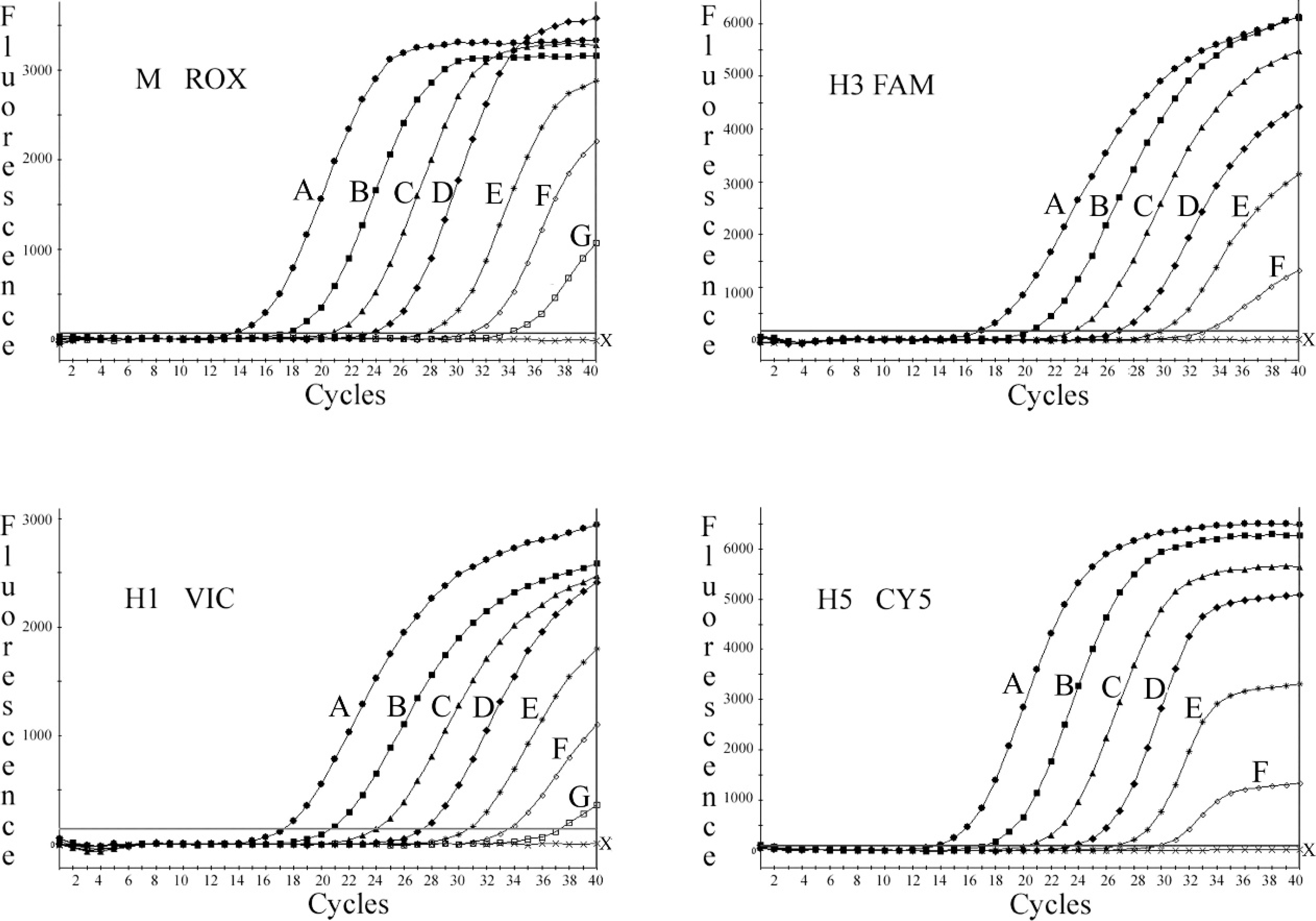
Amplification plots in tetraplex real-time reverse transcription polymerase chain reaction 2, displaying fluorescence signals of FAM, HEX/VIC, CY5, and ROX, corresponding to respective in vitro RNA transcripts of H3, H1, H5, and M serially diluted, representing 108 (A); 107 (B); 106 (C); 105 (D); 104 (E); 103 (F); and 102 (G) copies/μl; negative control (X).
The novel assays described in the current study are of potential value in supporting rapid and cost-effective molecular diagnostic efforts to track circulating subtypes of FLUAV, further genetic characterization, development of adequate vaccination strategies, and a better understanding of the emergence of novel influenza viruses with zoonotic potential. Further, the real-time RT-PCR assay designed for concurrent detection of H1 and H3, and H5, subtypes in swine offers a new strategic diagnostic approach, which could be routinely adapted for identifying not only the prevalent H1 and H3 subtypes in pigs but also the unusual HPAIV H5 subtype introduced from avian species. Increased concerns also remain about the recent outbreak of H1N1 FLUAV variant in humans and the detection of the same virus in swine on an Alberta farm (Canadian Food Inspection Agency: 2009, CFIA decodes genetic makeup of H1N1 in swine. Available at http://www.inspection.gc.ca/english/corpaffr/newcom/2009/20090515e.shtml. Retrieved on January 26, 2010).
The current study also demonstrated the feasibility of detecting in pigs the pandemic H1 subtype by the H1-specific primers-probes described in the present report. Further work is needed, however, to evaluate the different assays described on a larger number of clinical and field samples. The potential for the generation of a highly communicable novel influenza virus by reassortment among avian, human, and/or swine influenza virus gene segments with the ability to cross species appears to be greater in the recent context of a report of HPAIV H5N1 transmission to pigs. 22 Surveillance and identification of the prevalent and unusual subtypes are vital, because highly pathogenic H5N1 infection in pigs could be a silent event with no apparent clinical symptoms or mortality, 5,22 or as in the case of infection with recent H1N1 variant in pigs, an event with mild clinical symptoms and no apparent mortality (Canadian Food Inspection Agency: 2009, CFIA decodes genetic makeup of H1N1 in swine).
In conclusion, the novel single-step multiplex RT-PCR methods developed in the current study for simultaneous detection and subtype differentiation of FLUAV in North American pigs represent a significant advantage over other currently available screening methods. The present report should also allow extension of these alternative and rapid molecular diagnostic methods for surveillance studies to track a wide range of FLUAV subtypes, identify novel reassortants, and implement proper control measures.
Acknowledgements This study was supported by the Science Strategies Division of the Canadian Food Inspection Agency. The authors thank Dr. S. Carman, University of Guelph (Ontario, Canada), Dr. C. Gagnon, University of Montreal (Quebec, Canada), and Dr. O. Labrecque, MAPAQ (Quebec, Canada) for the provision of SIV reference, field isolates, and lung clinical specimens; and Dr. S. Alexandersen and Dr. J. Pasick, National Centre for Foreign Animal Diseases, CFIA (Winnipeg, Canada), for the pandemic H1N1 (A/Swine/Manitoba/MAFRI-17/2009) viral RNA. The authors also thank Peter Muller, Valerie Levesque, and Martine Antaya for technical assistance.
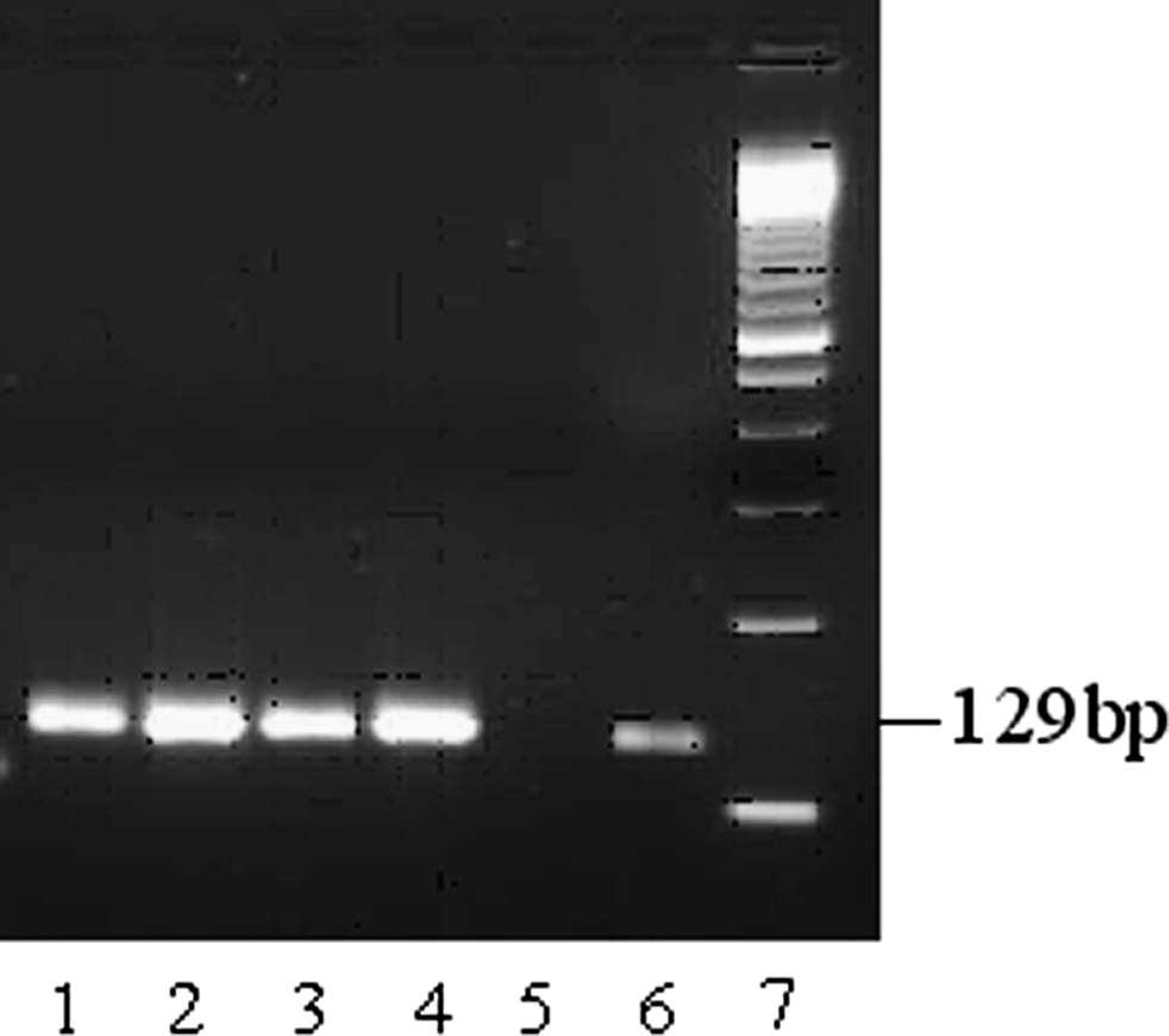
Band pattern on 3% agarose gel of the products of simplex H1 reverse transcription polymerase chain reaction. Lanes 1–4: products from pandemic Swine influenza virus H1N1 (A/swine/Manitoba/MAFRI-17/2009), H1N1 (A/swine/Iowa/15/30), H1N1 (A/swine/England/195852/92), and H1N2 (A/swine/Ontario/52156/03), respectively; lane 5: no template control; lane 6: H1 transcript-positive control; lane 7: 50-bp molecular size marker.
Footnotes
a.
QIAamp® viral RNA kit, Qiagen Inc., Mississauga, Canada.
b.
Qiagen RNeasy kit, Qiagen Inc., Mississauga, Canada.
c.
One-step RT-PCR kit, Qiagen Inc., Mississauga, Canada.
d.
QuantiTect multiplex RT-PCR NR kit, Qiagen Inc., Mississauga, Canada.
e.
Stratagene Mx4000, Stratagene Inc., La Jolla, CA.
