Abstract
To efficiently prevent and treat bovine mastitis and minimize its effect on the dairy industry, a sensitive, rapid, and specific test is required for identifying the mastitis-causing pathogens. In this study, a biochip capable of detecting 7 common species of mastitis-causing pathogens, including Corynebacterium bovis, Mycoplasma bovis, Staphylococcus aureus, and the Streptococcus spp. S. agalactiae, S. bovis, S. dysgalactiae, and S. uberis, within 6 hr was developed. The technique is based on DNA amplification of genes specific to the target pathogens and consists of 4 basic steps: DNA extraction of bacteria, polymerase chain reaction, DNA hybridization, and colorimetric reaction. To examine the accuracy and specificity of this biochip, a preliminary test with 82 random quarter milk samples were analyzed and compared with results from conventional microbiological methods conducted simultaneously. Results from all but 1 sample analyzed by the biochip were in agreement with those analyzed by bacteriology. The biochip could be a feasible tool for rapidly diagnosing mastitis-causing pathogens in milk and providing information for a more effective treatment to cure mastitis.
Introduction
Bovine mastitis is one of the most costly diseases of dairy cattle, resulting in a great deal of economic losses, mostly because of a reduction of milk yield, decreased milk quality, and higher production costs. The estimated annual loss associated with mastitis is about $184 per cow. 22 The disease is an inflammation of the mammary gland normally caused by the invasion of bacteria through the teat canal. The most common mastitis-causing pathogens are Staphylococcus aureus, Streptococcus agalactiae, coliform bacteria, and environmental Streptococcus spp. 23 Other pathogens that cause bovine mastitis, such as Corynebacterium bovis and Mycoplasma bovis, are emerging problems in dairy herds. 2,3,11,32
It has been demonstrated that early detection of mastitis can enhance curing rates and reduce the time required to recover the production of normal milk when coupled with appropriate antimicrobial therapies. 21 Identification of the pathogen is not only important to determine an effective antimicrobial treatment, but also to monitor and control the rate of infection at the farm level. Conventional microbiological methods have been the gold standard for identification of bacteria from milk. The methods, involving bacterial culture and subsequent biochemical tests on the isolate, 23 are able to identify viable bacteria and perform the test of antibiotic susceptibility. However, it is very time consuming, usually taking more than 48 hr to complete a detailed identification of pathogens. In addition, false results can be obtained when the mastitis is induced by bacteria capable of intracellular survival. 25 Other fast detecting methods, such as the California Mastitis Test and a commercially available test kit a can only detect certain pathogens, or provide limited information on the infection.
Application of DNA-based assays might circumvent some of these drawbacks associated with conventional microbiological procedures. DNA-based diagnostic assays target the unique genetic information of the bacterial genome 8,19,20 so that the specificity is higher compared with diagnostic methods based on phenotypic characterization. In addition, DNA-based assays allow rapid screening of a large number of pathogens simultaneously and provide definitive confirmation of pathogens. 8 Polymerase chain reaction (PCR) protocols have been developed for identification of various mastitis pathogens. 1,4,5,7,12,13,15,17,19,22,24–26,28 These PCR methods allow identification of bacteria within hours. After the first reports describing DNA microchip arrays appeared, microarray technology could be applied to the study of gene expression patterns in diverse organisms. 3,6 Arrays composed of oligonucleotides 24 or robotically spotted DNA 30 permit genome-scale analysis of gene expression patterns. The advantage of using microarray is that it combines powerful nucleic acid amplification strategies with its massive screening capacity, which dramatically increases the sensitivity, specificity, and efficiency. The aim of the current study was to develop a microarray with high sensitivity and specificity for rapid identification of the major pathogens involved in intra-mammary infections of cows. Therefore, a biochip capable of detecting 7 common species of mastitis-causing pathogens, including Streptococcus spp. S. bovis, S. uberis, S. agalactiae, and S. dysgalactiae, as well as C. bovis, Staph. aureus, and M. bovis, within 6 hr was developed.
The biochip is based on the DNA complementary property to capture specific target DNA. First, the genomic DNA of the pathogens is extracted from milk, and PCR technology is applied to amplify the specific target DNA. The amplicons, amplified PCR products of target DNA, are transferred to react/hybridize with specifically designed probes that are prespotted on the chip. Only the target DNA is able to hybridize with specific complementary probes, and the rest are washed away during stringent washes. Finally, subsequent colorimetric development demonstrates the pattern of pathogens presented in the sample.
Materials and methods
Bacterial strains
American Type Culture Collection (ATCC) b reference strains and Bioresource Collection and Research Center (BCRC) c reference strains were used as positive controls for the biochip assay, including Staph. aureus (ATCC 6538); Streptococcus spp. S. agalactiae (ATCC 13813), S. uberis (ATCC 19436), S. bovis (BCRC 14792), and S. dysgalactiae (ATCC 43078); C. bovis (ATCC 7715); and M. bovis (ATCC 25025). Seventy-five different ATCC reference strains, 22 BCRC reference strains, 31 U.S. Department of Agriculture strains, 20 U.S. strains, and 4 locally isolated strains d (Table 1) were used in the experiment for testing cross-reactivity.
Primers and probes
Primers and probes were designed with the use of Beacon Designer 2.1. e Primers were designed from highly divergent and species-specific regions of the DNA encoding 16S and 23S ribosomal RNA (rRNA), the gap gene, the deoxyribodipyrimidine photolyase (uvrC) gene, and the somatic cell (glyceraldehyde-3-phosphate dehydrogenase, G3p) gene on the basis of previously published sequence entries available in the GenBank database. The sequences and specificities are summarized in Table 2. The primer combinations and the lengths of the amplified products are summarized in Table 3. Primers were developed for multiplex PCR assays and tested for specificity and sensitivity previously (data not shown). All primers were resuspended to a final concentration of 200 μM in deionized autoclaved water.
Biochip preparation and assay
A tail composed of 19 T bases was added on each 5′ end of oligonucleotide probes, listed in Table 2. Ten micromolar of each probe was then spotted to each specific position on the microarray polymer substrate with an automatic spotting machine f and immobilized by a commercial ultraviolet crosslinker g with 1.2 J for 5 min. Specific probes were immobilized at known locations on a polymer substrate as the finished mastitis biochip. f The mastitis biochip contained 14 spots arranged in 5 columns as illustrated in Figure 1. In addition to the spots for specific pathogens, spots immobilized with the probes of somatic cell (C3), PCR-positive control (D5), and hybridization-positive control (A1, A5, E1, E5) were also included for comparison. The biochip assay is based on DNA amplification of genes specific to the target pathogens and consists of 4 basic steps: extraction of bacterial DNA, multiplex PCR reaction, DNA hybridization, and colorimetric reaction.
DNA isolation. Bacterial DNA was extracted directly from reference strains with the use of a commercial DNA kit h according to the manufacturer's directions. Briefly, 140 μl of bacterial broth was added to 560 μl of lysis buffer, and the mixture was incubated at room temperature for 10 min. Thereafter, 560 μl of ethanol was added to each sample, and the suspension was applied to the spin column. After centrifugation at 6,000 X g for 1 min, the column was washed with 60 μl of elution buffer. The supernatant was transferred to a new microcentrifuge tube, and 5 μl were used as the DNA template for the multiplex PCR assay.
Multiplex PCR. Amplification was made in a 50-μl reaction mixture that contained 10 mM of Tris-HCl (pH 8.3), 50 mM of KCl, 1.5 mM of MgCl2, 0.2 mM each of deoxynucleoside triphosphate (dATP, 2′-deoxyadenosine 5′-triphosphate; dCTP, 2′-deoxycytidine 5′-triphosphate; dGTP, 2′-deoxyguanosine 5′-triphosphate; dTTP, 2′-deoxythymidine 5′-triphosphate), 0.96 μM of 5′-biotinylated primer pairs (Streptococcus spp., C. bovis, and somatic cell), 0.64 μM of 5′-biotinylated primer pairs (Staph. aureus and M. bovis), and 2.5 U of Taq polymerase, i with an isolated DNA sample of 5 μl. Cycling conditions were as follows: an initial denaturation step of 95°C for 5 min; 30 cycles of 95°C for 30 sec, 60°C for 30 sec, 72°C for 30 sec; and then a final step of 72°C for 5 min. The PCR products were analyzed by electrophoresis on 2% (w/v) agarose gel to confirm their sizes.
DNA hybridization. The amplification reaction mixtures contained 5 sets of primers (Streptococcus spp., Staph. aureus, M. bovis, C. bovis, somatic cell), and the PCR products were hybridized together in the biochip. Amplicons (15 μl) were mixed with 200 μl of commercial buffer f (containing the 5′ end–biotinylated oligonucleotide complementary to the sequence of positive control probe), and the mixture was denatured in boiling water for 5 min. Thereafter, the mixture was immediately chilled on ice for 3 min and then centrifuged down. The hybridization mixture was transferred into the chip chamber and the chip was incubated at 47°C in a commercial hybridization oven f with vibration for 1 hr.
Bacterial strains evaluated for specificity by multiplex polymerase chain reaction for detection of Staphylococcus aureus; Streptococcus spp. S. agalactiae, S. uberis, S. bovis, and S. dysgalactiae; Corynebacterium bovis; and Mycoplasma.
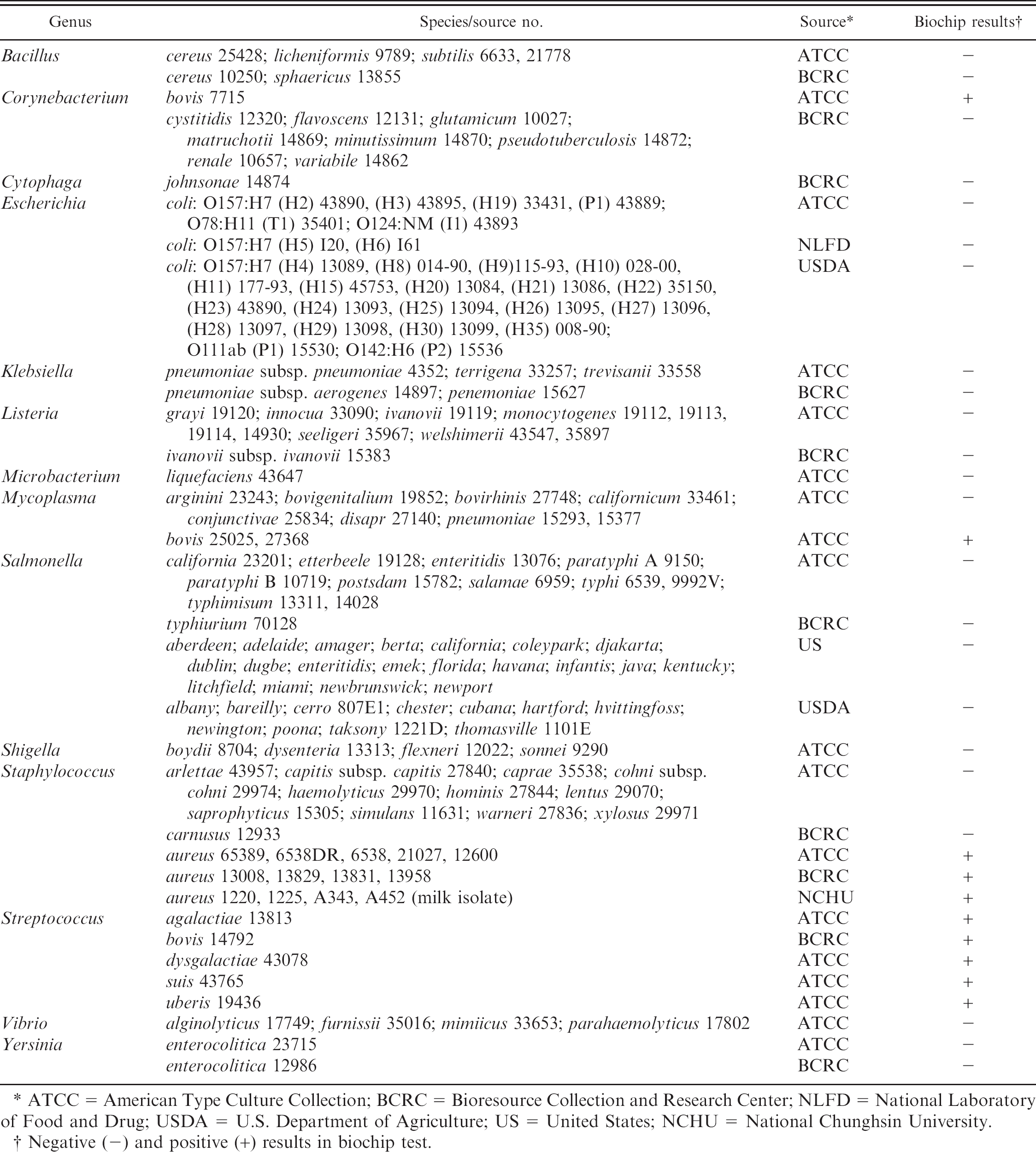
ATCC = American Type Culture Collection; BCRC = Bioresource Collection and Research Center; NLFD = National Laboratory of Food and Drug; USDA = U.S. Department of Agriculture; US = United States; NCHU = National Chunghsin University.
Negative (-) and positive (+) results in biochip test.
Colorimetric development. After hybridization, the chip was removed from the oven, and the hybridization liquid was completely discarded. Wash buffer (250 μl) was added to the hybridization chamber and discarded after 1 min. The wash was repeated 3 times. Then, the chip chamber was inverted and tapped on a paper towel to remove residual liquid. Thereafter, the mixture of 0.2-μl of Strept-AP f in 200-μl of blocking reagent was transferred into each chamber and allowed to react for 30 min at room temperature. The chamber was washed 3 times with 250-μl of wash buffer as described previously, followed by rinsing the chip with 200-μl detection buffer. After discarding the liquid, 4-μl of NBT/BCIP (nitro blue tetrazolium/5-bromo-4-chloro-3-indolyl phosphate) was mixed with 196 μl of detection buffer, and the mixture was added to the hybridization chamber. The colorimetric development was allowed to react in the dark for 10 min at room temperature, then the detection liquid was discarded. The chip was rinsed with water twice, and the pattern that developed on the chip was recorded with the naked eye.
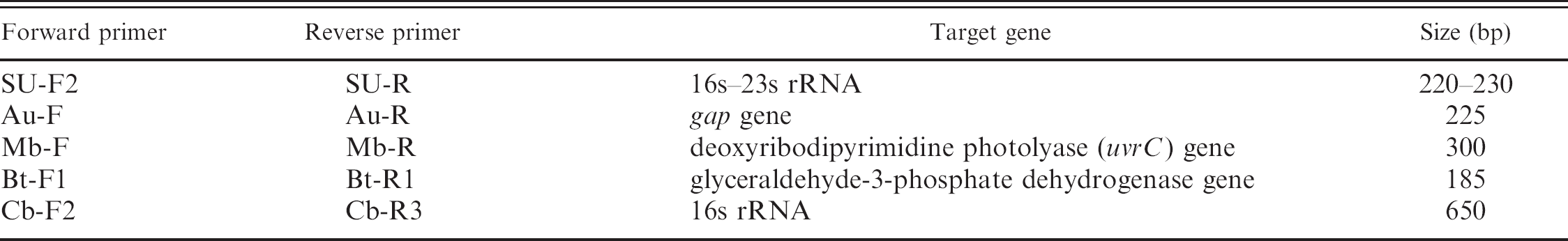
Primers and probes used in the biochip.

Length in number of nucleotides.
Establishing the detection limit of the biochip assay with pure bacterial cultures
Seven strains, including Staph. aureus (ATCC 6538); Streptococcus spp. S. agalactiae (ATCC 13813), S. uberis (ATCC 19436), S. bovis (BCRC 14792), and S. dysgalactiae (ATCC 43078); C. bovis (ATCC 7715); and M. bovis (ATCC 25025), were cultured overnight and numerated. The bacterial suspension of each strain was serial diluted (10-fold), from 107 to 102 CFU/ml, in ultra–heat-treated (UHT) milk purchased from a local grocery store in Taiwan. Bacterial DNA was extracted directly from the diluted suspensions with the use of a commercial DNA kit, h as described, and used as the template for determining the detection limit (sensitivity) of the biochip assay.
Comparison of conventional culture method and biochip assay for the detection of bacteria in milk samples
Sampling and collection of milk samples. Quarter milk samples (n = 82) were randomly collected from dairy farms participating in the Taiwan Dairy Herd Improvement program. These samples were collected before milking according to standard procedures described by the National Mastitis Council. 22 Before sample collection, teats of cows were dipped in premilking teat disinfectant, cleaned thoroughly, and dried with individual disposable paper towels; then, teat ends were sanitized with swabs containing 70% isopropyl alcohol. Milk samples were transported on ice and maintained at −20°C until they were analyzed.
Primers used for polymerase chain reaction (PCR) amplification and the PCR-amplified product sizes.
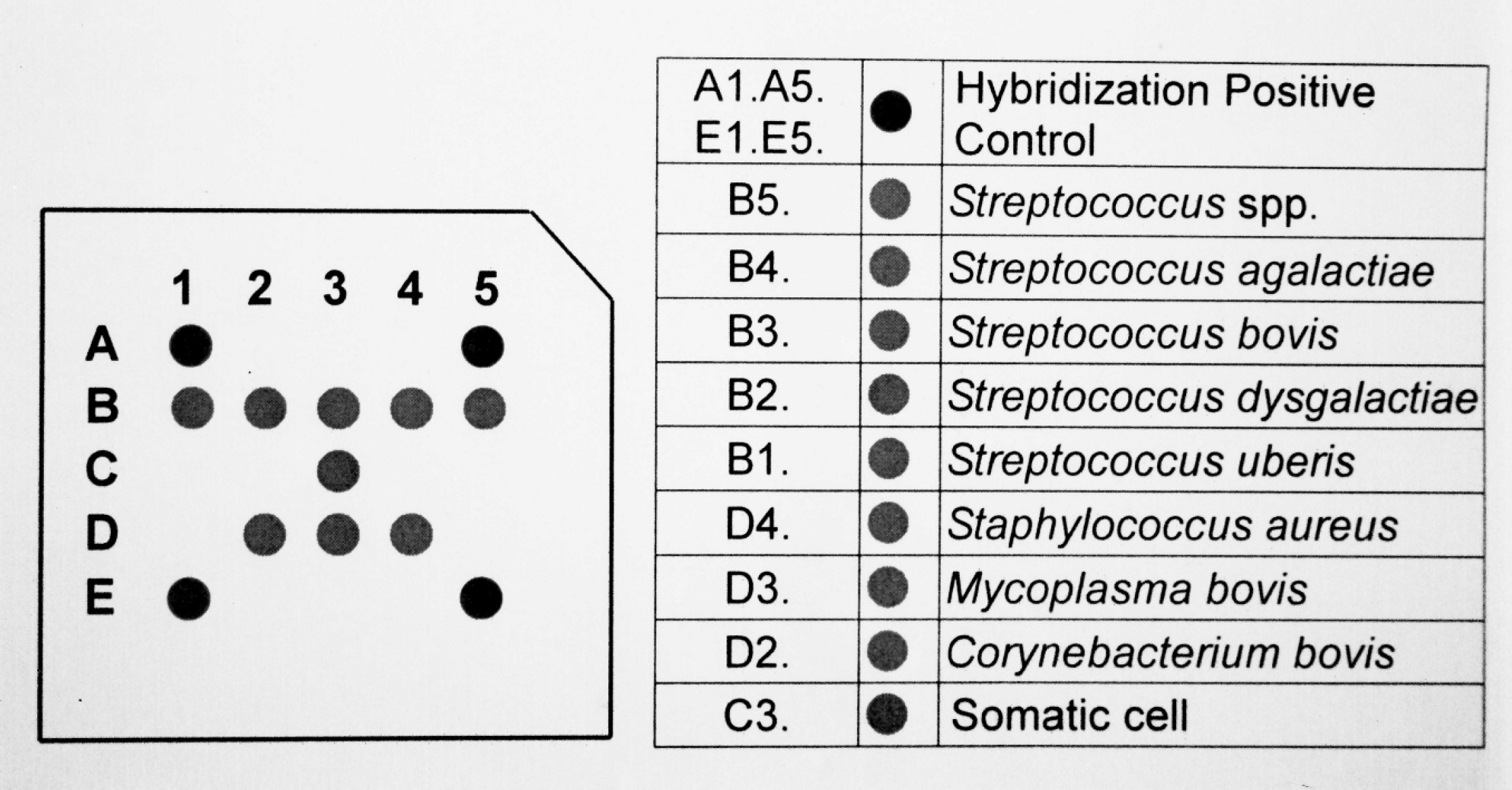
Mastitis biochip. The graph represents the biochip elements. The number inside the chip corresponds to controls or the probes listed in Table 2.
Conventional microbiological analysis and biochip assay, and determination of sensitivity and specificity. Bacterial culture of the milk samples was conducted following the procedures recommended by the National Mastitis Council. 14,22 Briefly, foremilk (10 μl) was plated onto 1 quadrant of a trypticase soy agar plate supplemented with 5% defibrinated sheep blood. j Plates were incubated at 37°C, and the growth of bacteria was observed at a 24-hr interval for 3 days. Bacteria on the primary culture medium were identified tentatively according to the colony morphology, hemolytic characteristics, catalase test, and Gram stain. k Isolates identified presumptively as staphylococci were tested for coagulase by the tube coagulase method, mannitol salt, and DNase agar. Isolates identified presumptively as streptococci were further confirmed by growth in 6.5% NaCl, hydrolysis of esculin, and a Christie, Atkins, and Munch-Petersen (CAMP) reaction. Isolates of Gram-negative bacteria were evaluated by their biochemical reactions on MacConkey agar, triple sugar, iron agar, urea agar, oxidase, motility, indole, and ornithine decarboxylase. Detailed strain identification for streptococci and Gram-negative isolates were carried out by the use of 2 commercial Analytical Profile Index identification systems. 1 The milk samples were tested with the biochip method as described above. With the use of bacterial culture as the gold standard, the relative sensitivity and specificity of the biochip assay were determined with the use of standard formulas. 18
Results
Specificities of PCR primers and biochip
Strain-specific primers were designed for Streptococcus spp., C. bovis, Staph. aureus, and M. bovis. Amplification of the DNA templates with these primers produced single products of the expected size, between 185 and 650 bp, for each strain (Fig. 2). To check the specificity of the biochip, cross-reactivity with DNA isolated from pure cultures of 148 type strains and 4 locally isolated Staph. aureus strains was examined. Only strains corresponding to these strain-specific probes showed positive results, and no cross-reactivity with other strains was observed (Table 1). The mastitis biochip contained 14 spots arranged in 5 columns (Fig. 1). The specificity can be determined by the hybridization pattern displayed on the chip. Each of the 7 strains included on the biochip has a unique hybridization pattern (Fig. 3). All tested strains show color on the spots of somatic cell (C3), PCR-positive control (D5), and hybridization-positive control (A1, A5, E1, E5). The B5 spot was common for all Streptococcus sp., and B1, B2, B3, and B4 spots were specific for S. uberis, S. dysgalactiae, S. bovis, and S. agalactiae, respectively. The D2, D3, and D4 spots were specific for C. bovis, M. bovis, and Staph. aureus, respectively. Strains other than the 7 targeted only showed color on spots for positive control and somatic cells, namely, A1, A5, C3, D5, E1, and E5.
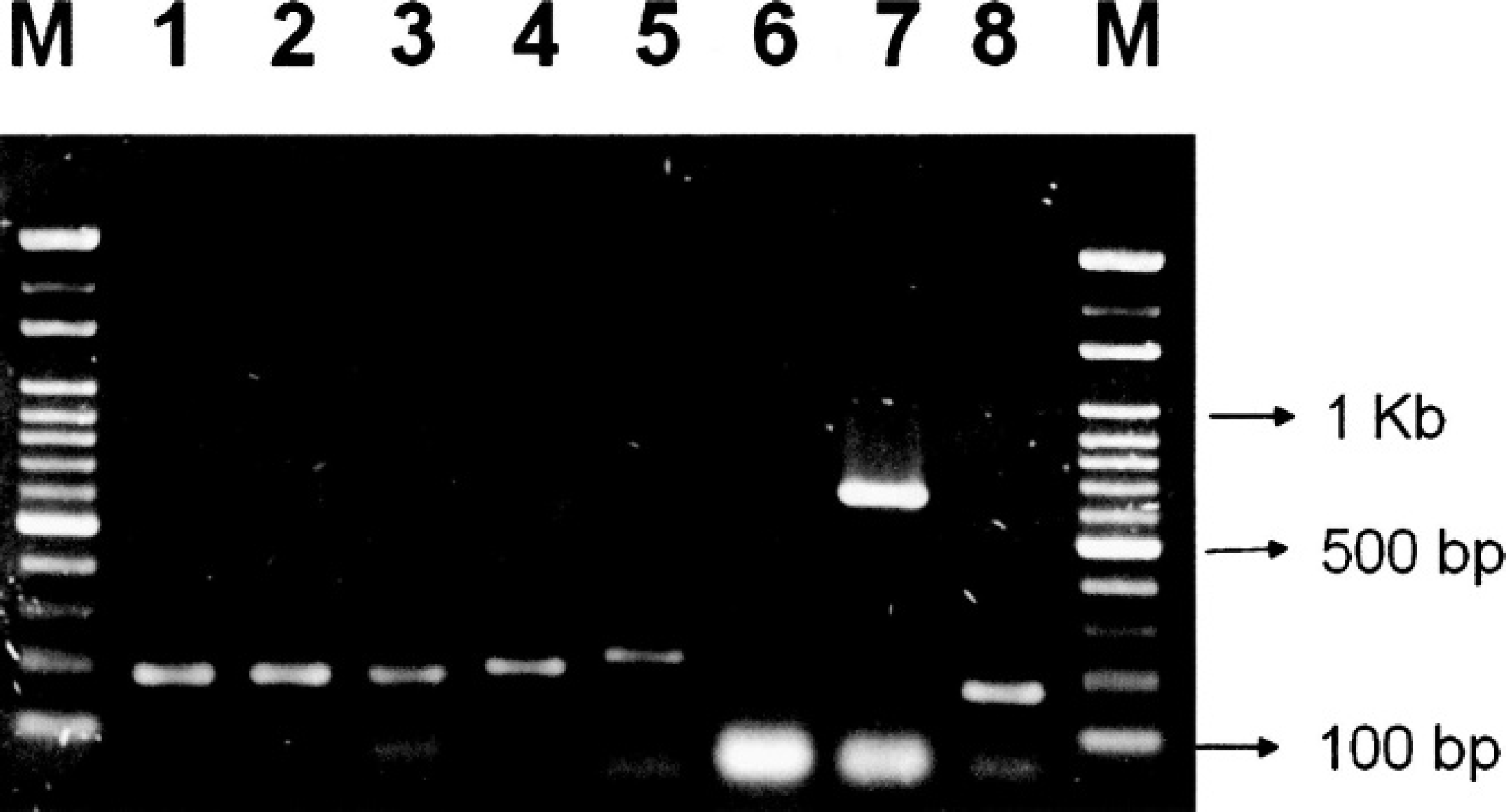
Specificity of multiplex polymerase chain reaction. Products amplified from DNA from Streptococcus spp.
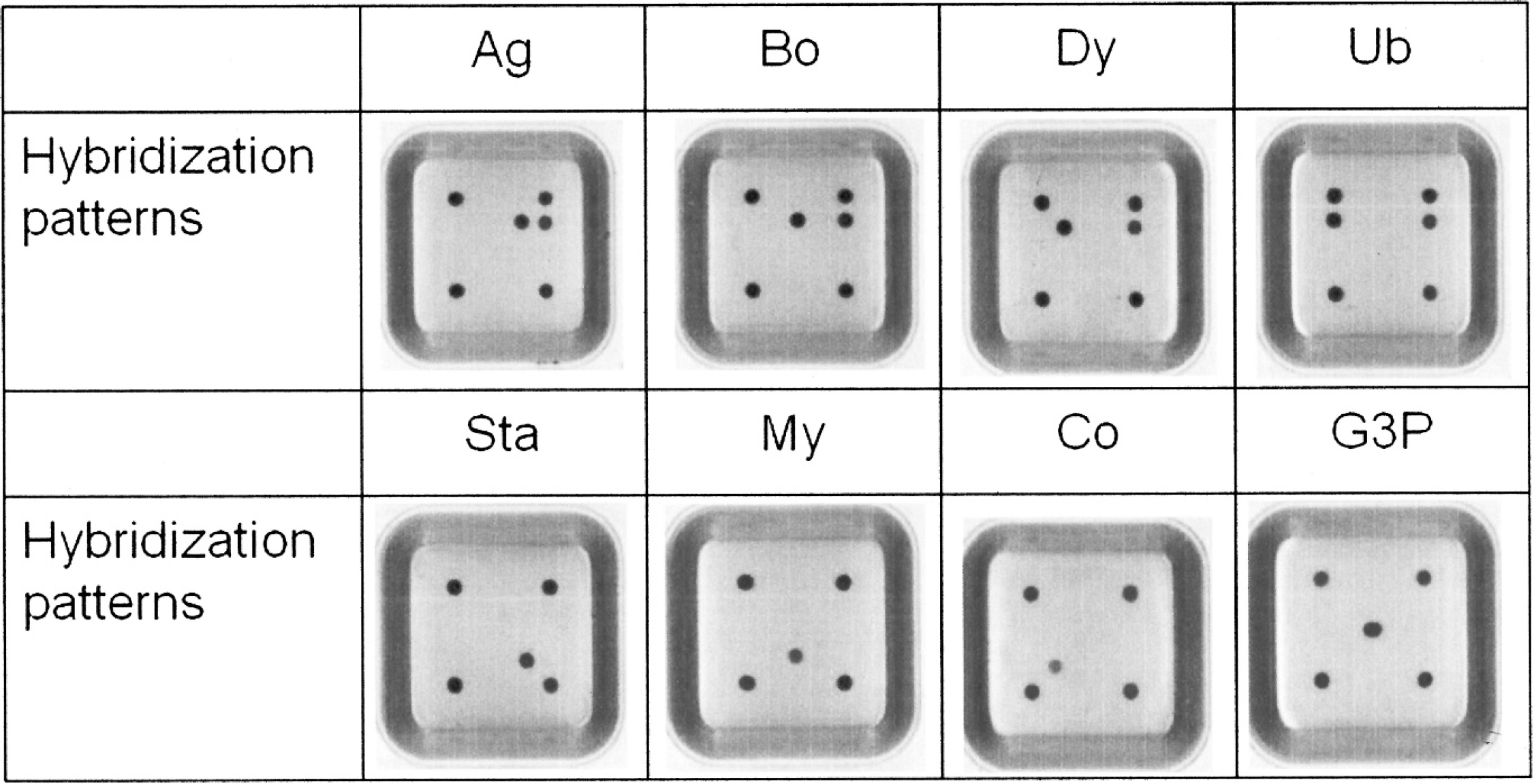

Hybridization patterns of polymerase chain reaction (PCR) amplicons on the biochip. Ag = Streptococcus agalactiae; Bo = Streptococcus bovis; Dy = Streptococcus dysgalactiae; Ub = Streptococcus uberis; Co = Corynebacterium bovis; Sta = Staphylococcus aureus; My = Mycoplasma bovis; G3p = glyceraldehyde-3-phosphate dehydrogenase gene for PCR- and hybridization-positive controls.
Detection limit of the biochip
The lowest detectable concentration of bacteria was determined for each strain (Table 4). The detection limit for S. bovis and C. bovis was 103 and 105 CFU/ml, respectively. For the other 5 strains, the detection limit was 104 CFU/ml.
Sensitivity and specificity of the biochip for milk samples
A trial of this biochip was implemented with the use of randomly collected quarter milk samples (n = 82) that were previously analyzed by conventional microbiological methods. From the bacteriology, 21 samples contained bacteria, including 10 S. uberis, 5 S. dysgalactiae, 2 S. bovis, and 4 Enterococcus faecalis, and the rest (61 samples) were negative. The number of positive samples identified by the biochip was 16, including 10 S. uberis, 5 S. dysgalactiae, and 1 S. bovis, with 1 false negative (S. bovis) and 4 E. faecalis samples that were not detectable by the biochip (Table 5). Therefore, the relative sensitivity of the biochip for the 7 targeted species was 94.1% (16/17). In contrast, those (n = 61) verified by the biochip as negative were in agreement with the results from bacteriology and no false positive occurred. Thus, the relative specificity of the biochip was 100% (61/61). The biochip assay was performed twice on each sample and the 2 runs yielded identical results.
The sensitivity and detection limits of the biochip assay determined with the use of pure bacterial cultures.
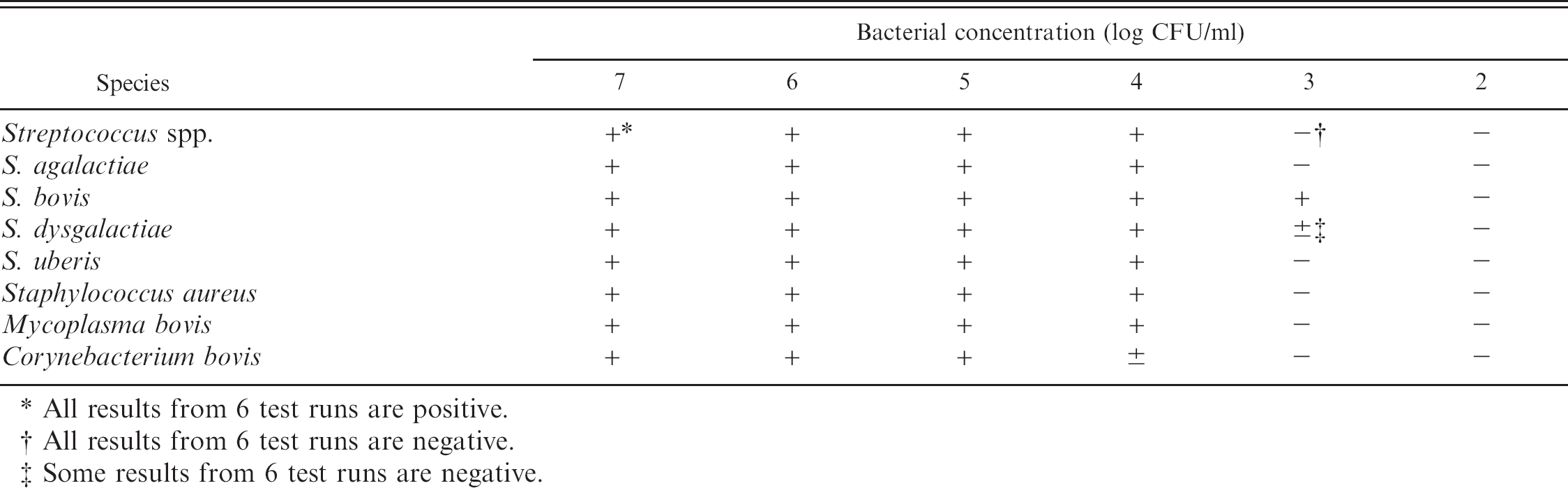
All results from 6 test runs are positive.
All results from 6 test runs are negative.
Some results from 6 test runs are negative.
Results of 82 quarter milk samples evaluated by the biochip for detection of Corynebacterium bovis, Staphylococcus aureus, Mycoplasma bovis, and Streptococcus spp. S. agalactiae, S. bovis, S. dysgalactiae, and S. uberis.
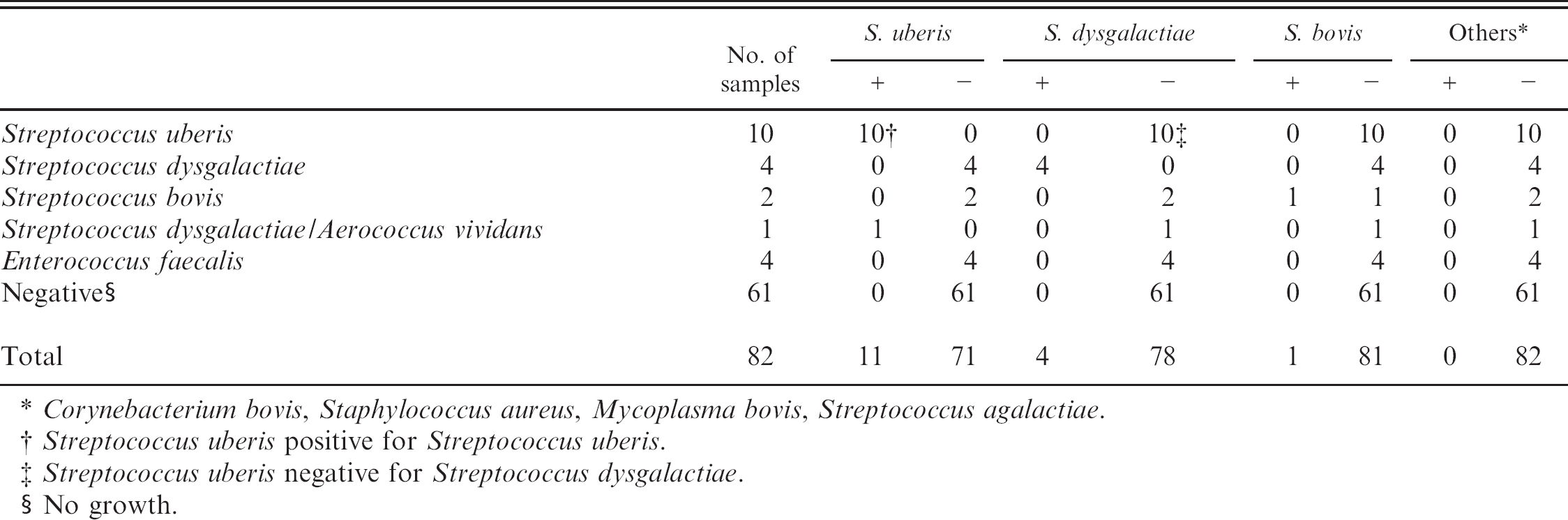
Corynebacterium bovis, Staphylococcus aureus, Mycoplasma bovis, Streptococcus agalactiae.
Streptococcus uberis positive for Streptococcus uberis.
Streptococcus uberis negative for Streptococcus dysgalactiae.
No growth.
Discussion
Despite a great deal of research devoted to preventing bovine mastitis, the incidence of mastitis is still high, which leads to tremendous economical losses to the dairy industry. Staphylococcus aureus, Streptococcus agalactiae, coliform bacteria, and environmental Streptococcus spp. remain the most prevalent mastitis-causing pathogens. 27 Identification of mastitis pathogens is usually performed by traditional culture followed by biochemical tests on bacterial isolates. 23 Identification of bacteria in most clinical laboratories is based on analysis of phenotypic characteristics with the use of biochemical tests, serotyping, and enzymatic profiles. Advantages associated with conventional culture methods are that viable bacteria can be identified as the causative agent of mastitis and antimicrobial susceptibilities can be performed to provide information for selection of appropriate antimicrobial therapy. However, several disadvantages are associated with conventional culture methods. A negative culture can result from residual antibiotics after antibiotic therapy or from low numbers of pathogens in the sample. Presence of leukocytes in milk from cases of clinical mastitis can also result in a negative culture. In addition, intracellular pathogens capable of surviving in mammary cells might not be detected by bacterial culture. 26 Conventional procedures for the identification of bovine mastitis pathogens are labor intensive and take at least 2–3 days to yield a positive result. 18 Inadequate pathogen detection or confirmation techniques have often delayed timely intervention in disease control. Therefore, new methods that are able to circumvent these drawbacks would facilitate a more efficient and precise identification. Molecular methods have been shown to be potential approaches for such purposes. Methods that use PCR based on the 16S–23S rRNA region sequences have been applied successfully for the identification of many bacteria. 6,9,14,22,26,29 Multiplex PCR methods targeting the 16S–23S rRNA spacer regions were developed to detect Staph. aureus and Streptococcus spp. S. agalactiae, S. dysgalactiae, and S. uberis. 25,26 Although DNA arrays have been most widely used by the genomics research community to study gene expression patterns and identify new genes, many researchers have speculated that they might also prove useful in DNA-based diagnostics. Recently, DNA microarray technology has proven to be a highly sensitive and accurate tool for distinguishing closely related bacterial species. 6,16 The major advantages of microarray derive from its ability to use only nanograms of nucleic acids and handle thousands of samples simultaneously, as well as its sensitivity, specificity, rapidity, and ease of analysis. In this study, the authors aimed to use microarray technology in developing a biochip capable of detecting and identifying various strains of bovine mastitis pathogens within hours. To the authors' knowledge, this is the first biochip described for this purpose used on pathogens directly from milk.
The results of this study have shown that robotically spotted DNA biochips can be used to facilitate the identification of mastitis pathogens in conjunction with multiplex PCR methods. Because the method exploits the specificity of both PCR and DNA hybridization reactions, it offers a degree of accuracy superior to those of many other methods commonly used to detect PCR-amplified DNA. The mastitis biochip was designed to detect Staphylococcus aureus, Streptococcus agalactiae, C. bovis, M. bovis, and environmental Streptococcus spp., including S. agalactiae, S. dysgalactiae, and S. uberis. The biochip test displays excellent specificity and good sensitivity.
The detection limit of the method was found to be in the range of 103–105 CFU/ml. A similar level of sensitivity (3.125 − 102 to 5 − 103 CFU/ml) for the detection of Staph. aureus in milk by PCR was previously reported. 26 In another study, the detection limit of multiplex PCR assay described for the detection of mastitis pathogens was reported to be 103–104 CFU/ml. 10 When DNA was extracted directly from dilutions of milk with the phenol-chloroform method, the multiplex PCR had a detection limit of 106 CFU/ml for all 4 organisms (Staph. aureus and Streptococcus spp. S. agalactiae, S. dysgalactiae, and S. uberis). 26 Presence of PCR inhibitors in milk and a variety of other clinical samples such as urine, blood, and feces have been reported. 13,29,31 However, the exact inhibitors in milk have not been identified. Other methods to overcome PCR inhibitors present in milk involve dilution and washing of the sample. Addition of an enrichment step has been reported and appears to be necessary for detecting low numbers of bacteria (<1,000 CFU/ml). For example, in a study 9 using the PCR method for detection of S. agalactiae in milk by targeting the conserved areas within 16S rRNA, the sensitivity of the assay increased from 104–105 to 1 CFU/ml after overnight selective enrichment. Another study 26 needed enrichment in a multiplex PCR for detection of Staph. aureus and Streptococcus spp. S. agalactiae, S. dysgalactiae, and S. uberis to detect levels of 1 CFU/ml, but it is more time-consuming.
Although the sensitivity of the DNA extraction method is important, many additional factors must be considered, including time required, cost per test, and the need for specific reagents. 4 Processing time varied among procedures. Sample processing by biochip assay required approximately 50 min; processing time by the other methods varied from 130 to 140 min, 4,7 and up to 3 hr for the DNA isolation kit. m In total, the method described herein requires a total time of less than 6 hr (for DNA extraction, 50 min; PCR amplification, 140 min; DNA hybridization, 70 min; and colorimetric reaction, 60 min).
Results are interpreted by the pattern formed on the chip. Sequence-specific oligonucleotide probes are designed and prespotted on the chip to capture bacteria strain–specific amplicons. After hybridization and colorimetric development, a perfectly matched probe–target hybrid will form a blue-purple precipitate on the chip as a result of enzymatic reaction. The pattern developed on the chip was recorded with the naked eyes. To improve the accuracy of test results, the chip patterns are captured and analyzed by a commercial, high-throughput genetic identification reader. f
Several disadvantages are associated with the mastitis biochip assay. The complete biochip system could be suitable for clinical laboratories with expertise in molecular biology techniques, despite the fact that the method requires relatively little hands-on training. Also, because much of the cost of the system lies in the DNA extraction of bacteria and multiplex PCR steps, the biochip system might not be affordable to farmers. The biochips are expected to cost between $15 and $20 (data not shown) each. Additionally, it can detect only 7 mastitis pathogens, and without quantities.
Results of this study indicate that the biochip has the potential to be a rapid, sensitive, and specific diagnostic method for simultaneous identification of multiple species of mastitis-causing pathogens in milk. The inclusion of probes for additional mastitis pathogens within the biochip could, in principle, further increase the overall practicability of the system, and the biochip system does have the ability to accommodate another 25–50 elements. Although this assay cannot replace conventional susceptibility testing, the high specificity of the system for detecting 7 mastitis pathogens could facilitate the early diagnosis and control of bovine mastitis.
Acknowledgements
The authors thank Sino Wang of DR. Chip Biotechnology Inc. for the use of their laboratory for conducting the biochip assay. This work was supported by the Council of Agriculture, Executive Yuan, Republic of China.
Footnotes
a.
ColimastTM Test, ICP Biotechnology Ltd., Auckland, New Zealand.
b.
American Type Culture Collection (ATCC), Manassas, VA.
c.
Bioresource Collection and Research Center (BCRC), Hsinchu, Taiwan, Republic of China.
d.
Department of Veterinary Medicine, Chunghsing University, Taichung, Taiwan, Republic of China.
e.
Premier Biosoft International, Palo Alto, CA.
f.
DR. Chip HybTM buffer, DR. Chip hybridization oven, Strep-AP (Streptavidin conjugated alkaline phosphatase), DR. AiMTM Reader, DR. Chip Biotechnology Inc., Chu-Nan, Miao-Li, Taiwan, Republic of China.
g.
UV Box, UV Light Enterprise Co., Taichung, Taiwan, Republic of China.
h.
QIAamp® DNA Mini Kit, Qiagen GmbH, TAIGEN Bioscience Corporation, Taipei, Taiwan, Republic of China.
i.
Promega Inc., Taipei, Taiwan, Republic of China.
j.
Becton Dickinson Co., Franklin Lakes, NJ.
k.
Baso Diagnostic Inc., Taipei, Taiwan, Republic of China.
l.
API 20 Strep System and API 20E System, respectively, bioMérieux, Marcy l'Etoile, France.
m.
PureGene DNA isolation kit (Gentra Systems), Biocompare Inc., San Francisco, CA.
