Abstract
In the current study, the development and validation of a real-time polymerase chain reaction (PCR) assay using a TaqMan-labeled probe for the detection of Actinobacillus suis from porcine lung samples is described. This real-time PCR amplified a 110-bp region of the 23S ribosomal RNA gene from A. suis but not from other bacteria. First, the assay was validated with 183 bacterial strains representing different species of bacteria. Subsequently, 85 porcine lung specimens that were declared A. suis–positive and –negative by bacterial culture and identification were tested to assess whether it can be performed directly on tissue specimens. The bacterial culture results and real-time PCR results agreed across all the samples tested assigning 100% positive and negative predictive values to the PCR. Further, the detection limit of the assay was 380 colony-forming units (CFU) per ml or approximately 1 CFU per reaction. In conclusion, the TaqMan real-time PCR assay described herein is a highly specific, sensitive, and reproducible test, which can be used to detect A. suis DNA in porcine lung specimens, thus providing a timely diagnosis.
Introduction
Actinobacillus suis is a Gram-negative bacterium that belongs to the Pasteurellaceae family. It is a member of the resident flora of tonsils, nasal cavity, and vaginal mucosa of healthy pigs.1,7 However, A. suis is regarded as an important swine pathogen due to its ability to cause fatal disease in pigs. Until recently, A. suis was considered as a pathogen of high status health herds, and specifically of sucking and recently weaned pigs (Sanford SE: 1995, Actinobacillus suis: an overview of an emerging disease. Proceedings of the American Association of Swine Practitioners).3-5 According to the latest evidence, A. suis infections pose a serious threat to both conventional and high health status swineherds, and also affect both young and adult pigs.4,8,9 Absence of clinical signs or lesions pathognomic to A. suis infection makes its diagnosis difficult (Oliveira S: Actinobacillus suis detection by PCR and strain characterization by genotyping. Available at: http://www.cvm.umn.edu/vdl/prod/groups/cvm/@pub/@cvm/@vdl/documents/asset/cvm_62339.pdf).2,7,8 In particular, A. suis septicemia in neonates can easily be confused with septicemic lesions caused by Escherichia coli.2,7,8 Similarly, pneumonic lesions caused by A. suis and Actinobacillus pleuropneumoniae are comparable.7,8 Also, fibrinous pleuritis and pericarditis lesions produced by Haemophilus parasuis resemble those caused by A. suis.7,8 Similarly, rhomboid erythematous skin lesions produced by Erysipelothrix rhusiopathiae and A. suis are very similar to each other and indistinguishable clinically.2,6-8 Therefore, diagnosis of A. suis infection requires isolation of the organism from clinical specimens of the affected pigs. However, isolation of A. suis from clinical specimens may not always be possible for many reasons, such as treatment of animals with antibacterial agents prior to sample collection and/or improper sample handling, storage, and transport techniques, which highlights the need for alternative methods to diagnose A. suis infection. Recently, a gel-based polymerase chain reaction (PCR) based on the 16S ribosomal (r)RNA gene has been developed (Oliveira S: Actinobacillus suis detection). Yet, this PCR is unable to differentiate A. suis from Actinobacillus equuli subsp. equuli, which is genetically very close to A. suis (Oliveira S: Actinobacillus suis detection). Therefore, the present study aimed to develop and validate a TaqMan real-time PCR assay for the routine detection of A. suis from clinical specimens.
Materials and methods
Bacteria
To determine the specificity and sensitivity of the real-time PCR assay, a total of 183 bacterial isolates belonging to 16 different species were tested (27 isolates of A. suis, members of the commensal flora of the swine respiratory tract, bacteria that cause clinical signs and lesions similar to those of A. suis infections, and/or bacteria that are closely related to A. suis; Table 1). Field strains were isolated from clinical specimens submitted to Iowa State University Veterinary Diagnostic Laboratory (ISU-VDL; Ames, IA) and Minnesota Veterinary Diagnostic Laboratory (St. Paul, MN).
Actinobacillus suis and other bacterial species and strains used in the current study.*
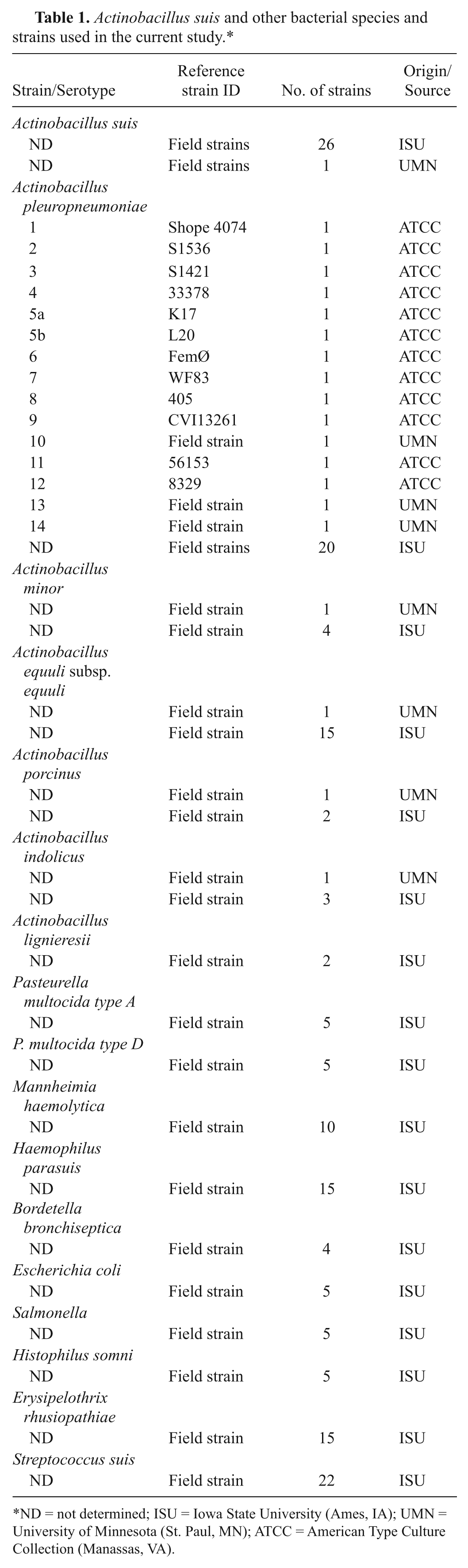

ND = not determined; ISU = Iowa State University (Ames, IA); UMN = University of Minnesota (St. Paul, MN); ATCC = American Type Culture Collection (Manassas, VA).
Clinical specimens
A total of 85 porcine lung samples submitted to ISU-VDL were tested in the current study. All samples were cultured for the isolation and identification of bacteria prior to being tested by PCR. The PCR technician was blinded to any clinical diagnosis or bacterial culture results. Samples were either processed immediately upon submission to the laboratory or the following day. Samples were stored at 4°C until they were processed.
Genomic DNA and polymerase chain reaction sample preparation
A commercial kit a was used in accordance with manufacturer’s recommendations to extract DNA. Specifically, to extract DNA from lung specimens, approximately 4 g of each tissue was incised with a sterile scalpel blade and placed in a stomacher bag containing 20 ml of Earle balanced salt solution. b The sample was then homogenized with a stomacher at a high speed for 60 sec. The resultant supernatant was stored at −20°C until it was used to extract DNA. At the time of DNA extraction, 100 µl of liquid homogenate was mixed with 180 µl of buffer ATL supplied with the kit, and DNA was extracted following the kit’s spin-column protocol for “Purification of Total DNA from Animal Tissues.” To extract DNA from bacterial isolates listed in Table 1, bacteria were grown in pleuropneumonia-like organism (PPLO) broth, c Columbia broth d supplemented with beta-nicotinamide adenine dinucleotide, b or brain heart infusion broth c overnight at 37°C, after which the broth cultures were used for DNA extraction. A bacterial culture containing approximately 109 cells were centrifuged at 5,000 × g for 10 min, and the pellet was resuspended in 180 µl of buffer ATL before proceeding with the kit’s protocol for “Purification of Total DNA from Animal Tissues.” Both sample types were eluted with 200 µl of elution buffer supplied with the kit. The elutes contained 4.8–10.2 µg of DNA as determined with a spectrophotometer. e
Primer and probe design
The 23S rRNA gene sequences of A. suis (EU333989) and related bacterial species were obtained from the NCBI GenBank database. The sequences were aligned using the ClustalW multiple sequence alignment program MegAlign f to determine the conserved and variable regions. The primers and probe were designed to amplify sequences present in A. suis but absent from other bacterial species, and were analyzed for guanine–cytosine content, melting temperature value, primer-dimer formation, and hairpin structure formation using the Primer Express 1.5 program. g The probe, a 27-mer oligonucleotide, was labeled with cyanine (Cy)5 as the reporter dye and was quenched with Black Hole Quencher 1. Primers and probes were proprietary. h The forward primer AS1 (5′-GAGCTGGGAAGCTCGACTAT-3′), the reverse primer AS2 (5′-CCCCCATCTTCAAACAGGAT-3′), and the probe AS (CY5-AGCTAACGACAAGTAGGGCG-BHQ2) amplified a 110-bp region of the 23S rRNA-encoding region of A. suis.
Real-time polymerase chain reaction
Amplification was performed in a total volume of 20 µl in 96-well plates. Each reaction contained 2.5 µl of extracted DNA, 10 µl of 2× TaqMan Universal PCR Master Mix, g 400 nM of each primer pair, and 100 nM of the probe. Finally, the volume was adjusted to 20 µl by the addition of autoclaved purified water. i The PCR was performed on a real-time PCR system g using the following program: 1 cycle of 15 min at 95°C, and 45 cycles of 15 sec at 94°C and 1 min at 60°C. The results were subsequently analyzed using the Sequence Detection System 1.7 software. g A true PCR amplification was reported as an increase in the reporter dye fluorescence of the probe paralleled to the decrease in its quencher dye fluorescence. All PCR assays were conducted in duplicate. The threshold cycle (Ct) was set at 35 based on the amplification curves generated by known positive and negative controls used in the assay (Table 1). If the Ct value was in excess of 35, the sample was considered negative for A. suis.
Results
Optimization of real-time polymerase chain reaction conditions
The primers and the TaqMan probe targeted a hypervariable region of the 23S rRNA gene. To optimize PCR conditions, a gel-based, conventional gradient PCR was performed using DNA extracted from a field isolate of A. suis as template DNA using different cycling parameters. The highest temperature that yielded a significant specific product (110 bp) on a 2% agarose gel visualized under ultraviolet light after ethidium bromide staining was used as the annealing temperature in the subsequent real-time PCR assay (data not shown).
Specific detection of A. suis by real-time polymerase chain reaction
The real-time PCR assay successfully amplified DNA from all A. suis isolates listed in Table 1. The Ct values ranged from 20.06 to 26.49. No positive result was produced by DNA extracted from bacterial species other than A. suis (Table 2). Similarly, positive PCR results were detected only with lung samples that were culture positive for A. suis by bacterial isolation and identification techniques. Accordingly, the positive and negative predictive values of this real-time PCR were determined to be 100% with both bacterial cultures and clinical specimens (Tables 2, 3).
Real-time polymerase chain reaction (PCR) results for Actinobacillus suis and non–A. suis bacterial strains used in the current study.*
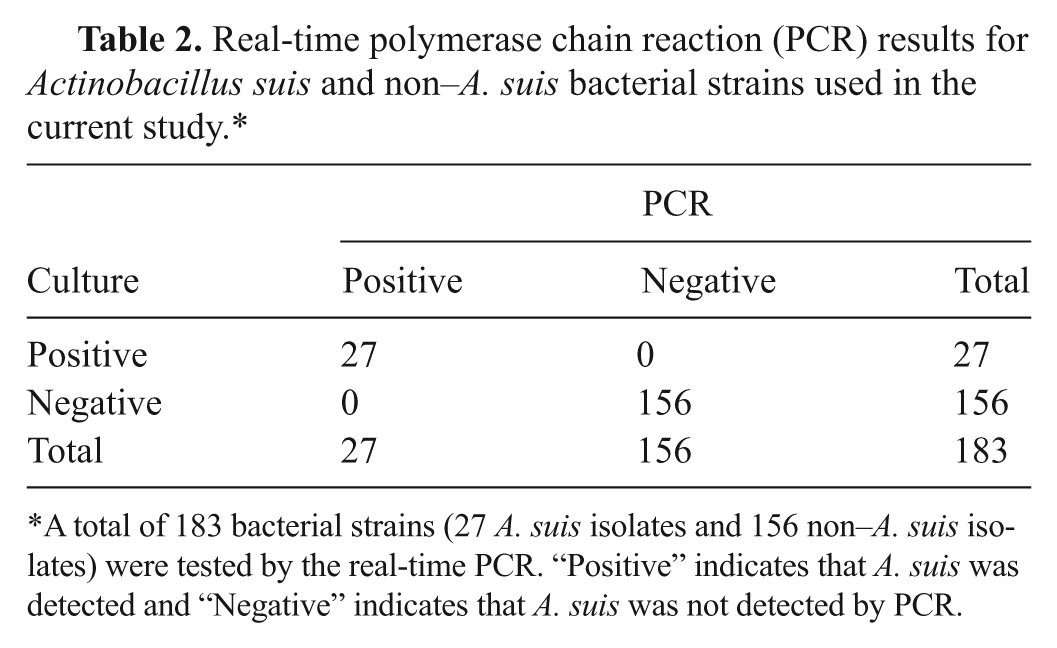
A total of 183 bacterial strains (27 A. suis isolates and 156 non–A. suis isolates) were tested by the real-time PCR. “Positive” indicates that A. suis was detected and “Negative” indicates that A. suis was not detected by PCR.
Agreement between Actinobacillus suis detection in lung specimens by real-time polymerase chain reaction (PCR) and bacterial culture.*
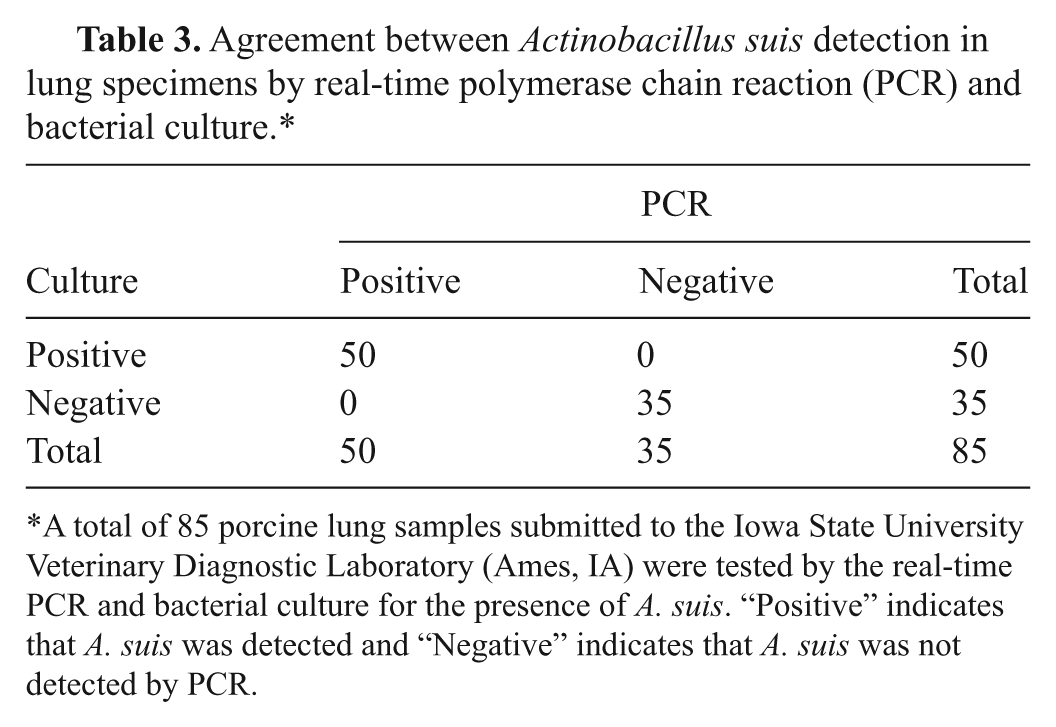

A total of 85 porcine lung samples submitted to the Iowa State University Veterinary Diagnostic Laboratory (Ames, IA) were tested by the real-time PCR and bacterial culture for the presence of A. suis. “Positive” indicates that A. suis was detected and “Negative” indicates that A. suis was not detected by PCR.
Analytical sensitivity of A. suis real-time polymerase chain reaction
The analytical sensitivity of the real-time PCR assay was evaluated by testing 2 different types of templates: A. suis broth cultures and A. suis–negative lung specimens that were spiked with a known number of A. suis. The sensitivity was first determined by testing DNA extracted from sequential 10-fold dilutions of broth cultures of 5 different A. suis field isolates. Each DNA extract was tested in triplicate. Samples containing as few as 380 colony-forming units (CFU) of A. suis/ml or 1 CFU of A. suis/reaction (2.5 µl) tested positive by the assay. Next, 7 lung specimens that were confirmed A. suis negative by bacterial culture and identification techniques were spiked with a pre-determined number of A. suis. Based on the results obtained with serially diluted A. suis cultures, 6 different concentrations of A. suis were used to spike the samples. Briefly, each lung specimen was spiked with approximately 1 × 104, 6 × 103, 5 × 103, 4 × 103, 3 × 103, or 2 × 103 CFU of bacteria per gram of tissue. DNA was extracted, and the samples were tested in duplicate by PCR. Actinobacillus suis was detected only from the samples spiked with ≥3 × 103 CFU/g of tissue giving a detection limit of approximately 1 CFU/reaction (2.5 µl).
Discussion
Diagnosis of A. suis infection in pigs mostly relies on culture and identification of A. suis from clinical specimens, which can take 2–3 days. Although bacterial culture is considered the “gold standard” diagnostic technique for the diagnosis of many bacterial infections including A. suis, a limitation is that it requires the presence of live bacteria in the specimen for detection. However, because animals are often treated with antibacterial agents prior to the collection of sample(s), bacteria in the specimen might be dead at the time of testing. This can also result from improper handling of the specimen and/or suboptimal specimen transport and storage conditions. The devastating effects imposed to the swine industry by A. suis infections and the absence of pathognomic clinical signs or lesions present in diseased pigs make it highly desirable to have a rapid detection method to diagnose A. suis infections.7,8 The diagnostic tool developed in the present study will offer the opportunity to implement timely intervention strategies to control A. suis outbreaks and minimize losses to producers. Currently available A. suis 16S rRNA gel-based PCR can be performed directly on clinical specimens such as lung and tonsillar tissues with a detection limit of 7.2 × 102 CFU/ml but one drawback of the assay is that it detects both A. suis and A. equuli subsp. equuli DNA, thus compromising its specificity (Oliveira S: Actinobacillus suis detection).
Real-time, fluorescence-based assays have several advantages over conventional gel-based PCR assays including high specificity provided by inclusion of a probe in addition to 2 primers. Many PCR assays that are available to detect bacterial pathogens use 16S and 23S rRNA genes as the target sequence. These 2 genes are universally present in bacteria but contain both conserved and variable regions within the sequences. Bacterial speciation techniques use primers complementary to 2 conserved regions flanking a hypervariable region followed by sequencing of the PCR amplified variable region. Similarly, PCR primers designed to contain the sequences complementary to hypervariable regions of a bacterial species can be used to detect a particular bacterium at the species level. Nevertheless, this is not always possible due to the highly conserved nature of rRNA genes among closely related species of bacteria and in particular, different species of bacteria belonging to the same genus or family.
In the current report, a real-time PCR assay using a TaqMan-labeled probe for the detection of A. suis DNA directly from clinical specimens is described. The assay, which amplified a 110-bp region of the 23S rRNA gene of A. suis, was highly specific when tested with numerous bacterial species that are known to inhabit the porcine respiratory tract and/or are closely related to A. suis. The assay detected exclusively A. suis DNA with positive and negative prediction values of 100% as compared to conventional bacterial culture technique, which is considered the current “gold standard” for the isolation and identification of A. suis. Further, the detection limit of the assay was evaluated using serial dilutions of an overnight broth culture of A. suis. The assay could detect as low as 380 CFU/ml or 1 CFU/reaction (2.5 µl). Lung tissues and cultures were the only specimen types included in the present assay validation. It was not determined, but is likely that this PCR is also effective for testing other specimen types such as tonsil, liver, spleen, and bronchoalveolar lavage. In the future, the assay developed in the current study may be used to develop a multiplex real-time PCR assay for the simultaneous detection of pathogens causing respiratory diseases in pigs. Such assays will significantly save time and money in terms of disease diagnosis; however, traditional techniques such as bacterial culture and virus isolation may still be required for downstream applications such as autogenous vaccine preparation and antimicrobial susceptibility testing, which demand the availability of live organism.
In summary, a real-time TaqMan PCR assay was developed to detect A. suis DNA directly from porcine lung specimens. The optimized assay was highly specific, sensitive, and reproducible, and produced results comparable to that of bacterial culture technique but within only 3 hrs. In the future, this PCR technique could be used as a diagnostic and research tool for the detection and quantification of A. suis DNA directly from clinical samples.
Footnotes
Acknowledgements
The authors wish to thank Dr. Simone Oliveira of the Minnesota Veterinary Diagnostic Laboratory and bacteriology section of the Iowa State University Veterinary Diagnostic Laboratory for providing bacterial strains used in the study.
a.
DNeasy® Blood and Tissue Kit, Qiagen Inc., Valencia, CA.
b.
Sigma-Aldrich, St. Louis, MO.
c.
Difco Inc., Detroit, MI.
d.
BBL™, Becton-Dickinson Inc., Sparks, MD.
e.
NanoDrop®, Thermo Fisher Scientific Inc., Waltham, MA.
f.
DNASTAR Inc., Madison, WI.
g.
ABI™ 7500 real-time PCR system: Applied Biosystems, Foster City, CA.
h.
Integrated DNA Technologies, Commercial Park, Coralville, IA.
i.
MilliQ water: Millipore, Billerica, MA.
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
The authors received no financial support for the research, authorship, and/or publication of this article.
