Abstract
The current study reports on a real-time reverse transcription polymerase chain reaction (real-time RT-PCR) ring trial for the detection of Classical swine fever virus (CSFV) genomic RNA undertaken by 10 European laboratories. All laboratories were asked to use their routine in-house real-time RT-PCR protocols and a standardized protocol commonly used by the Friedrich-Loeffler-Institute (FLI) on a panel of well-characterized samples. In general, all participants produced results within the acceptable range. The FLI assay, several in-house assays, and the commercial kits had high analytical sensitivity and specificity values. Nevertheless, some in-house systems had unspecific reactions or suboptimal sensitivity with only a single CSFV genotype. Follow-up actions involved either improvement of suboptimal assays or replacement of specific laboratory assays with the FLI protocol, with or without modifications. In conclusion, the ring trial showed reliability of classical swine fever diagnosis on an international level and helped to optimize CSFV-specific RT-PCR diagnostics.
Due to its economic impact, classical swine fever (CSF) ranks among the most important diseases of domestic pigs. Clinical and pathological signs are highly variable, and diagnosis must be confirmed by laboratory tests.5,16 During 2000–2010, the polymerase chain reaction (PCR) technique has become more and more important for routine CSF diagnosis,1,3,4,13,19 and reverse transcription (RT)-PCR has been accepted by the European Union as an official method for CSF confirmation (Commission Decision 2002/106/EC). Meanwhile, various real-time RT-PCR methods for diagnosing CSF have been described, * and the first commercial real-time RT-PCR kits are already available.3,11 Beside the published protocols, a variety of in-house protocols are also used. To guarantee consistent quality and diagnostic reliability, assay protocol optimization, validation, and standardization is needed. In this context, ring trials can help improve protocol standardization and harmonization. For CSF, such ring trials have been performed in the past.18,19
The current study reports on the results of a real-time RT-PCR ring trial for the detection of Classical swine fever virus (CSFV; family Flaviviridae, genus Pestivirus) genomic RNA among 10 European laboratories that are routinely involved in CSF diagnosis. Organized by the Friedrich-Loeffler-Institut (FLI; Riems Island, Germany), the trial was based on well-characterized samples evaluated with 1 standard method by all participating laboratories 7 in comparison to different routine in-house protocols (published and unpublished). The aim of the ring trial was to evaluate the reliability of CSF real-time RT-PCR diagnosis on an international level, and to optimize CSFV-specific RT-PCR diagnostics.
To this means, a test panel was sent out comprising 48 manually extracted a viral RNA samples in RNA-safe buffer (50 ng/µl of carrier polyA-RNA, b 0.05% Tween-20, c 0.05% sodium azide d in RNase-free water) with defined copy numbers including 2 negative controls. Viral RNA from CSFV strains of all available genotypes as well as from related pestiviruses were integrated into the test panel to assess analytical specificity in terms of inclusivity and exclusivity. Furthermore, 3 dilution series of CSFV RNA were used to define differences in the analytical sensitivity. Dilution series were prepared from CSFV strains Alfort (genotype 1.1), Spante (genotype 2.3), and Kanagawa (genotype 3.4), respectively. For the standardization of the ring trial, the protocol commonly used by the FLI 7 was included (hereafter, FLI protocol). The reagents for this assay were transferred together with the viral RNA.
Participating laboratories were asked to use their own routine real-time RT-PCR protocols for CSFV detection (hereafter, in-house protocols, not considering whether such protocols were published, unpublished, or commercially available). In total, 13 additional assays were used, of which only 3 were unpublished. Most protocols were CSFV specific,7,11,12,14,15,20 but 3 protocols (1 unpublished) were Pestivirus specific.22,23 In detail, 3 laboratories used a previously published assay,14,15 while 1 laboratory used a reference method 7 with in-house modifications, and another laboratory a different assay. 20 Two laboratories used a commercial real-time RT-PCR kit. e One of the laboratories combined the commercial assay with an additional commercial kit. f Each of the Pestivirus-specific protocols was used by 1 laboratory.
Most laboratories carried out 2 runs for each method to obtain a final result. For the purpose of the current ring trial, evaluation of results was done on the basis of mean threshold cycle (Ct) values. In cases where only 1 run was positive or Ct values were higher than the internal cut-off of the respective laboratory, results were reported as inconclusive. The same applied for samples with increasing fluorescent values that did not cross the threshold. Results of the organizing laboratory were only used for comparison and were not included in the comparative evaluation.
Results indicated that CSFV samples representing different genotypes were reliably detected in all participating laboratories that used the FLI protocol (Table 1, samples 1–13). Using the in-house protocols, only minor problems were noticed in this section. One laboratory (D) reported 4 inconclusive results, and 1 test system did not detect 1 of the group 3 CSFV strains in 2 laboratories (I1 and J1). While laboratory I1 failed to detect CSFV strain Kanagawa, laboratory J1 did not detect CSFV strain Congenital tremor. Both commercial kits (laboratories B and G) detected all strains included in the ring trial. Detailed results are presented in Table 2 (samples 1–13).
Classical swine fever virus detection: results of mean threshold cycle values obtained using the protocol routinely used by the Friedrich-Loeffler-Institut as reference method.*
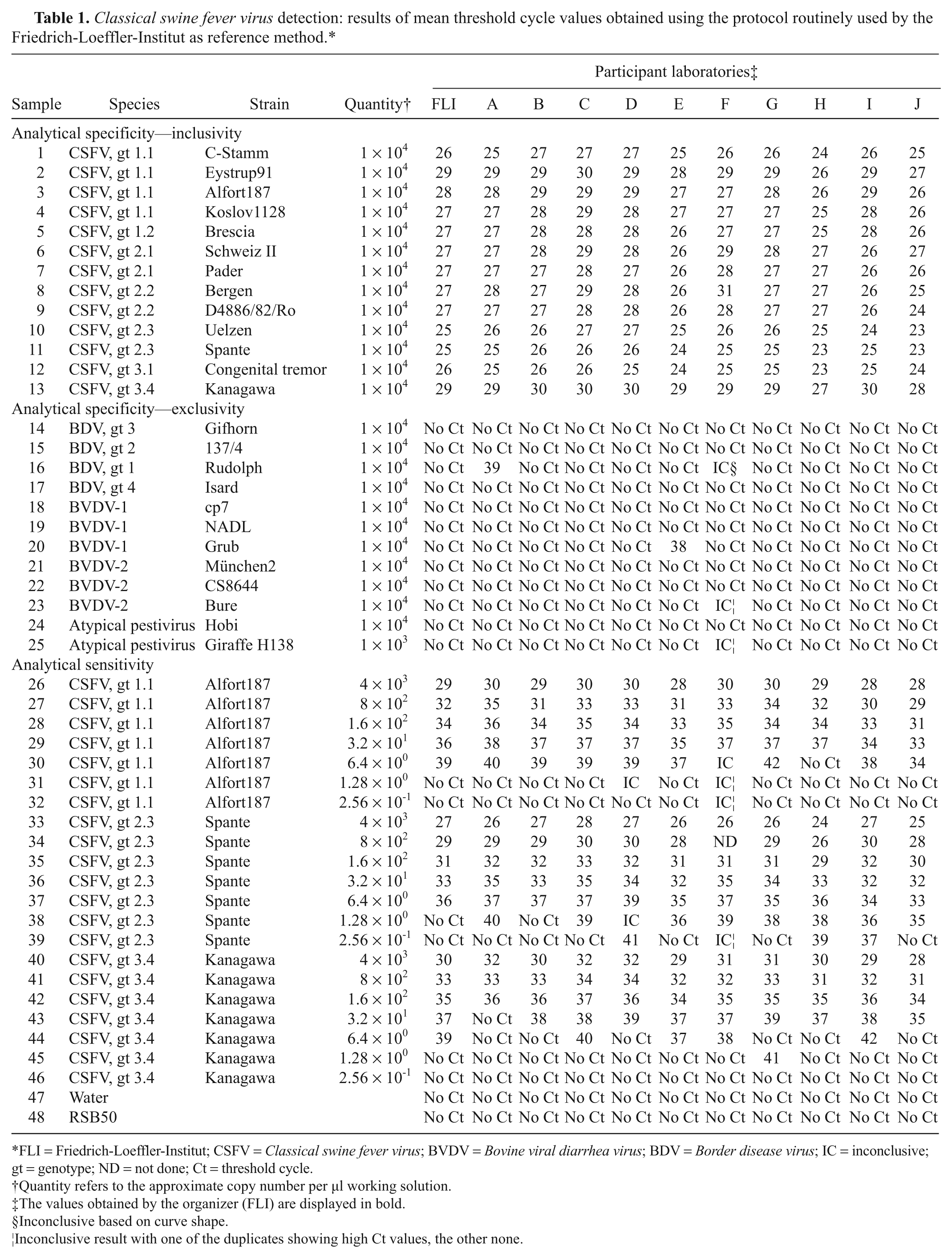
FLI = Friedrich-Loeffler-Institut; CSFV = Classical swine fever virus; BVDV = Bovine viral diarrhea virus; BDV = Border disease virus; IC = inconclusive; gt = genotype; ND = not done; Ct = threshold cycle.
Quantity refers to the approximate copy number per µl working solution.
The values obtained by the organizer (FLI) are displayed in bold.
Inconclusive based on curve shape.
Inconclusive result with one of the duplicates showing high Ct values, the other none.
Classical swine fever virus detection: results of mean threshold cycle values obtained using the in-house methods.*
FLI = Friedrich-Loeffler-Institut; CSFV = Classical swine fever virus; BVDV = Bovine viral diarrhea virus; BDV = Border disease virus; IC = inconclusive; gt = genotype; ND = not done; Ct = threshold cycle.
Quantity refers to the approximate copy number per µl working solution.
Coded participants with superscript figures refer to the reference list for all published protocols. The reference values obtained by the organizer FLI are displayed in bold.
Pestivirus-specific assays.
Inconclusive result with one of the duplicates showing high Ct values, the other none but a curve below cut-off.
In general, neither Border disease virus (BDV) nor Bovine viral diarrhea virus (BVDV), or other related pestiviruses, caused diagnostic problems using the FLI protocol. Only 2 laboratories (A and E) obtained a high Ct value in 1 sample each. Laboratory A reported high Ct values for BDV strain Rudolph, whereas laboratory E had a weak positive result for BVDV strain Grub (Table 1, samples 14–25). The results were not confirmed by the in-house methods employed by laboratories A and E.
One CSFV-specific in-house protocol had problems with 3 out of 4 BDV isolates as well as with BVDV strain CS8644 and Pestivirus Hobi (laboratory J1). In addition, some inconclusive or false-positive results with rather high Ct values occurred in 1 laboratory with 1 commercial test kit (B) and 1 in-house protocol (E). Not all pestiviruses were detected by the Pestivirus-specific protocols (laboratories H, I2, J2). Details can be found in Table 2.
Using the FLI protocol, all laboratories were able to detect the CSFV strain Alfort dilution with 3.2 × 101 copies/µl working solution. Eight out of 10 participating laboratories also detected dilutions containing 6.4 × 100 copies/µl; an additional laboratory scored this dilution as inconclusive. Furthermore, all laboratories were able to detect the dilution 6.4 × 100 copies/µl in the CSFV strain Spante dilution series. Eight out of 10 participating laboratories also scored the 1.28 × 100 copies/µl dilution positive. Three positive results were reported for the highest dilution of 2.56 × 10−1 copies/µl working solution. With the CSFV strain Kanagawa dilution series, all laboratories were able to detect the dilution containing 1.6 × 102 copies/µl using the reference method. 8 All but 1 laboratory also detected the dilution with 3.2 × 101 copies/µl, and 4 laboratories were able to detect the dilution containing 6.4 × 100 copies/µl. One positive result with a very high Ct value occurred in the dilution containing 1.28 × 100 copies/µl after a negative result for the previous dilution. Details are given in Table 1 (samples 26–46).
Most in-house systems did not have problems with the CSFV strain Alfort dilution series. All laboratories detected the second dilution containing 8 × 102 copies/µl with their in-house systems. The third dilution, containing 1.6 × 102 copies/µl, was detected by all but 1 assay. The fourth dilution (3.2 × 101 copies/µl) was detected by 11 out of 13 assays, and 10 assays also detected the fifth dilution containing 6.4 × 100 copies/µl. One assay scored all dilutions positive.
The CSFV strain Spante dilution series had similar results. All assays successfully detected the dilution containing 1.6 × 102 copies/µl, and 11 out of 13 had positive results in the next dilution step as well (3.2 × 101 copies/µl). The dilution with 6.4 × 100 copies/µl was detected in 11 assays again, but 2 results were inconclusive due to negative results in the previous dilution and unexplainable Ct values. The sixth dilution step (1.28 × 100 copies/µl) was detected with 4 assays. Four laboratories recorded high Ct values for the highest dilution step containing 2.56 × 10−1 copies/µl. A comparative analysis can be found in Table 2.
Several in-house assays had problems with the CSFV strain Kanagawa dilution series (Table 2). Two assays did not detect any of the dilutions, and another assay picked up only the first dilution containing 4 × 103 copies/µl. Nevertheless, 9 assays detected the third (1.6 × 102 copies/µl), 8 assays the fourth (3.2 × 101 copies/µl), 3 assays the fifth (6.4 × 100 copies/µl), and 3 assays the sixth (1.28 × 100 copies/µl). Some positive results in higher dilution steps were not accompanied by positive results in the previous dilution steps, or Ct values did not correspond with further dilution. Only 1 assay had problems with both negative controls.
Summarizing the results of the current ring trial, all participants produced results within the acceptable range, and the results indicate that the real-time RT-PCR assay is a rapid and extremely sensitive tool for CSFV detection. Combined with the possibility of using RNA extraction robots and real-time PCR machines with a 96- or 384-sample platform, high throughput, which is required during a crisis or outbreak situation, seems feasible.
In detail, the FLI protocol and several in-house assays (including the commercial kits) had high analytical sensitivity and specificity; however, some in-house systems also amplified related pestivirus RNA (unspecific reactions) or had suboptimal sensitivity with single CSFV genotypes (low sensitivity). The main problems were encountered with CSFV strain Kanagawa, which belongs to genotype 3.4 and is used as an out-group strain in phylogenetic analyses due to its variation from other CSFV strains. In order to understand the apparent failure or inconsistent results of some protocols, further investigations regarding primer and probe sequences as well as other laboratory variables, such as machines, sample handling, enzymes, and human factors, are needed. In general, the ring trial sample panel as well as the experiences with the real-time RT-PCR assays tested in the current ring trial can be used for further development and validation of molecular diagnostic assays for the identification of CSFV and other pestiviruses. 10
In the aftermath of the current ring trial, laboratories with a suboptimal protocol either improved their methods or replaced them with the highly sensitive and specific FLI method, with or without in-house modifications or use of commercial kits. In addition, the current study found that CSFV RNA is very stable in RNA-safe buffer. In conclusion, the current ring trial helped to optimize CSFV-specific real-time RT-PCR diagnostics within the European Union and confirmed that the real-time RT-PCR assay is a reliable and robust tool for CSF diagnosis. Based on the presented findings, minimum standards for diagnostic real-time RT-PCR assays for CSF diagnosis can be discussed in the future.
Footnotes
Acknowledgements
The authors thank all technical assistants involved in assembly of aliquots, standardization of samples, packaging, and shipping. The authors Bernd Hoffmann and Sandra Blome contributed equally.
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
a.
QIAamp® Viral RNA Kit, Qiagen GmbH, Hilden, Germany.
b.
RNA homopolymer, Amersham Biosciences Europe GmbH, Freiburg, Germany.
c.
Tween 20, Sigma-Aldrich Chemie GmbH, Munich, Germany.
d.
Sodium azide, Sigma-Aldrich Chemie GmbH, Munich, Germany.
e.
TaqVet PPC, Laboratoire Service International, Lissieu, France.
f.
Adiavet, Adiagene, Saint-Brieuc, France.
The Network of Excellence for Epizootic Disease Diagnosis and Control or EPIZONE (contract no. FOOD-CT-2006-016236) supported the current work.
