Abstract
A quantitative real-time PCR (qPCR) for detection of the neurotoxin of the Clostridium botulinum type C (BoNTC) encoding gene has been compared with a nested PCR (nPCR) and a conventional PCR (cPCR) using 2 toxigenic C. botulinum C1 reference strains and samples from bird tissues (n = 30) and sediments (n = 30) from wetlands where botulism outbreaks have been reported. A cPCR based on 16S ribosomal RNA sequences from 60 strains of Clostridium species was also developed to detect the genomic DNA of C. botulinum C in order to evaluate the presence of nontoxigenic strains. Quantitative PCR showed a similar sensitivity to nPCR (<0.5 pg of DNA), and both were more sensitive than the cPCR when using the C. botulinum reference strains. Quantitative PCR and nPCR revealed an equal number of positives in uncultured samples of sediments (3%) and bird tissues (40%), but these values tended to be higher after culture enrichment with the qPCR assay (10% and 80%, respectively). Associations between the presences of toxigenic C. botulinum C in the environment and in birds within the ecological conditions in wetlands could be studied further using the culture enrichment and qPCR techniques shown in the current study.
Clostridium botulinum is the spore-forming anaerobic bacterium responsible for botulism, a disease caused by the ingestion of a neurotoxin (BoNT). 12 The BoNT causes flaccid paralysis and death in human and animal populations. Seven bacteria subtypes have been classified by their neurotoxin production from A to G, 6 with type C being more frequently responsible for die-offs in waterfowl. Avian botulism has killed millions of birds around the world, and is considered to be the most important disease in waterfowl in terms of mortality. 8 In a wetland area (Mancha Húmeda) in central Spain, 13 recorded botulism outbreaks have occurred since 1978, and have affected more than 20,000 birds from several species, including the globally endangered white-headed duck (Oxyura leucocephala). 10
The use of a quantitative real-time polymerase chain reaction (qPCR) technique previously described for the detection of the gene encoding for the C. botulinum C toxin (BoNTC) 9 was applied to sediment and avian samples and evaluated in the current study. The aim of the present study is to present a quantitative technique that could help in future studies to consider the ecological conditions associated with the presence of C. botulinum in the environment and consequently aid risk assessments regarding the possibility of avian botulism outbreaks. Also, sensitivity results for the qPCR were compared with those of other PCR techniques previously described.
Sediments and samples from birds were collected at 4 sites (the Alcázar de San Juan Lagoons, the Vega del Jabalón Reservoir, the Navaseca Lagoon, and the Tablas de Daimiel National Park), which are all located in the wetland area known as the “Mancha Húmeda,” in central Spain. All samples were collected between 2005 and 2008. Waterbirds were collected dead or moribund during suspected botulism outbreaks by the Wildlife Rehabilitation Center of El Chaparrillo (Ciudad Real, Spain). The birds (N = 21) included 9 mallards (Anas platyrhynchos), 1 common teal (Anas crecca), 1 northern shoveler (Anas clypeata), 2 common pochards (Aythya ferina), 5 common black-headed gulls (Larus ridibundus), 1 black-winged stilt (Himantopus himantopus), 1 Eurasian coot (Fulica atra), and 1 cattle egret (Bubulcus ibis). Sediment samples (n = 30; 50–100 g) were collected from the upper 5-cm layer and immediately frozen at −30°C until DNA extraction and culture enrichment was undertaken. Samples from birds were collected at necropsy, were stored at −30°C until analysis, and included (n = 30) 10 gizzards, 10 small intestines, and 10 cecum contents selected from the 21 sampled birds. The animal tissues were sliced (individually) while frozen and then thawed and homogenized in sterile Petri dishes aseptically, taking 2 aliquots for direct DNA extraction and culture enrichment.
DNA extraction was performed using a DNA isolation kit for soil a and tissues, b following the manufacturers’ recommendations. To compare this direct extraction with results obtained after culture enrichment, sediment and samples from birds (1 g) were also cultivated in a cooked meat broth (9 ml) supplemented with glucose, hemin, and vitamin K c for 4 days at 40°C in anaerobic conditions using an anaerobe container. d A broth pellet was obtained by centrifuging 1 ml of the vortexed culture in a sterile microtube at 12,000 × g for 5 min. Post-culture DNA was then extracted using the same kits as mentioned above.
A qPCR previously described was then used for the C. botulinum C gene detection 9 using the DNA extracted from the samples (Table 1). DNA from 2 toxigenic C. botulinum C strains (C-468 f and CIP 104983 g ) was used as positive controls. A nontoxigenic strain of C. botulinum C (NCTC 10914) h was used to assess the specificity of the technique. Strains CIP 104983 and NCTC 10914 were cultured as mentioned above, and DNA extraction was undertaken using the same kit as mentioned above for tissues. The DNA extracted from all of the reference strains was measured on a spectrophotometer. i The limit of detection for the qPCR was calculated using the quantification cycle (Cq) 2 values observed when using serial 10-fold dilutions of the DNA extractions from both reference strains (C-468 and CIP 104983). All reference strains were analyzed in triplicate, and the results were considered positive if at least 2 of the 3 replicates were positive.
Primers and probes used in the current study.*
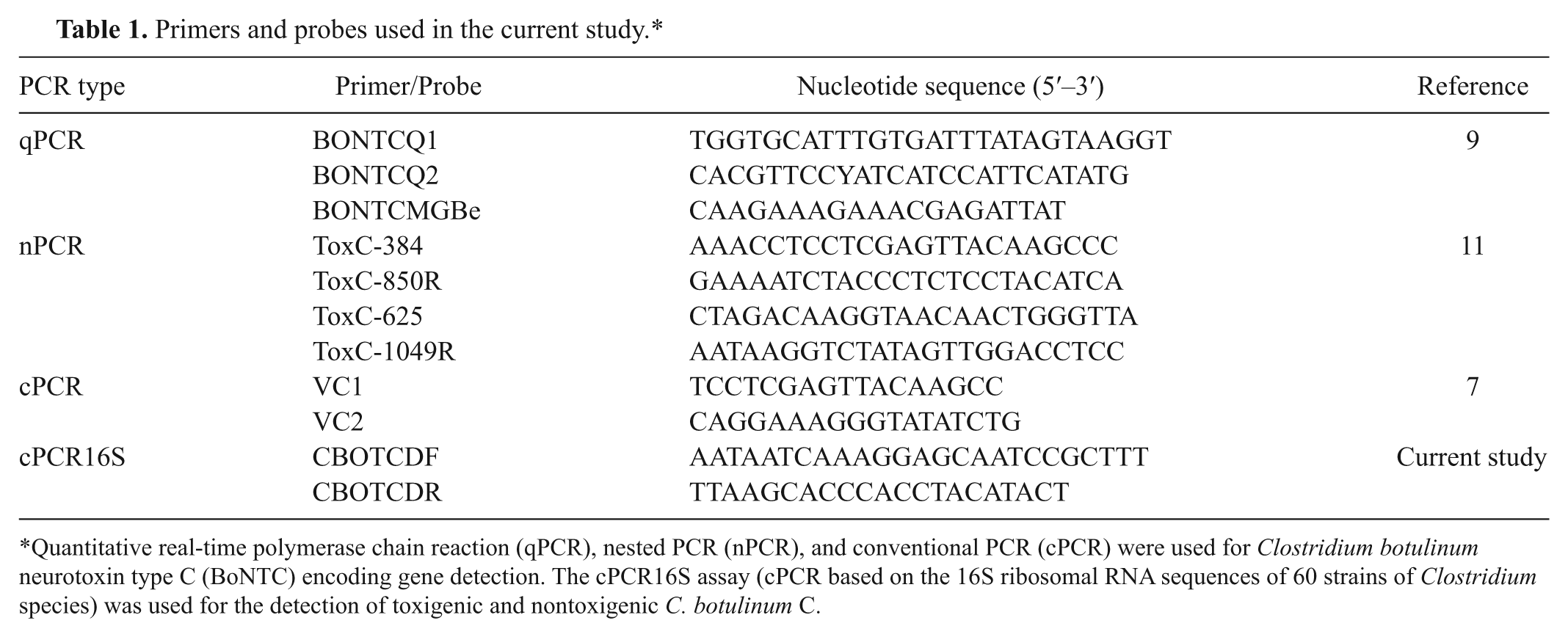

Quantitative real-time polymerase chain reaction (qPCR), nested PCR (nPCR), and conventional PCR (cPCR) were used for Clostridium botulinum neurotoxin type C (BoNTC) encoding gene detection. The cPCR16S assay (cPCR based on the 16S ribosomal RNA sequences of 60 strains of Clostridium species) was used for the detection of toxigenic and nontoxigenic C. botulinum C.
Results for the qPCR were compared with 2 other PCR techniques previously described in the literature, a nested (n)PCR 11 and a conventional (c)PCR, 7 using the reference strains and environmental samples. Proprietary reagents were used for the nPCR and cPCR. j All 3 PCR techniques are based on the amplification of different fragments of the BoNTC encoding gene (Table 1). Limits of detection for the 3 PCR assays were calculated using the reference strains mentioned above. DNA from sediment samples was also analyzed using the 3 PCR assays at 1:1, 1:10, and 1:100 dilutions (in order to reduce the effect of possible inhibitors) in nuclease-free water. k
In addition, a cPCR based on the 16S ribosomal (r)RNA sequences (cPCR16S) of 60 strains of Clostridium species (published in GenBank, http://www.ncbi.nlm.nih.gov) was also used to detect nontoxigenic C. botulinum when the other 3 PCR techniques all gave negative results (Table 1). These sequences were aligned, l and a specific cluster was selected, which contained C. botulinum C, C. botulinum D, and other uncultivable Clostridium species. A set of primers (Table 1) that amplified a 373-bp fragment was selected. m The cPCR16S assay was performed using the following parameters: 50–100 ng/µl DNA extraction, 0.2 µM primer concentration, 1.5 mM MgCl2, 200 µM deoxyribonucleotide triphosphate mix, and 1.25 U/µl of Taq polymerase. j This PCR reaction was performed in a thermocycler n under the following conditions: 1 denaturation cycle for 3 min at 94°C; followed by 35 denaturation cycles for 30 sec at 94°C, annealing for 30 sec at 59°C, and extension for 30 sec at 72°C; and a final extension for 5 min at 72°C. Finally, 10 µl from the amplification reaction was resolved by electrophoresis in a 2% agarose gel. DNA-amplified bands were visualized by ultraviolet transillumination o after staining with ethidium bromide. The limit of detection for the cPCR16S assay was obtained using toxigenic and nontoxigenic reference strains as described above for the other PCR assays. To assess the specificity of all the PCR techniques developed in the current study, several strains from the Centre de Recerca en Sanitat Animal (Barcelona, Spain) collection were analyzed, some of which were reference strains (including species of Clostridium), and others were isolated from the digestive tract of poultry (Table 2).
Strains used for assessing the specificity of polymerase chain reaction assays and their obtained results.*
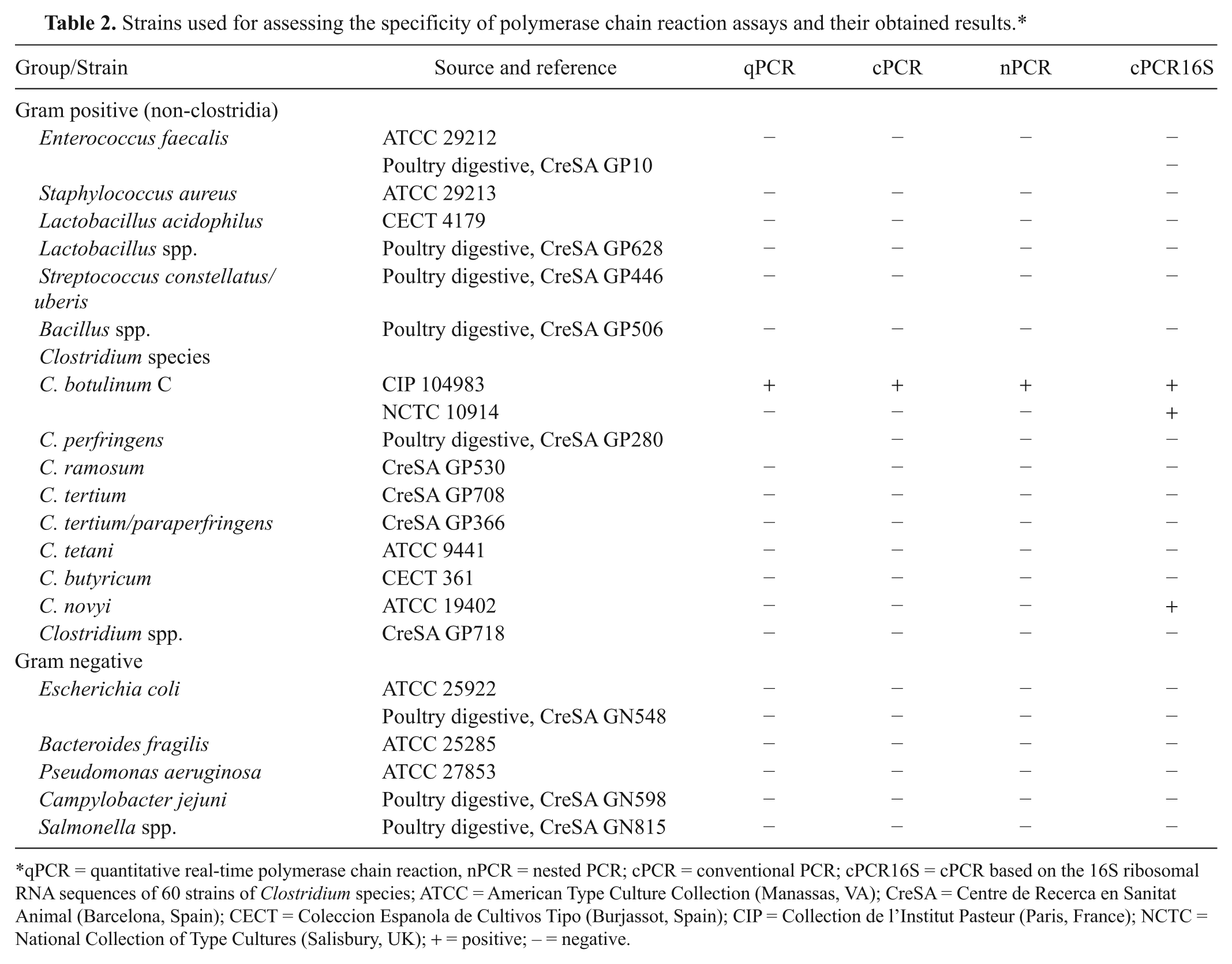
qPCR = quantitative real-time polymerase chain reaction, nPCR = nested PCR; cPCR = conventional PCR; cPCR16S = cPCR based on the 16S ribosomal RNA sequences of 60 strains of Clostridium species; ATCC = American Type Culture Collection (Manassas, VA); CreSA = Centre de Recerca en Sanitat Animal (Barcelona, Spain); CECT = Coleccion Espanola de Cultivos Tipo (Burjassot, Spain); CIP = Collection de l’Institut Pasteur (Paris, France); NCTC = National Collection of Type Cultures (Salisbury, UK); + = positive; − = negative.
For the qPCR and nPCR techniques, the limit of detection per reaction was <0.5 pg of DNA for the C-468 reference strain, and lower than for the cPCR (Table 3). The sensitivity obtained with the CIP 104983 reference strain was also similar for the qPCR and nPCR, and again better than when using the cPCR. The sensitivity obtained by a previous study 7 with cPCR was 1 pg of DNA. In terms of vegetative cells or spores, limits of detection for different PCR techniques can be as low as 10–50 cells/spores in 0.25–25 g of environmental sample, especially after culture enrichment as observed in the present study.3-5,11 Enrichment culture in clinical samples is routinely undertaken before the mouse bioassay is used for laboratory confirmation of botulism. 5 Moreover, enrichment culture has previously increased the sensitivity of an enzyme-linked immunosorbent assay developed to detect neurotoxins of C. botulinum types C and D in cases of bovine botulism. 1
Limits of detection for all polymerase chain reaction techniques used to detect toxigenic Clostridium botulinum type C.*
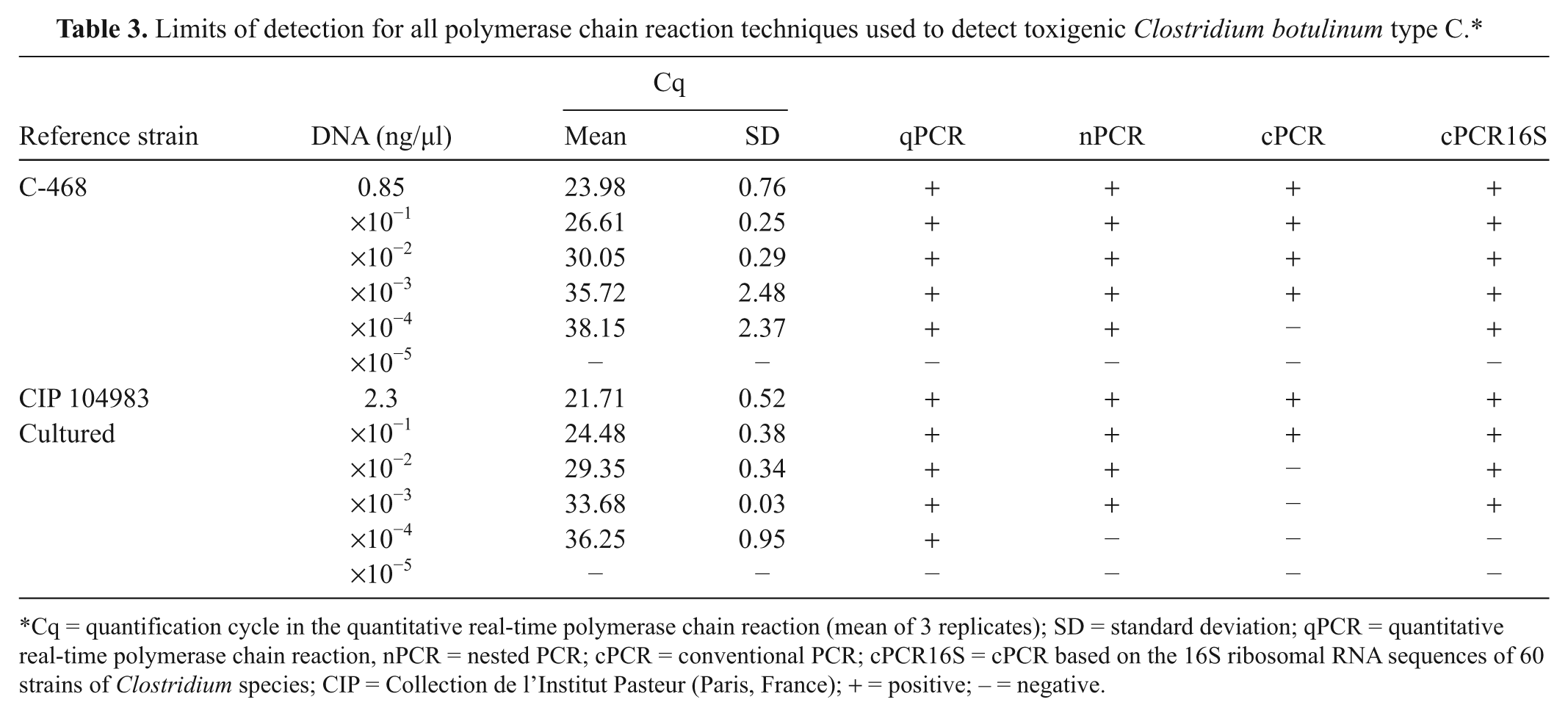
Cq = quantification cycle in the quantitative real-time polymerase chain reaction (mean of 3 replicates); SD = standard deviation; qPCR = quantitative real-time polymerase chain reaction, nPCR = nested PCR; cPCR = conventional PCR; cPCR16S = cPCR based on the 16S ribosomal RNA sequences of 60 strains of Clostridium species; CIP = Collection de l’Institut Pasteur (Paris, France); + = positive; − = negative.
Similar Cq values were obtained with the CIP 104983 reference strain and the C-468 strain (Table 3). The slope of calibration curves obtained with these 2 strains was also similar (Fig. 1). The nontoxigenic reference strain NCTC 10914 C. botulinum C was analyzed and tested negative when using the 3 toxigenic PCR techniques tested herein. However, it was positive when using the cPCR16S assay. Among all the Clostridium strains and other bacteria examined in the current study, only C. botulinum C and C. novyi strains gave positive results with the cPCR16S assay (Table 2).
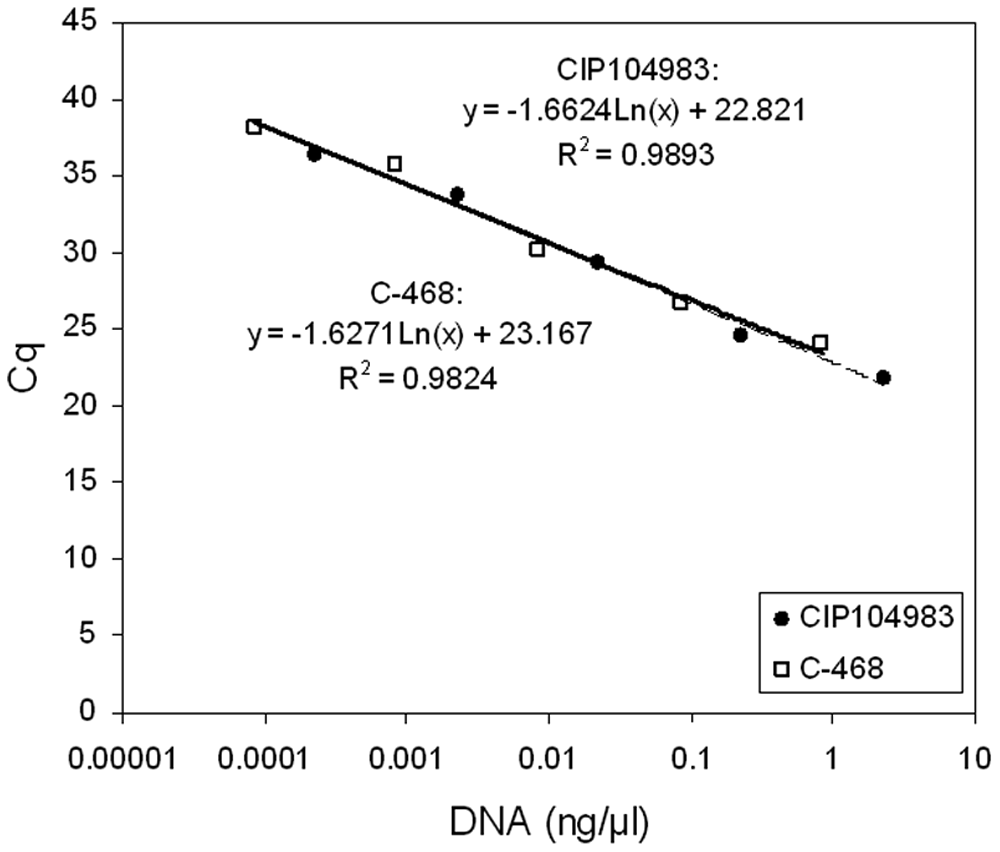
Quantitative real-time polymerase chain reaction calibration curves on semi-logarithmic scale obtained with DNA extracted from 2 toxigenic strains of Clostridium botulinum type C1 (CIP 104983 and C-468). Cq = quantification cycle.
Enrichment of environmental samples resulted in the detection of more positives than when samples were directly extracted, except when using nPCR (Table 4). This may be (at least in part) because the DNA extraction procedure used herein was only effective for vegetative cells of C. botulinum, and not for spores. 13 Therefore, some samples containing spores would only give positive results after culture enrichment. Although the objective of the present study was not to determine the prevalence of C. botulinum in these samples (a far larger sample set will be used for this purpose), if all the techniques used were considered, along with both DNA extractions, the prevalence of toxigenic C. botulinum C in sediments analyzed was 10%, while in animal samples the prevalence was 80% (Table 4). In sediment samples, it should be noted that 2 out of the 3 qPCR positives (after culture enrichment) were only positive at the 1:10 extracted DNA dilution. Hence, an as yet unidentified PCR inhibition effect (perhaps from a soil chemical component) may be present. None of the sediments was positive in the 1:100 dilution. For birds, when considering individuals, the total prevalence detected was actually 100% (i.e., 21 out of 21 birds tested were positive in at least one sample type tested).
Positive rate (number of positive/total number analyzed) obtained using 3 polymerase chain reaction techniques for detection of the neurotoxin of the Clostridium botulinum type C encoding gene in sediment and animal tissue samples before (direct) and after enrichment culture (cultured).*
qPCR = quantitative real-time polymerase chain reaction, nPCR = nested PCR; cPCR = conventional PCR.
In conclusion, further studies regarding possible associations between the presence or absence of toxigenic and nontoxigenic C. botulinum C in the environment and in birds, as well as the prevailing ecological conditions within wetlands, could be undertaken using the culture enrichment and subsequent qPCR and cPCR16S techniques shown in the current study. In addition, more research must be conducted to obtain DNA extraction methods for samples that do not involve a culture enrichment step. 11 Such methods will allow diagnosticians to establish the quantitative association between the number of C. botulinum cells or spores in samples and any ecological factors found in wetlands or linked to particular birds.
Footnotes
Acknowledgements
The authors thank Ramón González and José Barcenilla from the Institute of Industrial Fermentation (CSIC) for their help with laboratory analysis. The authors also appreciate the cooperation of staff from the Tablas de Daimiel National Park, of the Environmental Agents from Castilla-La Mancha (JCCM), and of personnel from the Wildlife Rehabilitation Centre of El Chaparrillo for their help with sample collection.
a.
PowerSoil™ DNA Isolation kit, MO BIO Laboratories Inc., Carlsbad, CA.
b.
DNeasy® Blood and Tissues kit, Qiagen GmBH, Hilden, Germany.
c.
BBL™ Cooked Meat Medium, BD SA, Madrid, Spain.
d.
GasPak™ EZ anaerobe system, BD SA, Madrid, Spain.
e.
TaqMan MBG, Applied Biosystems, Foster City, CA.
f.
National Wildlife Health Center, Madison, WI.
g.
Collection de l’Institute Pasteur, Paris, France.
h.
National Collection of Type Cultures, Salisbury, United Kingdom.
i.
NanoDrop® ND-1000, Thermo Fisher Scientific Inc., Waltham, MA.
j.
Biotools B&M Labs SA, Madrid, Spain.
k.
Ultrapure distilled water, Invitrogen Corp., Paisley, United Kingdom.
l.
BioEdit Sequence Alignment Editor 7.0, Ibis Therapeutics, Carlsbad, CA.
m.
Oligo Explorer™ 1.2, Gene Link, Hawthorne, NY.
n.
GeneAmp® PCR System 2700, Applied Biosystems, Foster City, CA.
o.
Gel Doc 2000, Bio-Rad Laboratories SA, Madrid, Spain.
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
This study was funded using a grant (99/2003) from the Autonomous Organisation of National Parks (Spanish Ministry of the Environment, MARM). Dolors Vidal was supported with a JAE DOC contract of the Spanish National Research Council (CSIC), and Mark A. Taggart was supported with a Juan de la Cierva Contract of the Spanish Ministry of Science and Innovation (MICINN).
