Abstract
A real-time polymerase chain reaction assay coupled with high resolution melting curve analysis (PCR-HRM) was developed for identifying and distinguishing Mycoplasma species commonly isolated from ruminant, avian, and canine samples. The real-time PCR used 1 set of universal primers specific for the spacer region between the 16S ribosomal RNA and the 23S ribosomal RNA genes; the melting curve analysis of the PCR product used a high-resolution melt fluorescent dye. The real-time PCR-HRM assay was able to distinguish M. arginini, M. bovigenitalium, M. bovis, M. bovirhinis, M. canadense, M. cynos, M. spumans, M. iowae, M. meleagridis, and M. agalactiae reference strains. The real-time PCR-HRM assay developed was evaluated by testing field isolates of M. bovis, M. arginini, M. bovirhinis, M. bovigenitalium, M. iowae, and M. spumans with results consistent with those of the fluorescent antibody test.
Many species of Mycoplasma are clinically important to animals, and are therefore of great concern in veterinary medicine.7,9 Culture isolation of Mycoplasma species is essential for a definitive identification of the organism.6,12 However, Mycoplasma species can be fastidious to grow and difficult to identify phenotypically. 9 Conventional methods of Mycoplasma species identification are based on serological tests, such as the indirect fluorescent antibody test (IFAT).3,6,7,9 The IFAT is based on staining of mycoplasma colonies on semi-solid media with fluorescein-conjugated antibodies. 13 Mycoplasma-specific antibodies are not commercially available and are laborious to produce and standardize. Furthermore, the IFAT is time-consuming and requires extensive interpretation by a trained professional. Recently, 16S ribosomal RNA (rRNA) gene sequence analysis has been applied for Mycoplasma speciation,7,9 but the test is expensive and time consuming.3,6,7,9 Although high-resolution melting curve analysis has recently been used for the strain typing of Mycoplasma gallisepticum 3 and Mycoplasma synoviae strains, 6 it has not been described for Mycoplasma speciation. Compared to the low-resolution double-stranded DNA (dsDNA) dyes such as SYBR Green I, high-resolution dsDNA dyes enable the melting curve analysis in much higher resolution and therefore is able to detect subtle changes in the melting properties resulting in more precise assessment of sequence variations.2,4,10,12 The aim of the current study was to develop a cost-effective and rapid assay to identify common veterinary Mycoplasma species employing real-time polymerase chain reaction (PCR) followed by high-resolution melting curve analysis (PCR-HRM) using 1 set of universal primers specific for the spacer region between the 16S and 23S rRNA genes.
Mycoplasma reference strains and field isolates used in the present study are listed in Table 1. All reference strains were cultured in Hayflick broth as previously described. 13 Clinical isolates were grown in Hayflick broth or Hayflick media plates with 0.8% agar. a Pure cultures from clinical specimens were acquired by re-culturing in Hayflick media plates until single colonies were obtained. Mycoplasma species were identified by IFAT. 11
Mycoplasma reference strains and their melting peaks determined by real-time polymerase chain reaction coupled with high-resolution melting curve analysis.
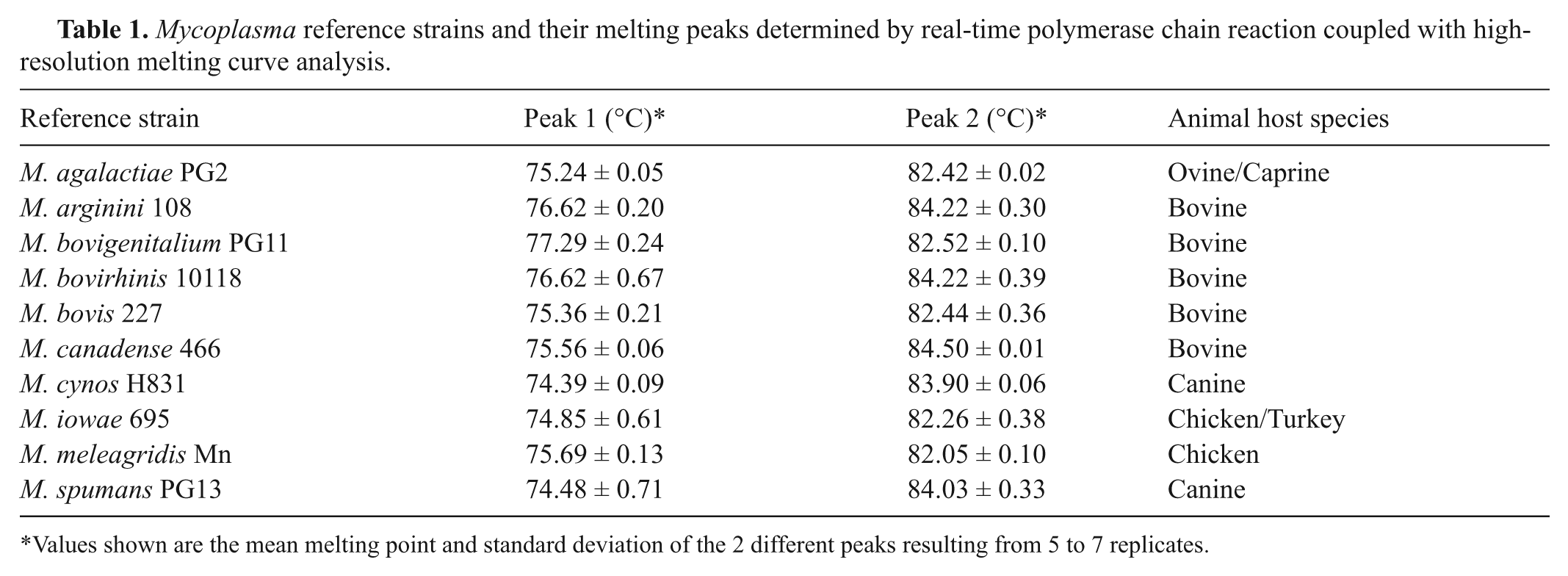
Values shown are the mean melting point and standard deviation of the 2 different peaks resulting from 5 to 7 replicates.
Mycoplasma broth culture (200 µl) or colonies scraped from the media plate and resuspended in 200 µl of phosphate buffered saline were used for genomic DNA extraction using an automated system b or manually. c DNA from lung and milk specimens was extracted as described previously. 1
Polymerase chain reaction was performed using a real-time PCR instrument d with a high resolution melting analysis kit. e Universal primers, F1(5′-ACACCATGGGAGY TGGTAAT-3′) and R1(5′-CTCCWTCGACTTYCAGACCCAAGGCAT-3′), specific for the spacer region between the 16S and 23S rRNA genes, were used to generate amplicons (350–490 bp). 4 The PCR was conducted in an 18-µl reaction volume containing the HRM master mix (in 1 time concentration) containing high resolution dsDNA dye, e 0.5 µM of each primer, 3.5 mM of MgCl2, and 2 µl of genomic DNA. The PCR thermocycling conditions were 95°C for 10 min followed by 35 cycles of 95°C for 10 sec, 60°C for 30 sec, then 72°C for 30 sec.
Following the completion of PCR, products were heated to 95°C for 1 min at 4.8°C/sec, and cooled to 60°C for 1 min at 2.5°C/sec. High-resolution melting curve analysis data was collected by increasing the temperature from 60°C to 95°C at 0.01°C/sec with 40 acquisitions per degree Celsius. Optical measurements were done in the 6-carboxyfluorescein (FAM) channel (excitation at 465 and detection at 510 nm). High-resolution melting curve analysis was performed using the HRM software that came with the real-time PCR instrument.d,f Five to 7 replicates of the real-time PCR-HRM assay were performed simultaneously for each Mycoplasma reference strain, clinical isolate, and negative control tested. Mycoplasma species differentiation was based on HRM profiles and/or normalized HRM profiles, which consists of melting temperatures and the overall shape of the melting curves (Fig. 1).
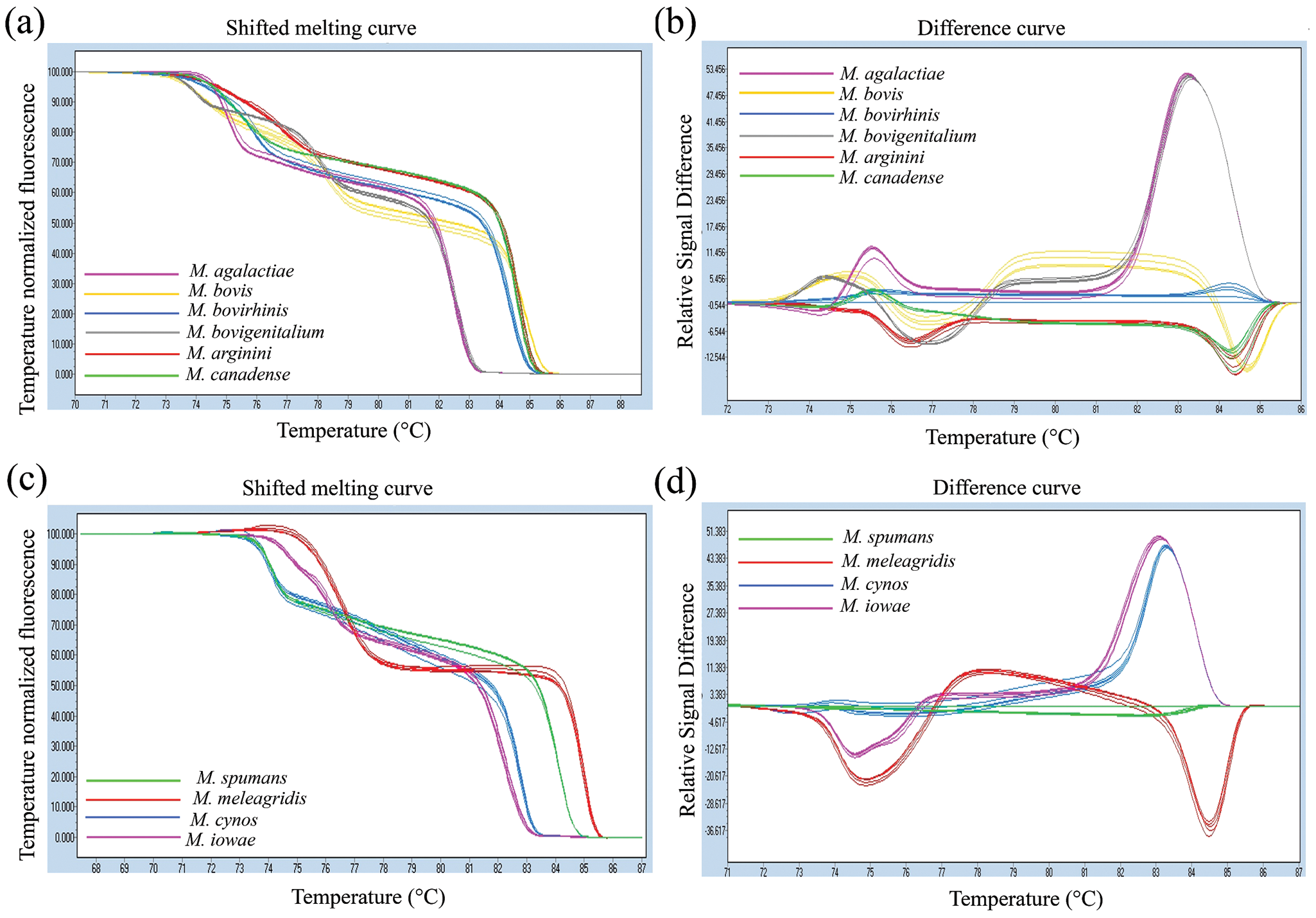
All 10 Mycoplasma reference strains including M. agalactiae, M. arginini, M. bovigenitalium, M. bovirhinis, M. bovis, M. canadense, M. cynos, M. iowae, M. meleagridis, and M. spumans produced clearly distinct HRM profiles and normalized HRM profiles (Fig. 1). The PCR amplicons for each Mycoplasma species tested generated 2 melting peaks, the first peak ranging from 73°C to 77°C and the second peak ranging from 82°C to 85°C (Table 1). The observed peaks were in accordance with the in-silico analysis predictions using a melting curve analysis program. g Mycoplasma arginini and M. bovirhinis produced very similar melting peak temperatures. However, these organisms did generate distinctive HRM profiles (Fig. 1a) and normalized HRM profiles (Fig. 1b), making the differentiation of each organism quite clear.
Mycoplasma species tested in the present study were chosen based on the authors’ routine IFAT panel so as to develop an alternative assay to the IFAT. Although not significant pathogens, M. arginini and M. bovirhinis are often isolated from clinical samples and were therefore included in the real-time PCR-HRM assays. The HRM analysis software was able to differentiate 6 species at a time, therefore Mycoplasma species were differentiated in 2 subsets, 1 subset for bovine, ovine, and caprine Mycoplasma species and 1 subset for canine, turkey, and chicken Mycoplasma species. Both subsets were set up in the same real-time PCR-HRM run.
The real-time PCR-HRM assay developed was evaluated by testing field isolates of M. bovis, M. arginini, M. bovirhinis, M. bovigenitalium, M. iowae, and M. spumans. The real-time PCR-HRM results were consistent with those of the IFAT (Table 2). Field isolates of M. agalactiae, M. canadense, M. cynos, and M. meleagridis were not tested because clinical samples were not available.
Identification of Mycoplasma clinical isolates by real-time polymerase chain reaction coupled with high-resolution melting (HRM) curve analysis.
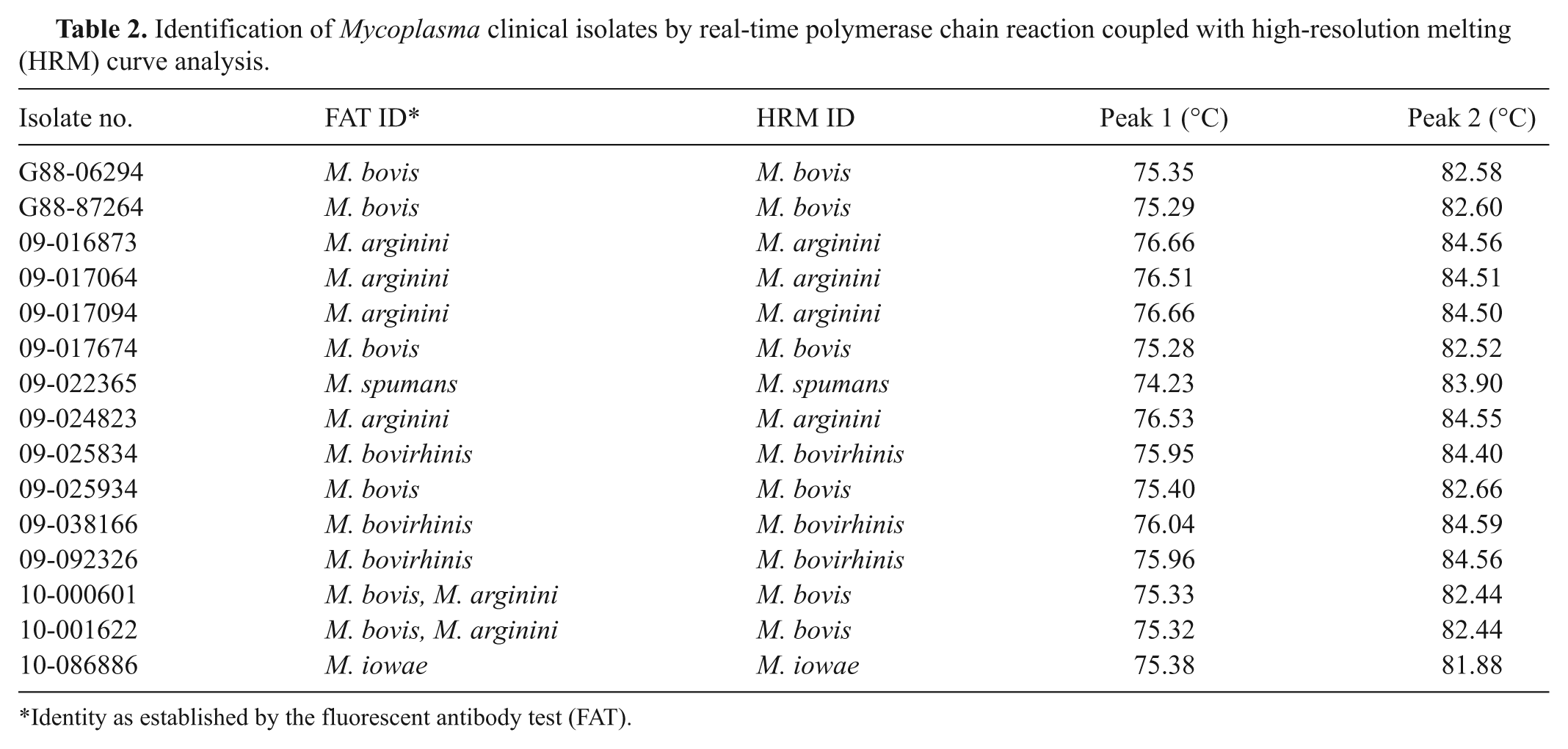
Identity as established by the fluorescent antibody test (FAT).
In order to evaluate if the real-time PCR-HRM assay works on DNA of mixed cultures and DNA extracted directly from clinical samples, mixed cultures of M. bovis and M. arginini, and clinical samples infected with M. bovis, were selected for testing, as M. bovis is one of the most commonly isolated Mycoplasma species. Only M. bovis was identified from the mixed cultures, and the assay was not able to detect M. bovis from the DNA extracted from lung and milk specimens (data not shown). It was determined by IFAT analysis that M. bovis was the organism present in higher concentration compared to M. arginini, which may have overshadowed the detection of M. arginini (Table 2). Similarly, the background DNA in the clinical samples may have interfered with the assay. No further experiments were carried out to test mixed cultures or samples consisting of other Mycoplasma species.
The approach using universal primers to amplify 16S and 23S rRNA spacer region coupled with amplicon melting curve analysis has been previously described for the differentiation of cell culture Mycoplasma contaminants, however, with low-resolution dsDNA dye SYBR Green I, which generated lower resolution melting curve profiles,5,8 and lacked a precise differentiation for similar PCR products. The present study investigates HRM analysis with a high-resolution dsDNA dye for the speciation of Mycoplasma species. In contrast to low-resolution dsDNA dye such as SYBR Green I, high-resolution dsDNA dye does not interference with polymerase and primer binding and can therefore be used at a higher concentration to saturate the dsDNA to avoid dye shifting during the melting analysis. As a result, with a high-resolution dsDNA dye, the dsDNA melting profile has higher resolution, which gives more accurate sequence differentiation.2,4,7,9
The spacer region between the 16S and 23S rRNA genes contains stretches of high sequence conservation; however, the size of the sequence varies for different Mycoplasma species. 5 Sequence variation at the spacer region between the 16S and 23S rRNA genes allowed for a very clear and reproducible HRM curve analysis differentiation among all 10 reference Mycoplasma species that were analyzed in the present study (Fig. 1). Another conserved gene, the 16S rRNA gene, was also tested but failed to generate good discriminatory results (data not shown). All Mycoplasma reference strains tested generated HRM profiles with 2 melting peaks. A major peak is observed at the temperature of 82–84°C and a minor peak at the temperature of 74–77°C (Fig. 1b, 1d). Some Mycoplasma species produced similar melting peaks. However, the HRM profile and/or normalized HRM curve gave a clear distinction. For instance, M. bovirhinis and M. arginini produced very similar melting temperatures. However, these 2 Mycoplasma species had distinct HRM profile and normalized HRM profile (Table 1; Fig. 1a, 1b).
Comparing to the current gold standard IFAT and 16S rRNA gene sequence analysis, the real-time PCR-HRM was less expensive and fast (1–2 hr) but was only able to identify pure cultures. The IFAT takes approximately 6 hr to identify 6–10 isolates and can work on colonies of mixed species, but it is technically demanding, and can have cross-reactions and autofluorescence (false positive). Sequencing of the 16S rRNA gene is expensive, takes 2–3 days to obtain a result, and only works with pure cultures.
In summary, the current study demonstrated that the real-time PCR-HRM technique was a cost effective and rapid assay for the differentiation of Mycoplasma species in pure cultures. For the identification of less commonly isolated Mycoplasma species not tested in the present study, the related reference strains would need to be tested to generate reference melting curve profiles as positive control.
Footnotes
Acknowledgements
The authors would like to thank Drs. Mark Yoshimasu and Victor Chong (Roche) for their technical support, and the Animal Health Laboratory technicians Rebeccah McDowall, Patricia Bell-Rogers, and Patricia McRaild for their technical assistance.
a.
Purified agar, Oxoid, Ottawa, Ontario, Canada.
b.
MagNA-Pure LC, Roche Diagnostics, Montreal, Quebec, Canada.
c.
DNeasy Blood and Tissue Kit, Qiagen Inc., Mississauga, Canada.
d.
LightCycler® 480, Roche Diagnostics, Montreal, Quebec, Canada.
e.
LightCycler® 480 High Resolution Melting Master kit containing ResoLight® dye, Roche Diagnostics, Montreal, Quebec, Canada.
f.
LightCycler® 480 Software (Version 1.5.0.39) Roche Diagnostics, Montreal, Quebec, Canada.
g.
The authors declared that they had no conflicts of interest with respect to their authorship or the publication of this article.
This research was financially supported by the Ontario Ministry of Agriculture, Food and Rural Affairs (OMAFRA) Animal Health Laboratory, Animal Health Strategic Investment (AHSI) program.
