Abstract
Alloherpesviruses affect freshwater and marine fish species. The aim of the current study was to characterize a novel alloherpesvirus in Atlantic cod (Gadus morhua). Samples were processed for histopathology, transmission electron microscopy (TEM), virus isolation, molecular characterization, and in situ hybridization (ISH). Histopathology revealed that the infection was restricted to the gills and that it induced cytomegaly in infected cells. By TEM, numerous viral particles with morphology compatible with a herpesvirus were observed inside the cytomegalic cells. To characterize this new agent, polymerase chain reaction amplified regions of the ATPase subunit of the terminase, and DNA polymerase genes were sequenced. Phylogenetic analysis revealed strongest similarity with alloherpesviruses belonging to the genus Ictalurivirus and Salmonivirus. The ISH showed specific labeling of nuclear inclusions in the cytomegalic cells. While virus isolation was unsuccessful, the results obtained through different diagnostic tests in the present study confirm the discovery of a new alloherpesvirus affecting Atlantic cod. The authors propose the formal species designation Gadid herpesvirus 1 (GaHV-1) to be considered for approval by the International Committee on the Taxonomy of Viruses.
Introduction
Information on viruses affecting fish lags behind knowledge of mammalian viruses. However, in recent years, some fish viruses have been intensively studied due to their involvement in significant economic loss in farmed fish.15,21
The order Herpesvirales includes double-stranded DNA viruses known to affect a large number of animal species.9,45 The order is divided into 3 families: Herpesviridae, which includes mammalian, avian, and reptilian viruses; Alloherpesviridae, which includes fish and amphibian viruses; and Malacoherpesviridae, which includes a single invertebrate virus affecting a range of bivalve species.9,45 Alloherpesviruses have been described in both freshwater and marine fish, although few of them have been formally characterized.15,45 Due to their negative impact on the aquaculture industry, some of the most well-known herpesviruses affect different species of salmonids, cyprinids, catfish, sturgeons, eels, and turbot. There is evidence that several alloherpesviruses induce neoplasia or proliferative epithelial lesions and long-term infections.18,45
Atlantic cod (Gadus morhua) is one of the best-known cold-water fish species in the world. Recent interest in farming this commercially important species has raised awareness about their diseases including viral infections. 38 The only known herpesvirus infection in cod was observed in Pacific cod (Gadus macrocephalus).30,31 In these reports, hypertrophied cells in the skin lesions and herpesvirus-like particles inside the giant cells were described. 30 In Atlantic cod, a previous study 24 reported hyperplastic lesions in the skin, in which the epidermis was 4 times thicker than normal. Adenovirus-like particles were found to be the cause. Iridovirus infections have also been reported in Atlantic cod. A previous study 25 reported the isolation of iridovirus from Atlantic cod affected with “ulcus syndrome,” an ulcerative skin disease of wild cod populations. It was subsequently demonstrated 3 that the iridovirus isolated from ulcus syndrome affected fish belongs to the genus Ranavirus (family Iridoviridae). An iridovirus is also the causative agent of viral erythrocytic necrosis, which is characterized by intracytoplasmic inclusions and nuclear degeneration in affected erythrocytes.20,36,49 More recently, nodavirus infection, producing viral encephalopathy and retinopathy (VER), has been reported in Atlantic cod. 42 Although VER affects mainly larval and juvenile stages, mortality in older fish has been reported. 38 Major viral diseases in salmonids such as infectious pancreatic necrosis 40 and viral hemorrhagic septicemia28,34,41 have also been reported in Atlantic cod.
In September 2009, formalin-fixed samples from Atlantic cod broodstock in Canada suffering from significant mortality were submitted to the Institute of Aquaculture (IoA; University of Stirling, Scotland, United Kingdom) for routine diagnostic examinations. Although the histopathological examination identified vibriosis as the main cause of the disease, it also revealed the presence of hypertrophied single cells in the gills (cytomegaly). The affected cells showed pronounced cytoplasmic and nuclear enlargement, findings not incompatible with certain viral agents, such as herpesviruses. Transmission electron microscopy (TEM) confirmed the presence of numerous viral particles inside the cytomegalic cells. Morphological features of the viral particles and their locations within affected cells indicated the possibility of herpesvirus infection. The present study describes the histopathology and TEM findings and presents the results of molecular analyses and in situ hybridization (ISH) to verify and characterize the suspected herpesvirus etiology.
Materials and methods
Sample collection
In September 2009, tissue samples from moribund Atlantic cod broodstock were collected in 10% buffered formalin and sent from Canada to the IoA for histopathological diagnosis. Together with severe multifocal caseonecrotic granulomatous lesions, typical of those associated with vibriosis in this species, histopathology also revealed hypertrophic single cells in the gills, mostly in the lamellar base. Additional samples from affected fish were subsequently collected for further testing. Four extra gill samples in 10% (w/v) formalin were submitted to the IoA for histopathological evaluation and TEM examination. Another 12 gill samples in 70% ethanol, plus a duplicate of the second set of formalin samples sent to the IoA, were sent to the Fish Health Laboratory of the University of California–Davis (FHL-UCD; Davis, California) for molecular virology testing and ISH.
During the same period, 16 moribund Atlantic cod were sent to the New Brunswick Research & Productivity Council (RPC; Fredericton, New Brunswick, Canada) for virological analysis using cell culture. Fish were necropsied, and organs (gills, pyloric caeca, spleen, and kidney [front, mid, and hind]) were combined into 5 pools. Pools consisted of the following: pool 1 (5 fish), pool 2 (2 fish), pools 3–5 (3 fish each). Intestine samples from the same fish were pooled separately. Pooled tissues were then frozen at −80°C until analysis by cell culture.
Cell culture
In January 2010, pooled tissue samples were thawed, macerated, diluted 1:4 (w/v) using Hanks balanced salt solution (HBSS), and further macerated at high speed for 120 sec. Homogenates were then diluted 10-fold with HBSS and centrifuged at 2,000 × g at 4ºC for 15 min. Supernatants were filtered through a 0.45-µm filter, and 0.1 ml from each pool was plated in duplicate on each type of cell line. Negative controls of HBSS were run for each cell line. The cell lines used were Atlantic salmon kidney (ASK), Chinook salmon embryo (CHSE; Oncorhynchus tshawytscha), epithelioma papulosum cyprini (EPC), and rainbow trout gonad (RTG-2; Oncorhynchus mykiss). Following inoculation, plates were incubated at 15ºC for 4–6 weeks and observed for cytopathic effect (CPE). All cultures showing CPE were then subjected to a second passage on the same cell lines. DNA extracted from aliquots of the first passage was sent to FHL-UCD for polymerase chain reaction (PCR) testing.
Histopathology
Gills, visceral organs, skin, and muscle samples stored in 10% (w/v) formalin were received at the IoA. The samples were routinely processed into paraffin wax blocks, sectioned (3–5 µm), and stained with hematoxylin and eosin.
Transmission electron microscopy
Gill samples were immersed in 2.5% glutaraldehyde in 100 mM cacodylate buffer (v/v) for 2–4 hr at 4°C. Samples were then kept in 100 mM cacodylate buffer with sucrose overnight at 4°C to remove excess fixative. Samples were post-fixed for 1 hr with 1% osmium in the same buffer, washed 3 times in distilled water for 10 min, and stained with 2% (w/v) uranyl acetate in 30% (v/v) acetone for 1 hr in the dark. Samples were then dehydrated through a series of acetone concentrations: 60% acetone for 30 min, 90% acetone for 30 min, 100% acetone for 30 min, and an additional 100% acetone bath of 1 hr. Samples were infiltrated first in agar low viscosity resin (ALVR) with acetone (1:1) for 1 hr and then twice in 100% ALVR for 1 hr each. Samples were embedded with new ALVR in green block molds and allowed to polymerize at 60°C for 24 hr. Sections 90-nm thick were cut, stained for 4 min with 4% uranyl acetate in 50% ethanol and for 7 min with Reynolds lead citrate, and then viewed with a transmission electron microscope. a
DNA extraction, polymerase chain reaction amplification, cloning, and sequencing
DNA was extracted from ethanol-preserved, formalin-fixed gill samples as well as from cell culture samples using a commercial kit b as per the manufacturer’s recommendations. DNA samples were stored at 4°C until used for PCR. The ethanol-preserved samples were subjected to a previously described degenerate PCR protocol 16 that targets a portion of the viral DNA-dependent DNA polymerase gene (hereafter, DNA polymerase). Using the aforementioned PCR protocol, a second primer pair designed to amplify a portion of the first exon of the ATPase subunit of the terminase gene (hereafter, the terminase) in alloherpesviruses was employed (primers: TermExon1F 5′-CAGGTBGARCTMATGMGRGGGTTTT T-3′ and TermExon1R 5′-CATRATKKYDGTYTTVCCRCA YTGTC-3′). Using the DNA polymerase sequence generated (described below), a new specific primer pair was developed to screen the ethanol-preserved, formalin-fixed samples and tissue culture samples, using the aforementioned PCR degenerate protocol.
Amplicons of the expected molecular size were excised from agarose gels and purified. b Purified DNA fragments were then cloned c according to manufacturer’s instructions. Transformed bacterial colonies were screened by PCR using the M13 primers, and plasmids containing the desired gene insert were purified. b Purified plasmids were sequenced with M13 primers on an automated sequencer. d
BLASTP, molecular dataset, sequence alignment, and phylogenetic analysis
Following cloning, sequencing, and primer sequence removal, general BLASTP searches (http://www.ncbi.nlm.nih.gov/blast/Blast.cgi) using the amino acid sequences deduced from the partial DNA polymerase and terminase genes were performed to acquire appropriate outgroups for phylogenetic analysis. The deduced amino acid sequences were aligned using Mafft 5.8 26 followed by minor manual adjustments in ClustalW. 43 The E-INS-I alignment strategy was used with the following parameters: scoring matrix (BLOSUM62), gap open penalty (1.53), and offset value (0).
To assess gene concordance, Bayesian analyses were performed independently for each gene. Phylogenetic trees were constructed using Bayesian analyses (MrBayes version 3.1.2). 22 A mixed prior was used on amino acid models and default priors for topology (uniform) and branch lengths (Exp and 10). The Markov chain was run for a maximum of 10 million generations, with a stopping rule implemented so that the analysis would halt when the average deviation of the split frequencies was less than 0.001%. Four independent analyses were conducted, each with 1 cold and 3 heated chains with the default heating parameter (temperature = 0.2). Every 50 generations were sampled and the first 25% of mcmc samples discarded as burn-in.
DNA probes for in situ hybridization
The probe was labeled with digoxigenin (DIG) using a PCR DIG probe synthesis kit. e The aforementioned plasmid containing the partial viral DNA polymerase was used as template (approximately 50 pg) in the PCR for probe generation, using ACHV-1F (5′-TGGGCAACCGTTCCCCTGG-3′) and ACHV-1R (5′-TGAGCCCTGAGATCCAGGT-3′) primers. The PCR for probe preparation produced a 145-bp amplicon, and conditions were as previously described for the PCR screening protocol. DIG labeling of the probes was conducted according to the manufacturer’s instructions. Following PCR, subsamples of the labeled probe were run on a gel to confirm successful tagging of the DIG. Prior to use in ISH, the probes were denatured for 5 min at 95°C, immediately placed on ice, and stored at −20°C until used.
In situ hybridization using DNA probes
Gill sections from the first reported case sent to the IoA and from the formalin-fixed samples forwarded to the FHL-UCD were used for non-radioactive ISH. Fish tissues infected with Epizootic epitheliotropic disease virus (EEDV), an alloherpesvirus from lake trout (Salvelinus namaycush), were used as a negative control. Samples were processed to paraffin wax blocks, cut to 5-µm sections, and placed on SuperFrost plus slides. Except for modifications on the probe labeling, as described above, hybridization temperature (52°C), and signal development at 4°C overnight with the substrate solution, all steps in the ISH assay were followed according to a previous protocol established for fish pathogens at the FHL-UCD. 1
Results
Histopathology
The first samples received at the IoA had caseonecrotic granulomatous lesions in different organs. The heart was the most seriously affected organ, but granulomas were also present in skin and kidney. Histopathology also revealed the presence of cytomegalic cells in the gills, showing nuclear and cytoplasmic enlargement and nuclear inclusions (Fig. 1A). Cytomegalic cells were not present in all the examined filaments, but affected filaments showed hypertrophic cells in high numbers both at the lamellar base and along the epithelium. Multifocal hyperplasia and fusion of the lamellar epithelium were also observed, although these changes were not associated with the cytomegalic cells. Evidence of similar cytomegalic cells was not found in any of the other sampled organs.
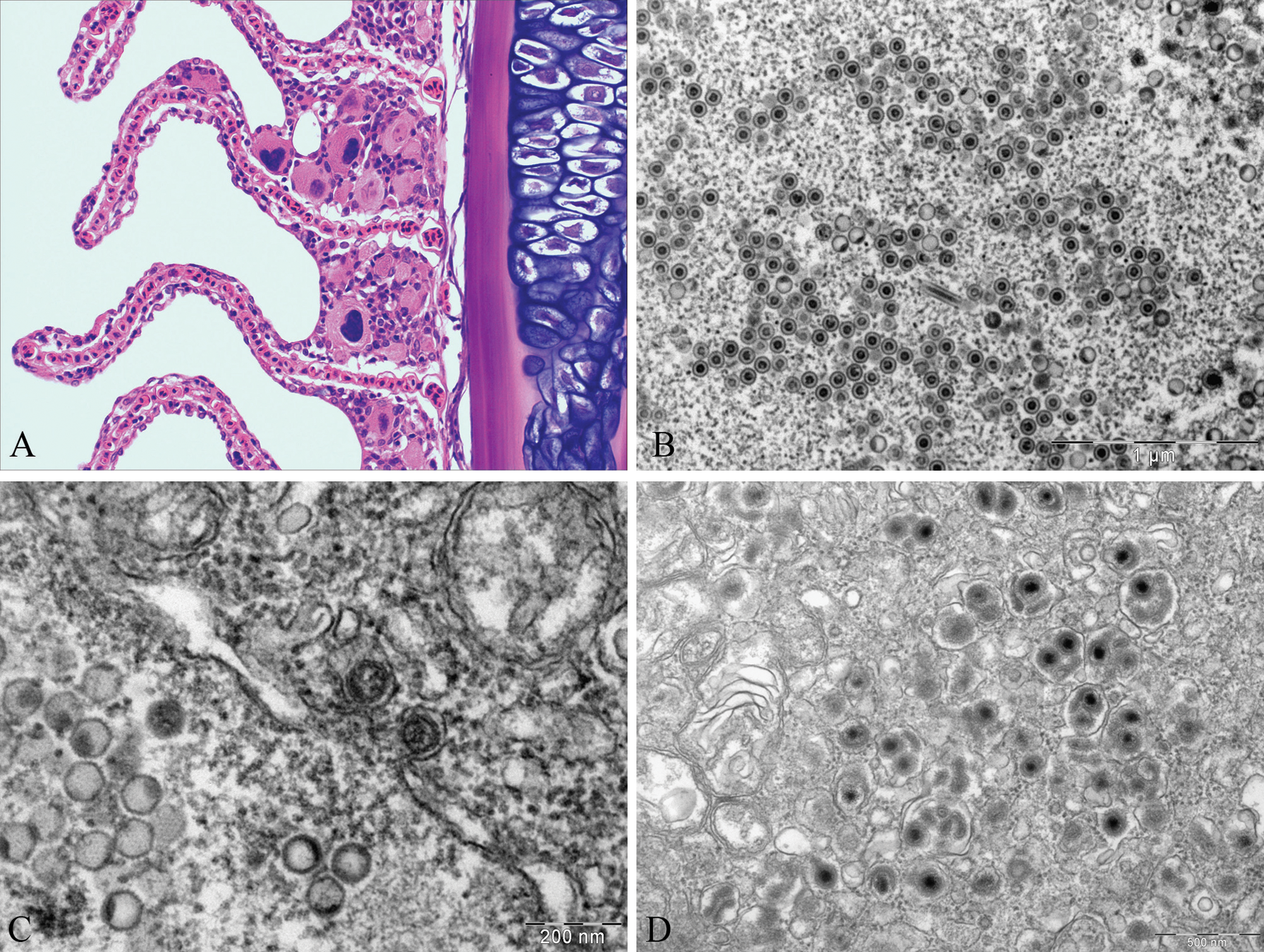
Atlantic cod (Gadus morhua).
Transmission electron microscopy
Transmission electron microscopy focused on the cytomegalic cells in the gills. Affected cells showed hypertrophy of the cytoplasm and the nucleus, and both contained viral particles. Viral particles were restricted to the nucleus in some cells, while in others the particles were found in both nucleus and cytoplasm. Intranuclear particles had an icosahedral shape and an average size of 75 nm (n = 20, standard deviation [SD] = 3.5 nm; Fig. 1B). Two different types of nuclear particles were mainly observed: empty particles (capsids) and particles containing an electron dense core (nucleocapsids). Three different types of nucleocapsid cores were present: concentric, eccentric, and cores emerging from the nucleocapsid. A few atypical rod-shaped viral particles were also seen in the nucleus. In the perinuclear space, naked capsids and nucleocapsids budding at the inner nuclear membrane (Fig. 1C) were observed, and loss of nuclear membrane definition was detected in some areas.
In the cytoplasm, viral particles appeared enveloped and highly pleomorphic, with diameters ranging from 115 to 170 nm (Fig. 1D). In some cases, a single envelope was found surrounding 2 nucleocapsids. Enveloped nucleocapsids tended to form aggregates rather than appearing randomly distributed through the cytoplasm. The bi-layered envelope and the surface projections were clearly seen in some of the particles.
Cell culture
Cytopathic effect on the first passage was detected in pools 1–4 for each of the 4 types of cell lines (ASK, CHSE, EPC, and RTG-2) and in pool 5 for CHSE. All cultures showed weak CPE, except pools 1 and 5 in CHSE, which exhibited strong CPE. On the second passage, all cultures failed to yield any detectable CPE after 6 weeks of incubation, which is why aliquots of the first passage were used for DNA extraction. All intestine samples tested were negative for CPE on the first passage and were not analyzed further. None of the negative controls showed any signs of CPE.
Polymerase chain reaction, amplification, BLASTP, molecular dataset, and phylogenetic analysis
The degenerate DNA polymerase primers generated a 487-bp PCR product (including primers) from 6 out of 12 ethanol-preserved samples. The terminase primer pair yielded a 243-bp PCR product (including primers) from one of the ethanol-preserved DNA samples that had previously produced a strong positive band by agarose gel electrophoresis using the aforementioned degenerate PCR. A new specific primer pair (ACHV-3F 5′-GGGACGTGGACGGCAACCCTGTGG-3′ and ACHV-3R 5′-GCTTCACCTTGTGCTGCCCCTGG-3′), was designed from the DNA polymerase sequence generated. The new specific primer pair amplified products of the expected size (246 bp) in 9 out of 12 ethanol-preserved samples, in 1 out of 4 formalin-preserved samples, and in none of the cell culture samples.
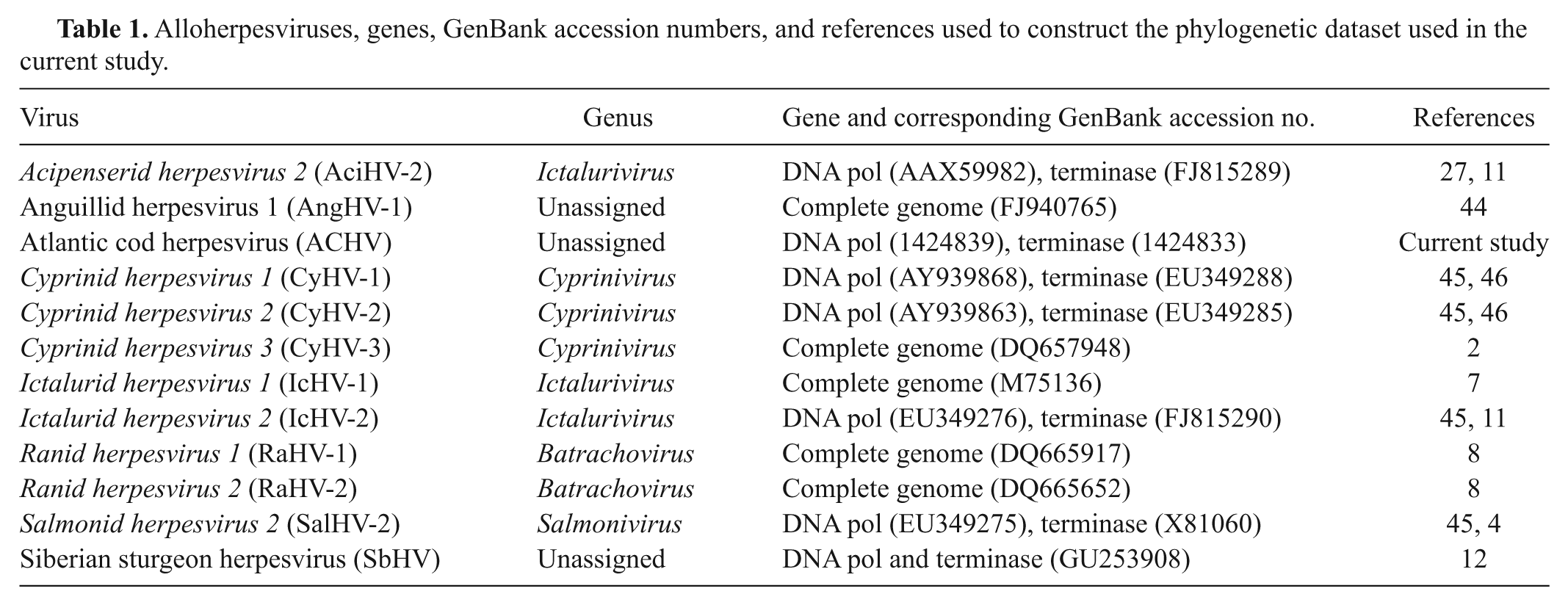
Overall, general BLASTP searches for the polymerase and terminase sequences revealed strongest sequence identity with alloherpesviruses belonging to the genus Ictalurivirus (Ictalurid herpesvirus 1, Ictalurid herpesvirus 2, and Acipenserid herpesvirus 2), lower similarity to alloherpesviruses belonging to the genera Salmonivirus (Salmonid herpesvirus 1, Salmonid herpesvirus 2, and Salmonid herpesvirus 3) and Batrachovirus (Ranid herpesvirus 1 and Ranid herpesvirus 2), and even lower similarity to members of the genus Cyprinivirus (Cyprinid herpesvirus 1, Cyprinid herpesvirus 2, and Cyprinid herpesvirus 3; data not shown). Hereafter, the virus will be referred to as the Atlantic cod herpesvirus (ACHV). BLASTP searches revealed 11 additional alloherpesviruses, for which each of the 2 homologous sequences were obtained (Table 1).
Alloherpesviruses, genes, GenBank accession numbers, and references used to construct the phylogenetic dataset used in the current study.
Preliminary phylogenetic analysis revealed that there was no significant incongruence among individual gene trees (defined by the presence of incompatible bipartitions that received a posterior probability of 90%, respectively) allowing concatenation of the sequence for the 2 genes into 1 final dataset. This dataset contained 225 amino acid characters (including gaps) for 12 viral taxa with at least 1 representative for each of the 4 genera within the family Alloherpesviridae. The concatenated 2-gene Bayesian analysis demonstrated with a high level of confidence that ACHV is an alloherpesvirus most closely related to the ictaluriviruses and salmoniviruses and more distantly related to the batrachoviruses and cypriniviruses (Fig. 2). Although not all regions of the tree were statistically supported, the overall tree topology was in agreement with recently published analyses,44,45 which further supports the value of partial sequence analyses in unraveling the evolutionary relationships of members of the order Herpesvirales.13,14,32,48
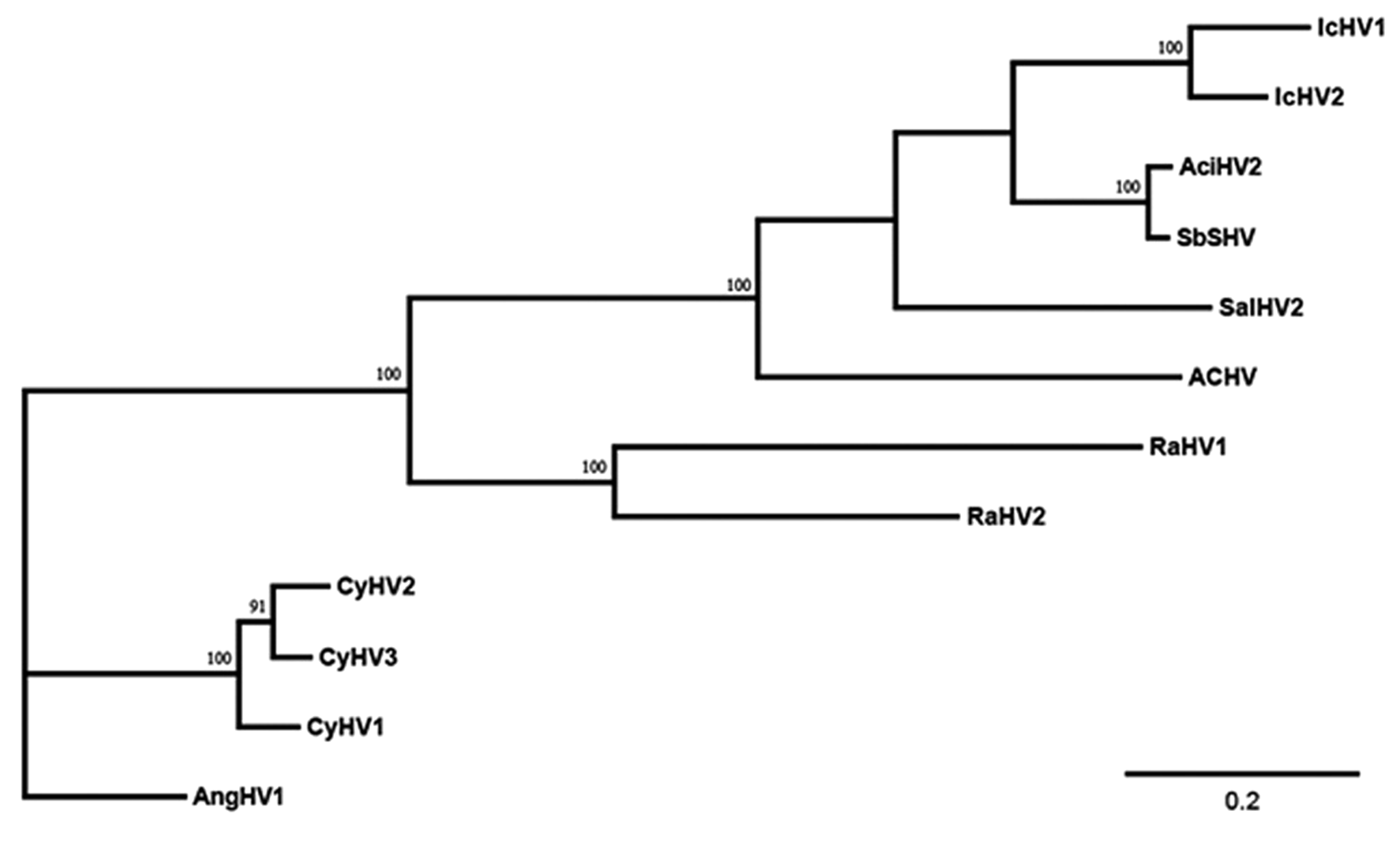
Phylogram depicting the relationship of the Atlantic cod herpesvirus (ACHV) to other fish and amphibian alloherpesviruses, based on the concatenated partial deduced amino acid (aa) sequences of the DNA polymerase and terminase genes (225 aa characters including gaps). Numbers above each node represent posterior probabilities (values >90 shown) of the Bayesian analysis. Branch lengths are based on the number of inferred substitutions, as indicated by the scale bar. Abbreviations: Ictalurid herpesvirus 1 (IcHV1), Ictalurid herpesvirus 2 (IcHV2), Acipenserid herpesvirus 2 (AciHV2), Siberian sturgeon herpesvirus (SbHV), Salmonid herpesvirus 2 (SalHV2), Atlantic cod herpesvirus (ACHV), Ranid herpesvirus 1 (RaHV1), Ranid herpesvirus 2 (RaHV2), Cyprinid herpesvirus 1 (CyHV1), Cyprinid herpesvirus 2 (CyHV2), Cyprinid herpesvirus 3 (CyHV3), Anguillid herpesvirus 1 (AngHV1).
In situ hybridization
In situ hybridization showed specific labeling of nuclear inclusions in the cytomegalic cells (Fig. 3). Nonspecific binding was not observed, as hybridization signals were not present in nonaffected cells. In addition, no cross-reaction occurred in sections positive for EEDV (data not shown).
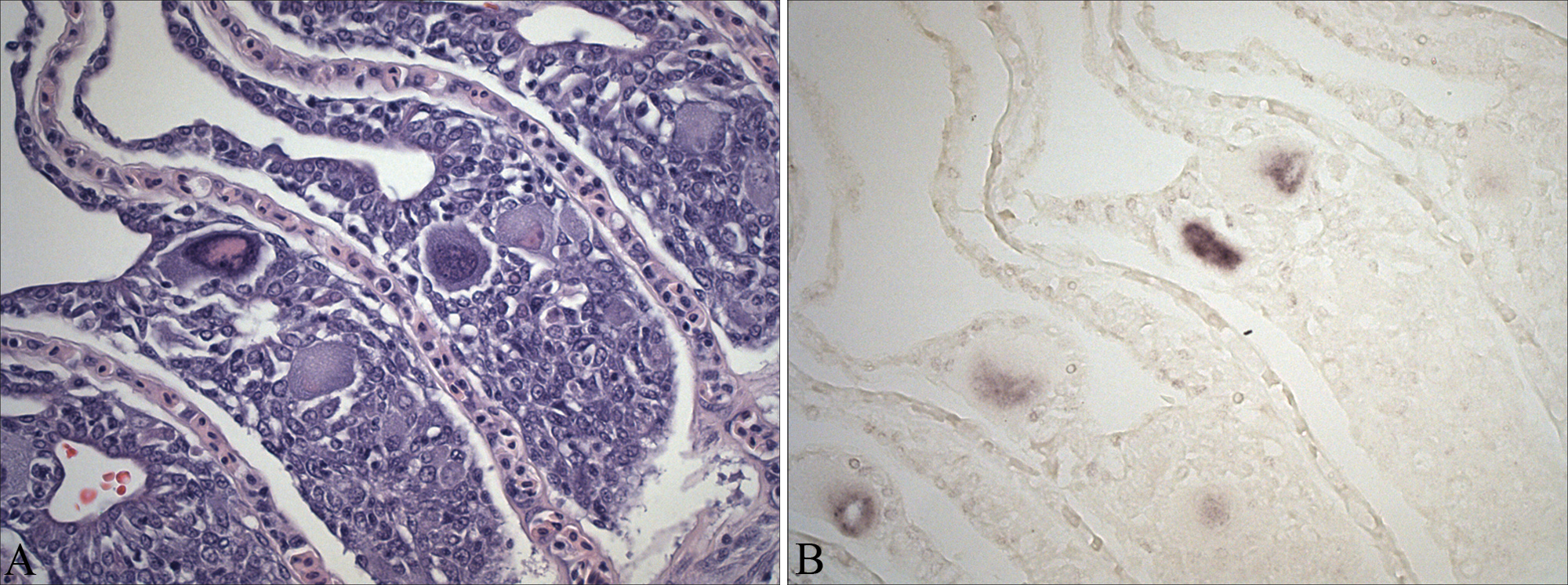
Atlantic cod (Gadus morhua). A, gill filament presenting some cytomegalic cells at the lamellar base. Section used for the in situ hybridization. Hematoxylin and eosin. Bar = 50 µm. B, in situ hybridization results. Hybridization signal evident in the cytomegalic cells. 0.005% Bismarck brown Y counterstain. Bar = 50 µm.
Discussion
The first Atlantic cod samples received at the IoA showed lesions that were compatible with bacterial infection due to Vibrio spp., and this infection was thought to be the probable cause of the observed mortality. The presumed viral infection in the gills was considered to be an incidental finding in this particular case. Nevertheless, the significance of the herpesvirus and whether it may have predisposed the fish to subsequent bacterial disease are unknown. Virus isolation and experimental infections would help to answer these questions.
Transmission electron microscopy was used as the first approach to identify the viral etiology. Viruses are classified according to their virion architecture, morphogenesis, and the nature and structure of their genetic material. DNA-containing viruses assemble in the nucleus, except nucleocytoplasmic large DNA viruses that include the viral families Poxviridae, Asfarviridae, Iridoviridae, and Phycodnaviridae. 23 The particles were herpesvirus-like due to the observed histopathology, size, icosahedral shape of the capsid, intranuclear location, and the presence of enveloped virions in the cytoplasm. Herpesviruses are double-stranded DNA viruses exhibiting icosahedral symmetry and consisting of 162 capsomeres. The viruses assemble in the nucleus and appear enveloped in the cytoplasm. Envelope shape can vary, and differences observed in TEM can be the result of variations in fixation and processing. The size of the mature virion given by the International Committee on Taxonomy of Viruses (ICTV) for herpesviruses is 120–200 nm, with the capsid diameter ranging from 100 to 110 nm. 10 Within this range are found some of the most well-known alloherpesviruses, such as Koi herpesvirus (virion size: 170–200 nm, capsid diameter: 110 nm) 33 and Channel catfish virus (virion size: 175–200 nm, capsid diameter: 100 nm). 50 The particles described in the present study are smaller, but other aquatic animal herpesvirus capsids are also below the expected range, such as the herpesvirus affecting common carp (82 nm), 6 the herpesvirus described in Pacific cod (80–100 nm),30,31 and Salmonid herpesvirus 1 (90 nm). 51 In bivalve mollusks, a previous study 35 described herpesvirus-like capsids measuring 71–86 nm in Pacific oyster (Crassostrea gigas) and Manila clam (Ruditapes philippinarum).
Herpesvirus infections in the gills have been reported in other fish species. For example, Koi herpesvirus is a severe and highly contagious viral disease characterized by hyperplasia and necrosis of the gill epithelium.17,39 The nuclei of affected cells appear with marginated chromatin and faint intranuclear inclusions. 17 Herpesvirus anguillae infecting fibrocytes of the gill filament has been associated with gill filament necrosis in farmed Japanese eel (Anguilla japonica). 29 In turbot (Scophthalmus maximus), herpesvirus scophthalmi infection results in the formation of giant cells in skin and gill epithelial cells.5,19,37 The associated gill lesions observed in turbot resemble those reported in the current study. In 1995 and 1998–1999, 2 epizootics causing huge mortalities in South American pilchard (Sardinops sagax) populations were reported in the waters of southern Australia, and these mortalities were associated with herpesvirus infection. 47 Lesions were confined to gills and, as with turbot, the lesions bore a striking similarity to those reported herein.
Most fish herpesviruses are epitheliotropic, inducing changes such as hypertrophy, hyperplasia, and necrosis of the epidermis, and in some cases, neoplasia. 18 In the present study, the infected cells located at the base of the lamellae, beneath the epithelium, were most likely progenitor epithelial cells. Hypertrophic cells more distally located in lamellae were probably epithelial cells of the inner layer of the lamellar epithelium or progenitor cells that had migrated up the lamellae. An infected chloride cell was also observed by TEM.
Different cell lines were employed in an attempt to isolate the viral agent. However, although CPE occurred, no virus was isolated, and DNA samples extracted from the cell cultures proved to be negative by PCR. Detection of CPE indicates a cytopathic effect within the samples, but CPE does not necessarily indicate which virus may be causing the effect or that a virus is, in fact, responsible for the observed effect, as cytotoxic effects could resemble those caused by a virus. On the other hand, samples having no CPE (i.e., pool 5 of ASK, EPC, and RTG-2) may still include a virus that does not induce observable CPE. The results observed suggest that the initial CPE observed resulted from cytotoxic effects, since no additional CPE was observed following a second passage. Alternatively, it is possible that a different virus had caused the initial CPE but was unable to produce CPE in the second passage. If cytotoxicity was responsible for the observed CPE, it is unknown why the intestine samples showed no detectable signs.
Specific primers designed against the DNA polymerase sequence confirmed ACHV in both the ethanol-preserved specimens (9/12) and in the formalin-preserved specimens (1/4). The nucleotide sequences obtained from formalin- and ethanol-positive samples confirmed that the amplicons detected by the screening PCR were identical to the original sequence generated by the initial degenerate PCR.
The positive results obtained in the ISH confirmed that ACHV detected by PCR was the cause of the observed cytomegaly. Although the specificity of the ACHV probe was assessed by using only EEDV-positive tissues, fish tissue sections infected with other closely related viruses will be included in the future to further confirm the probe specificity. The fact that all tested samples were not PCR positive is consistent with the histopathological observations in which the typical virally induced cytomegaly was not observed in every sample. The results may indicate that the virus establishes focal infections that are readily missed by sampling a few gill filaments for PCR and/or histology, or that the sampled fish were in fact uninfected with ACHV.
The phylogenetic analysis revealed that this new alloherpesvirus is most closely related to members of the genera Ictalurivirus and Salmonivirus. Adequate sequence data for comparison of the ACHV with other marine herpesviruses that induce similar pathology in turbot and pilchard are not currently available. Given the observed cellular pathology, virion architecture and morphogenesis, and the phylogenetic analysis, the authors propose the formal species designation Gadid herpesvirus 1 (GaHV-1) to be considered for approval by ICTV. Based on different diagnostic tests, molecular analyses, and ISH, the results presented in the current study confirm the discovery of a new alloherpesvirus affecting Atlantic cod.
Footnotes
Acknowledgements
a.
Tecnai™ G2 Spirit BioTwin TEM, FEI Co., Hillsboro, OR.
b.
DNeasy® DNA Blood and Tissue Extraction kit, Qiagen Gel Extraction kit, QIAprep® Spin Miniprep Kit; Qiagen Inc., Valencia, CA.
c.
pGEM®-T Easy Vector System, Promega Corp., Madison, WI.
d.
ABI 377 automated sequencer, Applied Biosystems, Foster City, CA.
e.
PCR DIG probe synthesis kit, Roche Applied Science, Indianapolis, IN.
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Mar Marcos-Lopez received funding from a Fundacion Caja Madrid scholarship. Funding for research was provided by the New Brunswick Department of Agriculture and Aquaculture Total Development Fund (project AA10046).
