Abstract
A novel siadenovirus was identified in the Sulawesi tortoise (Indotestudo forsteni). A group of 105 Sulawesi tortoises was obtained by the Turtle Survival Alliance. Many of the tortoises were in poor health. Clinical signs included anorexia, lethargy, mucosal ulcerations and palatine erosions of the oral cavity, nasal and ocular discharge, and diarrhea. Initial diagnostic tests included fecal testing for parasites, complete blood count and plasma biochemical analysis, mycoplasma serology, and polymerase chain reaction (PCR) testing for intranuclear coccidia and chelonian herpesvirus. Treatment included administration of antibiotics, antiparasitic medications, parenteral fluids, and nutritional support. Tissue samples from animals that died were submitted for histopathologic evaluation. Histopathologic examination revealed systemic inflammation and necrosis associated with intranuclear inclusions consistent with a systemic viral infection in 35 tortoises out of 50 examined. Fecal testing results and histopathologic findings revealed intestinal and hepatic amoebiasis and nematodiasis in 31 animals. Two of 5 tortoises tested by PCR were positive for Chlamydophila sp. Aeromonas hydrophila and Escherichia coli were cultured from multiple organs of 2 animals. The mycoplasma serology and PCR results for intranuclear coccidia and chelonian herpesvirus were negative. Polymerase chain reaction testing of tissues, plasma, and choanal/cloacal samples from 41 out of 42 tortoises tested were positive for an adenovirus, which was characterized by sequence analysis and molecular phylogenetic inference as a novel adenovirus of the genus Siadenovirus. The present report details the clinical and anatomic pathologic findings associated with systemic infection of Sulawesi tortoises by this novel Siadenovirus, which extends the known reptilian adenoviruses to the chelonians and extends the known genera of reptilian Adenoviridae beyond Atadenovirus to include the genus Siadenovirus.
Keywords
Introduction
Adenoviral infections have been reported in all classes of bony vertebrates and are a significant cause of disease in many species. In reptiles, adenoviral infections have been described in lizards, snakes, and Nile crocodiles. 14,28,30,34,41,43,44,52 Adenoviridae are nonenveloped, double-stranded DNA viruses with a medium-sized genome of 26–45 kilo base pair (kbp). Like many DNA viruses, phylogenetic analyses of adenoviruses (AdVs) indicate that many elements in the branching patterns of these viruses are congruent with branching patterns for the corresponding host species. 3 This finding suggests that these viruses have often coevolved along with their hosts, although host switches have certainly occurred. Adenoviruses are classified into 4 genera: Mastadeno-virus, Aviadenovirus, and the 2 recently accepted genera Atadenovirus and Siadenovirus. 4 A fifth genus is proposed for fish AdVs. 4 Members of Mastadenovirus are found in mammals, and aviadenoviruses are found in birds. Atadenoviruses have been found in birds 23 and mammals, 8,49 but it appears that their most probable origin is in reptiles. 2,14,22,34 Siadenoviruses have been found in birds 42 and amphibians. 9 The origin of the Siadenovirus genus is not clear.
The only characterized reptile AdVs to date have been from squamates, and all have been in the genus Atadenovirus. 2,14,52 Although AdV-like particles have been seen in crocodilians, they have not been characterized. 29 Findings seen in reptiles associated with AdV-like agents include hepatitis, 29,33 enteritis, 24 esophagitis, 27 splenitis, 24 and encephalopathy. 44
The current study describes disease in a group of confiscated wild-caught Sulawesi tortoises (Indotestudo forsteni) that had been imported into the United States. The Sulawesi tortoise inhabits the islands of Sulawesi and Halmahera in Indonesia and is currently listed under appendix II of the Convention on International Trade of Endangered Species (CITES). Moreover, this species is considered an endangered species under the International Union for Conservation of Nature (IUCN) Red List. As with many other chelonian species, pressures from loss of native habitat and harvesting for consumption and the pet trade have made this species rare in its home range. 7,36 The large-scale commercial trade of chelonians has created concern for the survival of wild populations, and the spread of pathogens to naïve populations.
Material and methods
Animals
One hundred and five illegally imported Sulawesi tortoises were confiscated by the U.S. Fish and Wildlife Service upon entry into the United States. Thirty tortoises died prior to veterinary examination, and their carcasses were not available for diagnostic studies. Tortoises were sent to institutions where veterinary care was readily available including Bronx Zoo (Bronx, NY), Fort Worth Zoo (Fort Worth, TX), Lake Howell Veterinary Clinic (Maitland, FL), Dr. Charles J. Innis (Boston, MA), and Zoo Atlanta (Atlanta, GA). Medical therapy varied at the discretion of individual clinicians; however, all tortoises were treated with systemic antibiotics, antiparasitics, fluid therapy, and nutritional support.
Clinical pathology
Fecal samples were collected for parasite screening. Blood samples were collected from 19 tortoises and submitted for complete blood count (CBC) and plasma biochemical analysis.
Pathology
Necropsies were performed at the participating institutions. Tissue samples were submitted for histopathologic evaluation, and additional samples were frozen. Tissue selection occasionally varied between institutions depending on choices made by prosectors. Tissues submitted for histopathologic evaluation were fixed by immersion in 10% neutral buffered formalin. Samples containing bone were decalcified before processing according to procedures currently in use by the several recipient pathology services. Tissues for histologic sectioning were processed in a routine manner, embedded in paraffin, sectioned at 4–5 μm thickness, mounted on glass slides, stained with hematoxylin and eosin (HE), and examined by light microscopy.
Transmission electron microscopy
Sections of selected tissues (e.g., liver, bone marrow) were finely minced, and fixed in a solution containing 2% paraformaldehyde, 2% glutaraldehyde, and 0.2% picric acid in 0.1 mol of cacodylate-HCl buffer. After fixation, the tissues were embedded in resin. Thin sections were prepared and stained with osmium tetroxide and Reynolds' lead citrate for examination by transmission electron microscopy.
Formalin-fixed samples of spleen from 2 specimens, each tissue of which had been identified by prior histopathologic examination as having nuclei with features suspect of viral inclusions, were processed using a microwave protocol modified from previously published techniques 20,21 and using a commercial microwave system with variable wattage, a microwave processing surface and water load, a and specimen holders. a Tissues were trimmed to 1–2 mm 3 dimensions, then immersed in McDowell–Trump fixative. 38 Tissue processing included postfixation in 1% OsO4 under a vacuum of 25 mm Hg, en bloc staining with 1% uranyl acetate, dehydration in a graded series of ethanol, then infiltration and embedment using London Resin (LR) White resin (medium grade) b under a vacuum of 25 mm Hg. Thin sections were stained with uranyl acetate and lead citrate, then viewed in an electron microscope c at 80 kV.
Adenovirus PCR amplification and sequencing
Samples from 42 Sulawesi tortoises (5 nasal flush, 5 choanal swab, 2 choanal/cloacal swab, 20 plasma from live animals; and 9 oral/nasal mucosal tissues, 1 colon, 1 small intestine, 11 liver, 2 lung, 5 kidney, and 6 spleen from deceased animals) were collected for PCR analysis. DNA was extracted from samples using a commercial extraction kit. d Adenoviral consensus nested PCR amplification of a partial sequence of the DNA-dependent DNA polymerase gene was performed using previously described methods. 52 Products were resolved on 1% agarose gels, and bands of the expected size were excised and purified using a commercial gel extraction kit. d Direct sequencing was performed using a commercial cycle sequencing kit e and analyzed on automated DNA sequencers e at the University of Florida Interdisciplinary Center for Biotechnology Research DNA Sequencing Facilities (Gainesville, FL). Similarly, PCR identification, sequencing, f and sequence analysis were independently performed at the Department of Pathobiology and Veterinary Science at the University of Connecticut (Storrs, CT) using a different set of tissue samples. All products were sequenced in both directions. Primer sequences were not included in subsequent sequence analysis.
In situ hybridization using riboprobes
A 272–base pair (bp) sequence of the AdV DNA–dependent DNA polymerase gene was cloned using a commercial TOPO (topoisomerase I) TA (Taq-amplified) cloning kit and plasmid pCRII-TOPO. g Orientation of the plasmid insert was confirmed by DNA sequencing. Plasmids were used to create labeled RNA probes (riboprobes) with the Sp6 and T7 transcription kits h and digoxigenin-uridine triphosphate RNA labeling mix i as described by the manufacturer's instructions. Dot blot analyses were used to assess appropriate digoxigenin labeling and to estimate concentrations of transcribed riboprobes, which were compared with known concentrations of digoxigenin-labeled control RNA i detected by alkaline phosphatase-conjugated antidigoxigenin Fab fragments i and the 5-bromo-4-chloro-3-indolyl-phosphate/4-Nitro blue tetrazolium (BCIP-NBT) color substrate system. i
Tissue sections, including the spleen from 2 tortoises, were chosen. The in situ hybridization (ISH) protocol was adapted from previously published protocols. 5,11,18 Serial sections were mounted unstained on glass slides. j Slides were heated to 70°C for 30 min in a slide moat to melt paraffin, then subjected to three 3-min washes in xylene. Tissues were allowed to dry, and then rehydrated with 1× phosphate buffered saline (PBS) plus 5 mM of MgCl for 15 min at room temperature. Tissues were digested by incubation with 20 μg/ml proteinase K in 10 mM of Tris plus 2 mM of CaCl2, pH 7.5, for 5 min at 42°C. Enzyme activity was stopped by immersing slides in 0.2 M of Tris, pH 7.5, plus 0.1 M of glycine for 1 min, and a second time for 2 min. Two hundred microliters of prehybridization solution k was applied, and slides were cover-slipped and incubated at 42°C for 1 hr. Coverslips were removed, prehybridization solution was drained, and slides were covered with 100 μl of prehybridization buffer containing 85–200 ng of riboprobe and cover-slipped. Target DNA was denatured by heating slides to 95°C for 3 min, after which slides were cooled to 85°C and covered with parafilm to prevent dehydration. The temperature was then decreased to 38°C for an overnight incubation. The next day, slides were washed in 2× saline-sodium citrate solution (SSC) with 1% sodium dodecyl sulfate (SDS) at 50°C for 20 min, followed by 1× SSC with 0.1% SDS at 50°C for 20 min, then 1× SSC at room temperature for 10 min, and finally 0.1× SSC at room temperature for 15 min. Slides were rinsed in buffer 1 (0.1 M of Tris, 0.15 M of NaCl, pH 7.5) for 1 min and then incubated in buffer 1 plus 10% 5 mM levamisole l for 30 min. Slides were then covered with 150 μml of buffer 1 containing 2% sheep serum l and alkaline phosphatase-conjugated anti-digoxigenin Fab fragments at a 1:250 dilution, cover-slipped, and incubated for 2 hr at 37°C. Following incubation, slides were washed in buffer 1 at room temperature for 15 min, then in TBST (0.15 M of NaCl, 2.7 mM of KCl, 25 mM of Tris, and 0.05% of Tween 20, pH 7.6) for two 5-min washes. Hybridization signal development was achieved using Permanent Red m with 4.5% 5 mM levamisole. Slides were counterstained with Gill's modified hematoxylin, air dried, cleared with alcohols and xylene, and cover-slipped according to manufacturer's directions. m The control template provided in the EPICENTRE T7 transcription kit h was similarly transcribed to generate a nonhomologous control riboprobe used in ISH experiments.
Other diagnostic work
Plasma samples were submitted for mycoplasma serology. Cloacal swab samples were submitted for bacterial culture and sensitivity testing. Samples from various organs (kidney, liver, ovary, and oviduct) and coelomic effusion were also submitted for bacterial culture and sensitivity from 2 tortoises. Chlamydiales consensus PCR analysis was performed on specimens from 5 tortoises (2 spleen, 5 liver) using previously described methods. 13 Herpesviral consensus PCR analysis was performed on specimens from 13 tortoises (4 nasal flush, 2 oral mucosa, 5 choanal swab, 8 liver, 2 spleen) using previously described methods. 50 Coccidial consensus PCR analysis was used to detect any intranuclear coccidia. This technique was performed on specimens from 16 tortoises (4 nasal flush, 7 nasal mucosa, 5 choanal swab) using previously described methods. 19
Phylogenetic analysis
The sequences of the adenoviral DNA–dependent DNA polymerase gene were compared with those in GenBank (http://www.ncbi.nlm.nih.gov/Genbank/index.html), EMBL Nucleotide Sequence Database (http://www.ebi.ac.uk/embl), and DNA Data Bank of Japan (http://www.ddbj.nig.ac.jp) databases using TBLASTX for protein coding sequence or BLASTN for nonprotein coding sequence. 1 Predicted homologous 89–91 amino acid sequences of adenoviral DNA–dependent DNA polymerase were aligned using 3 methods: CLUSTAL W, 48 T-COFFEE, 39 and MUSCLE. 12
Bayesian analyses of each amino acid alignment were performed using MrBayes version 3.1 46 (http://mrbayes.csit.fsu.edu/index.php) with gamma distributed rate variation and a proportion of invariant sites, and mixed amino acid substitution models. Statistical convergence was assessed by looking at the standard deviation of split frequencies as well as potential scale reduction factors of parameters. The first 10% of 1,000,000 iterations were discarded as a burn in.
Maximum likelihood (ML) analyses of each amino acid alignment were performed using PHYLIP version 3.66 16 (PHYLogeny Inference Package; http://evolution.genetics.washington.edu/phylip.html), running each alignment in proml with amino acid substitution models JTT, 31 PMB, 51 and PAM 35 further set with global rearrangements, 5 replications of random input order, less rough, gamma plus invariant rate distributions, and unrooted. The values for the gamma distribution were taken from the Bayesian analysis. Sturgeon adenovirus was designated as the out-group. The combination of alignment producing the most likely tree was then used to create data subsets for bootstrap analysis to test the strength of the tree topology (200 resamplings), 15 which was analyzed using the amino acid substitution model producing the most likely tree.
Results
Animals
The initial evaluation of the surviving tortoises (75 animals) revealed that all individuals were in poor health. Abnormal clinical signs documented included anorexia, lethargy, mucosal ulcerations and palatine erosions of the oral cavity, nasal and ocular discharge, and diarrhea. Body weights ranged from 0.23–1.65 kg. Despite aggressive treatment, many tortoises died within 8 weeks of initiation of treatment, with the majority of the deaths occurring within the first 3 weeks. A small group of tortoises responded to therapy and started eating within a short period, but several later died from adenoviral infections. The mortality rate was 82%. Thirteen (17% of the original group) survived 20 months after initial presentation.
Complete blood cell count results from 19 sick Sulawesi tortoises.
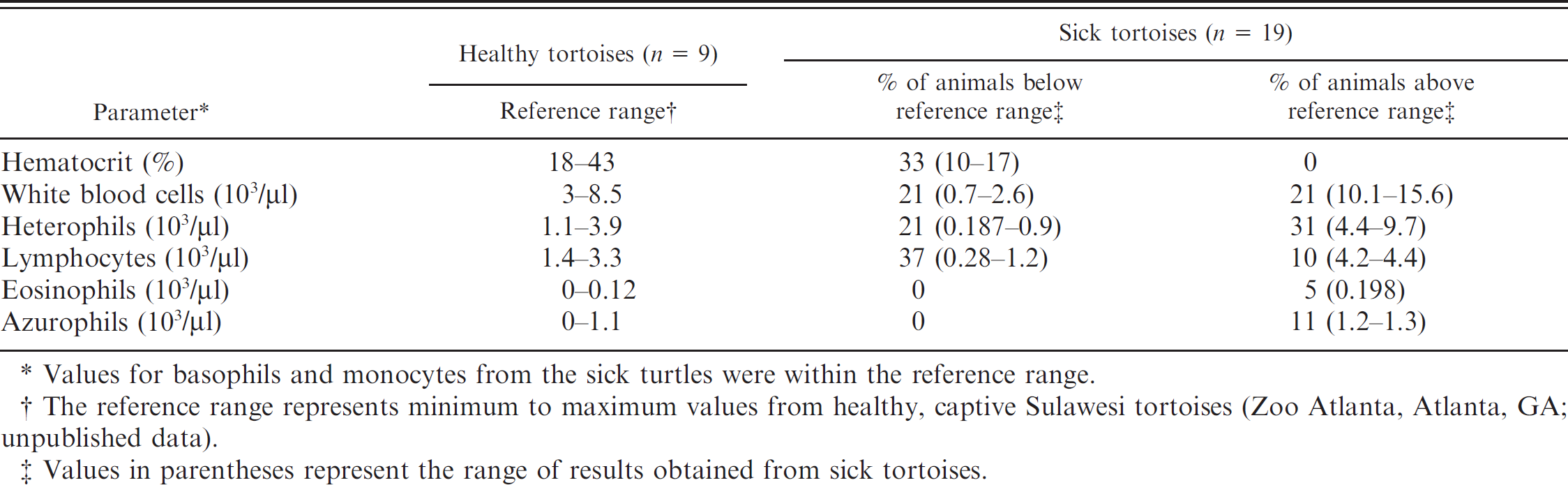
Values for basophils and monocytes from the sick turtles were within the reference range.
The reference range represents minimum to maximum values from healthy, captive Sulawesi tortoises (Zoo Atlanta, Atlanta, GA; unpublished data).
Values in parentheses represent the range of results obtained from sick tortoises.
Clinical pathology
The CBC and plasma biochemical analysis results were compared with in-house results obtained at Zoo Atlanta from healthy Sulawesi tortoises (n = 9). Based on these comparisons, the abnormal values seen in the CBC results were anemia (33% of affected animals), leukopenia (21%), leukocytosis (21%), heteropenia (21%), heterophilia (31%), lymphopenia (37%), lymphocytosis (10%), eosinophilia (5%), and azurophilic monocytosis (11%; Table 1). The most common abnormalities found in the plasma biochemical analysis results were elevated aspartate amino-transferase activity (28%), elevated creatinine phosphokinase activity (5%), hypoglycemia (21%), hyperglycemia (58%), elevated blood urea nitrogen (BUN) concentration (92%), hyperkalemia (22%), and elevated uric acid concentration (21%; Table 2).
Pathology
Sixty-two of 75 tortoises died despite aggressive treatment. Full or partial necropsies were performed on 50 animals. Gross necropsy lesions included hepatosplenomegaly, thick fibrinonecrotic membranes in the lumen of the large intestine, segmental darkening of the serosal surfaces of the small and large intestine, oronasal fistulae, and tongue and oral mucosa ulceration. The major pathologic findings are presented in Table 3.
Intestine. Severe, multifocal to diffuse, variably hemorrhagic, necrotizing enterocolitis with pseudo-membrane formation was consistently found in animals that died early in the course of the disease outbreak. Forty-one of 50 tortoises examined had significant lesions in the digestive system (Table 3). In the most severely affected animals, widespread mucosal effacement was accompanied by severe edematous expansion of the submucosa, heterophilic and lymphoplasmacytic infiltration of submucosal and muscularis layers, and severe intraluminal and interstitial proliferation of bacilli and cocci. Variable numbers of degenerating or intact amoebic trophozoites or cysts and many submucosal heterophilic granulomas, which were often centered on degenerating or intact amoebae and fragments of ascarids, were also found in 31 tortoises. One specimen also had severe zygomycete invasion of the muscularis mucosa of the intestine. As later cases survived longer and partially responded to therapeutic efforts before death, the numbers of parasites decreased, and the intestinal mucosae appeared more intact. Intact enterocytes often contained intranuclear viral inclusions. The inclusions were variably shaped, basophilic or amphophilic, refractile, and associated with margination of chromatin and abnormal nuclear shapes and sizes.
Plasma biochemistry results from 19 sick Sulawesi tortoises
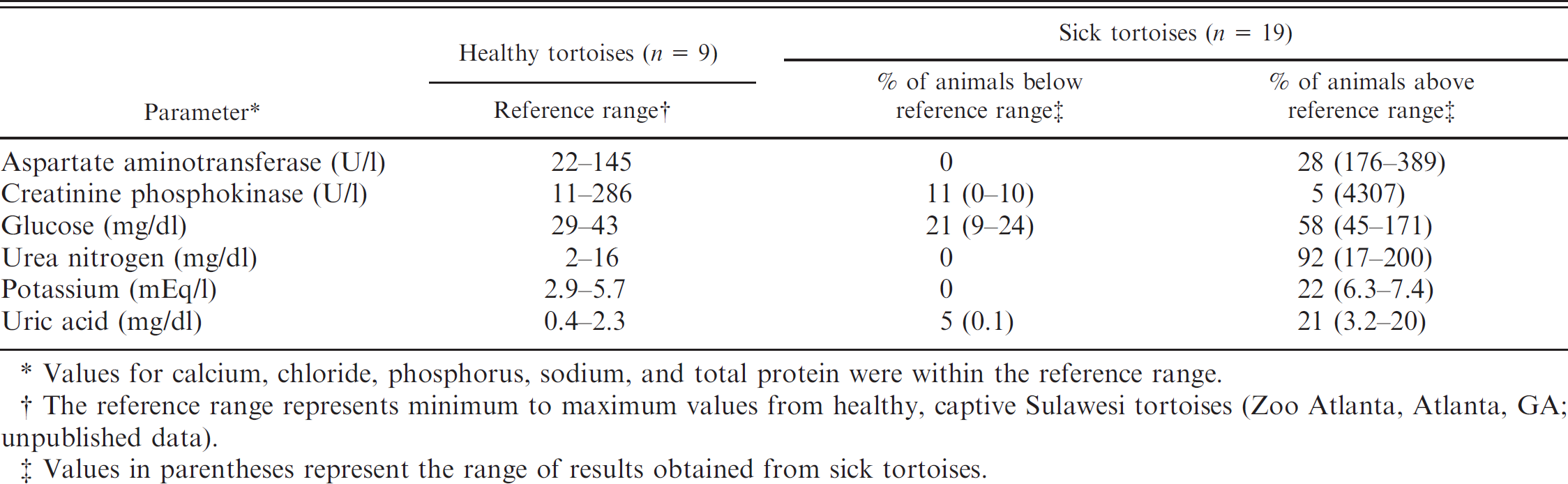
Values for calcium, chloride, phosphorus, sodium, and total protein were within the reference range.
The reference range represents minimum to maximum values from healthy, captive Sulawesi tortoises (Zoo Atlanta, Atlanta, GA; unpublished data).
Values in parentheses represent the range of results obtained from sick tortoises.
Major pathologic findings in 50 Sulawesi tortoises diagnosed with systemic adenovirus infection by polymerase chain reaction (n = 26) and/or identification of viral inclusions (n = 35).
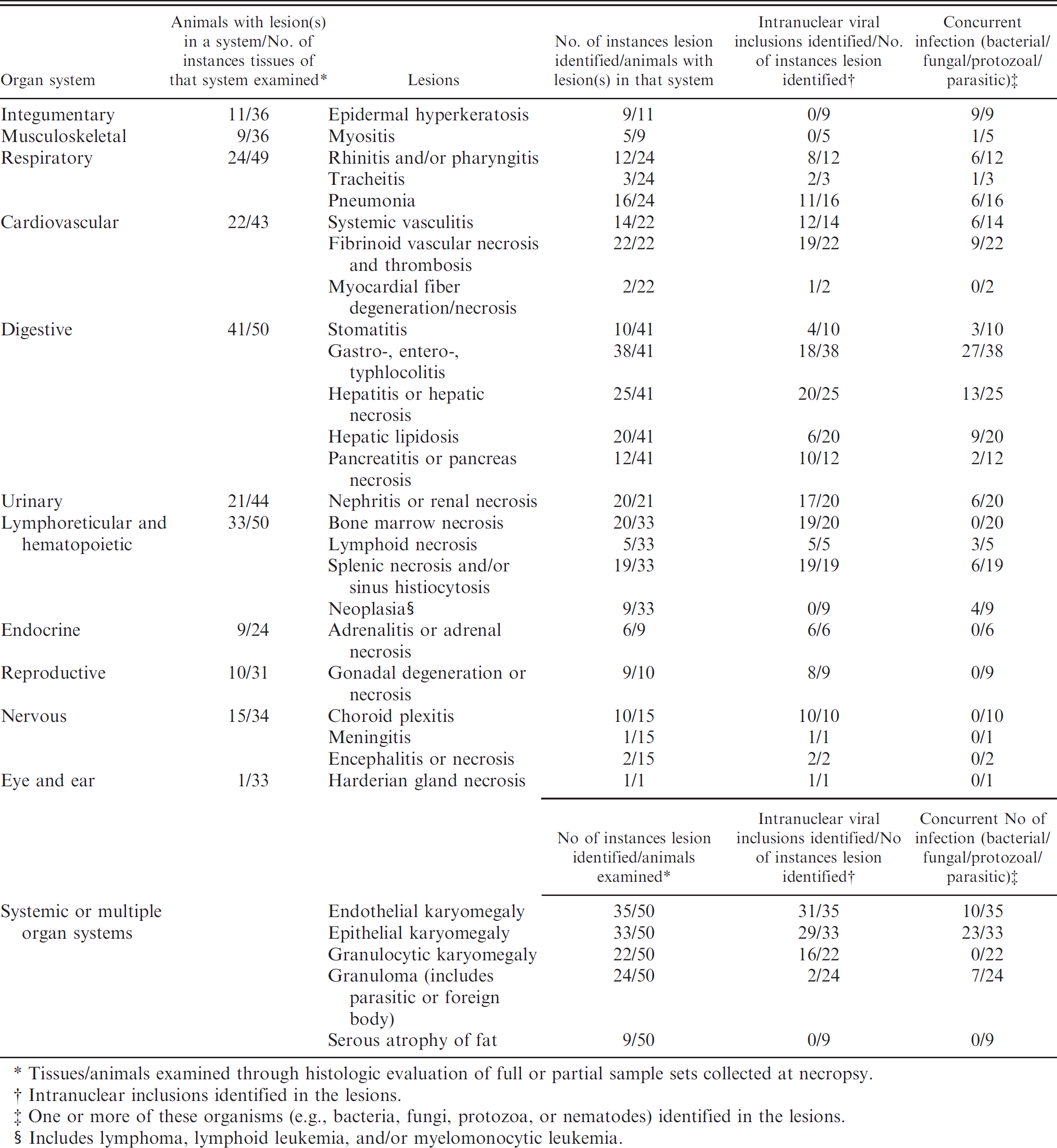
Tissues/animals examined through histologic evaluation of full or partial sample sets collected at necropsy.
Intranuclear inclusions identified in the lesions.
One or more of these organisms (e.g., bacteria, fungi, protozoa, or nematodes) identified in the lesions.
Includes lymphoma, lymphoid leukemia, and/or myelomonocytic leukemia.
Liver. In addition to amoebic hepatitis in some animals, early cases of disease had individual hepatocellular necrosis or multifocal to submassive coagulative hepatocellular necrosis with admixed heterophils. Hepatocyte nuclei were often enlarged and many contained variably shaped, basophilic or amphophilic, refractile inclusions associated with margination of chromatin. Intranuclear inclusions were seen in 20 of 25 instances where hepatitis or hepatic necrosis was identified (Table 3). These intranuclear inclusions also were seen in the hepatic sinusoidal endothelium, Kupffer cells, and leukocytes. Moderate, diffuse, glycogen-type and lipid-type hepatocyte vacuolation and variable numbers of melanomacrophage aggregates were observed.
Bone marrow. Microscopic examination of the decalcified bone sections revealed myeloid necrosis. Bone marrow necrosis was identified in 20 of 33 animals examined and was associated with the presence of intranuclear inclusions in myeloid and stromal of the hematopoietic tissue in 19 of 20 instances (Table 3; Fig. 1).
Viral inclusions in other tissues. In animals that lived longer with intensive clinical support, intranuclear inclusions were identified in many other tissues, including spleen, pancreas, testis, ovary, kidney, respiratory epithelium, renal epithelium, vascular and cardiac endothelium, and cerebral glia and choroid plexus (Table 3). Viral inclusions were not identified in animals that died at least 5 months after the initial diagnosis of adenoviral infection.
Other histologic findings. Severe diffuse splenic necrosis was found in 19 of 33 animals examined (Table 3). Neoplasia (5 cases of lymphoma, 1 case of lymphoid leukemia, and 3 cases of myelomonocytic leukemia) was identified in AdV PCR-positive animals that survived several months after the initial clinical diagnosis. A Chlamydophila-like agent was identified within the renal tubular epithelium in 1 tortoise. A few large ciliates and other protozoal organisms also were seen in the renal epithelium of some animals.
Transmission electron microscopy
Electron microscopy performed by several of the pathology services independently confirmed the presence of nonenveloped, 70–90 nm diameter, viral particles in nuclei with marginated chromatin and abnormal nuclear shapes and sizes (i.e., those interpreted as having inclusions on histopathologic examination; Fig. 2A). On occasion, viral particles were observed to form paracrystalline arrays in nuclei (Fig. 2B).
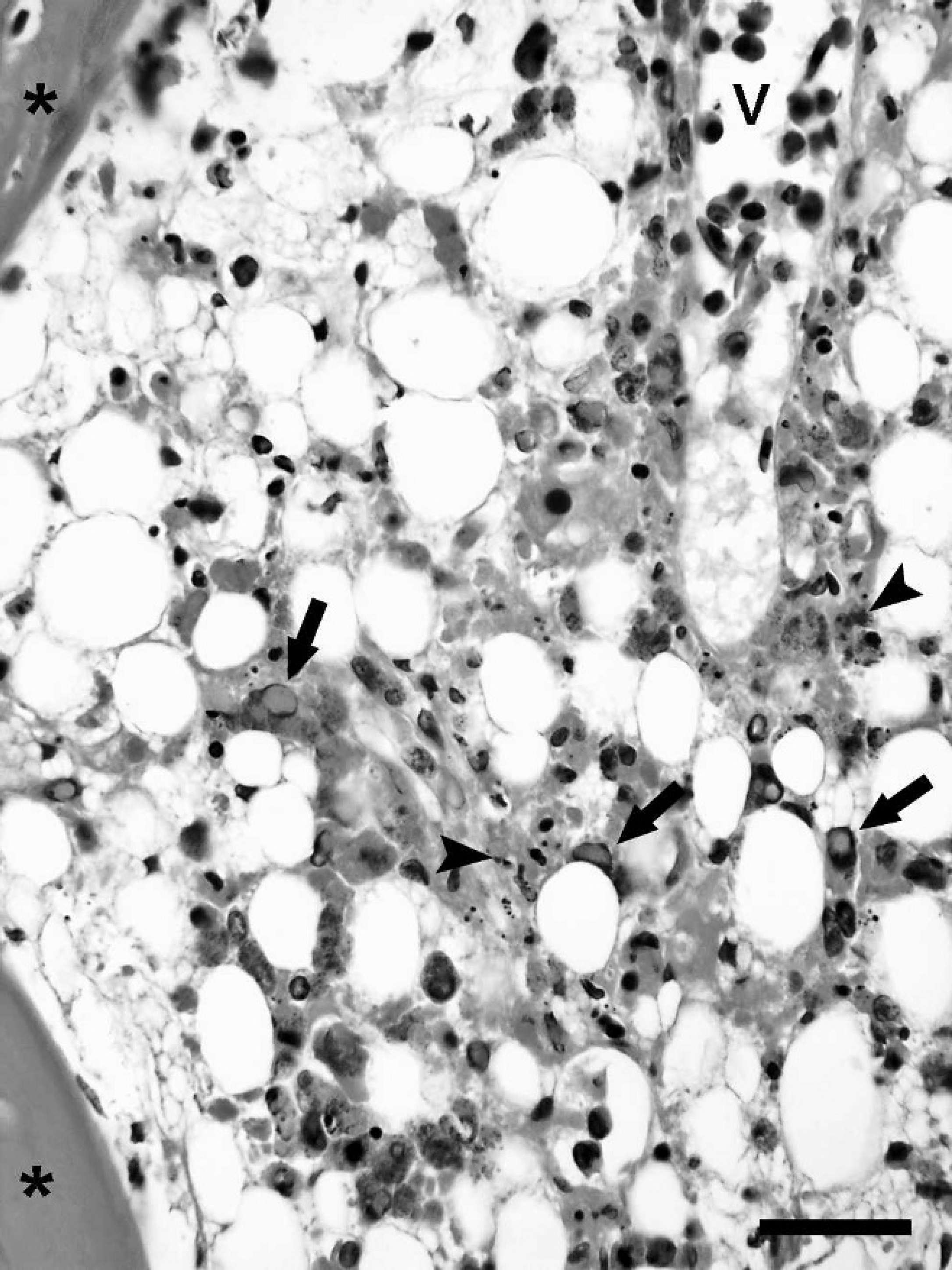
Sulawesi tortoise, bone (*) and bone marrow. Several myeloid cells have enlarged round-to-ovoid nuclei with marginated chromatin and ground glass appearance of the nucleoplasm (arrows), typical of the intranuclear inclusion observed in systemic infections by Sulawesi tortoise adenovirus-1. Necrotic cells, pyknotic and karyorrhectic debris (arrowheads) are present among myeloid cells and along the margin of a blood vessel (V). Hematoxylin and eosin. Bar = 50 μm.
Adenovirus PCR amplification and sequencing
Adenovirus PCR amplicons were detected from 41 of 42 (97.6%) tortoises (5 nasal flush, 9 oral/nasal mucosal tissue, 5 choanal swab, 2 choanal/cloacal swab, 19 of 20 plasma samples, 1 colon, 1 small intestine, 11 liver, 2 lung, 5 kidney, and 6 spleen). Adenovirus PCR amplification yielded a 272-bp product after exclusion of primer sequences. The sequence of all PCR products was identical (Fig. 3). All nasal flush, nasal mucosa, choanal and choanal/cloacal swab, colon, liver, lung, kidney, and spleen samples were positive. Nineteen of 20 plasma samples were positive. The negative tortoise only had a plasma sample examined. This 272-bp sequence was submitted to GenBank under accession number EU056826.
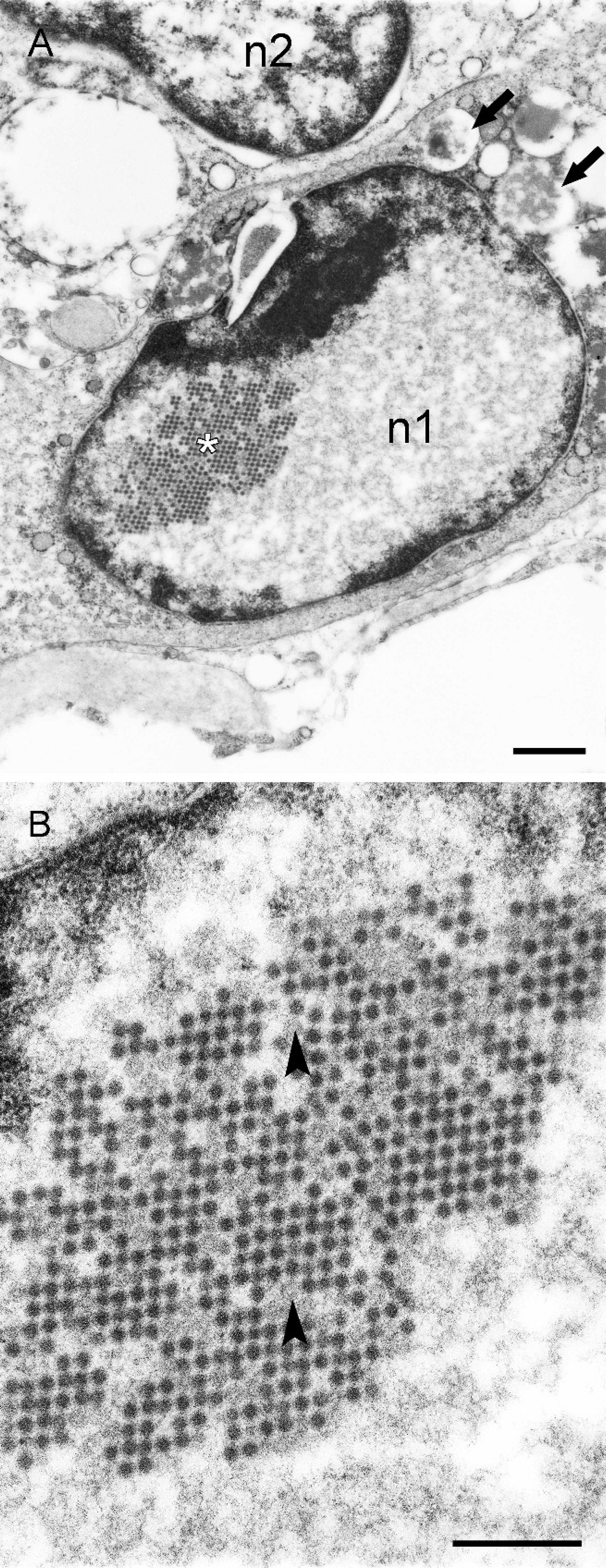
Sulawesi tortoise, spleen; transmission electron photomicrographs. A, this virus-infected cell has an oval nucleus (n1) with marginated chromatin and a paracrystalline array of virus particles (*). Note the presence of vacuoles containing amorphous flocculent material (arrows) in the cytoplasm. The nucleus of an adjacent, noninfected cell (n2) is visible. Lead citrate and uranyl acetate. Bar = 1 μm. B, higher magnification of the paracrystalline array of virus particles in the nucleus. Virions are hexagonal and approximately 70 nm in width; occasional empty nucleocapsids are present (arrowheads). Lead citrate and uranyl acetate. Bar = 500 nm.
Adenoviral in situ hybridization using riboprobes
Digoxigenin-labeled riboprobe transcribed from the cloned fragment of the AdV DNA–dependent DNA polymerase gene generated hybridization signal on cells with nuclear sizes and shapes interpreted as having viral inclusions in sections of liver, kidney, and/or spleen in several runs (Fig. 4A). Hybridization signal was variable in intensity among these runs. No hybridization signal was detected in successive tissue sections to which the digoxigenin-labeled riboprobe transcribed from the EPICENTRE T7 control template was applied (Fig. 4B).
Other diagnostic results
Fecal testing results revealed amoeba trophozoites and nematode ova. A. hydrophila and E. coli were cultured from multiple organs (liver, kidney, ovaries, oviduct, and coelomic effusion) in 2 tortoises. The mycoplasma serology results were negative. All herpesvirus and coccidia PCR results were negative. Chlamydial PCR products were detected from 2 of 5 (40%) tortoises that were tested. One of 2 spleen samples and 2 of 5 liver samples were PCR test positive. Chlamydial PCR amplification yielded a 260-bp product after exclusion of primer sequences. The 3 chlamydia PCR products had an identical sequence. This sequence was submitted to GenBank under accession number EU128171.
Phylogenetic analysis
Comparison with other sequences using TBLASTX revealed that the AdV PCR product sequence was similar to, but distinct from, other known AdVs present in the available databases. The highest score obtained was with Frog AdV 1 (GenBank accession no. AF224336), a member of the genus Siadenovirus.
Bayesian phylogenetic analysis of the Adv showed the greatest harmonic mean of estimated marginal likelihoods using the T-COFFEE alignment (Fig. 3). The Rtrev model of amino acid substitution was found to be most probable with a posterior probability of 1.000. 10 A Bayesian tree using the T-COFFEE alignment is shown (Fig. 5).
Maximum likelihood analysis found the most likely tree from the CLUSTAL W alignment and the PMB model of amino acid substitution. These parameters were used for bootstrap analysis. Bootstrap values from ML analysis are shown on the Bayesian tree (Fig. 5).

T-COFFEE alignment of partial adenoviral DNA polymerase amino acid sequences. Lines separate genera. Sulawesi tortoise adenovirus 1 is in boldface.
Comparison with other sequences using BLASTN revealed that the sequence of the chlamydiales PCR product was consistent with Chlamydophila or Chlamydophila psittaci. The highest scores obtained were tied with 5 isolates of C. abortus and C. psittaci. (GenBank accession nos. D85709, D85710, AB001809, AB001810, and AB001811).
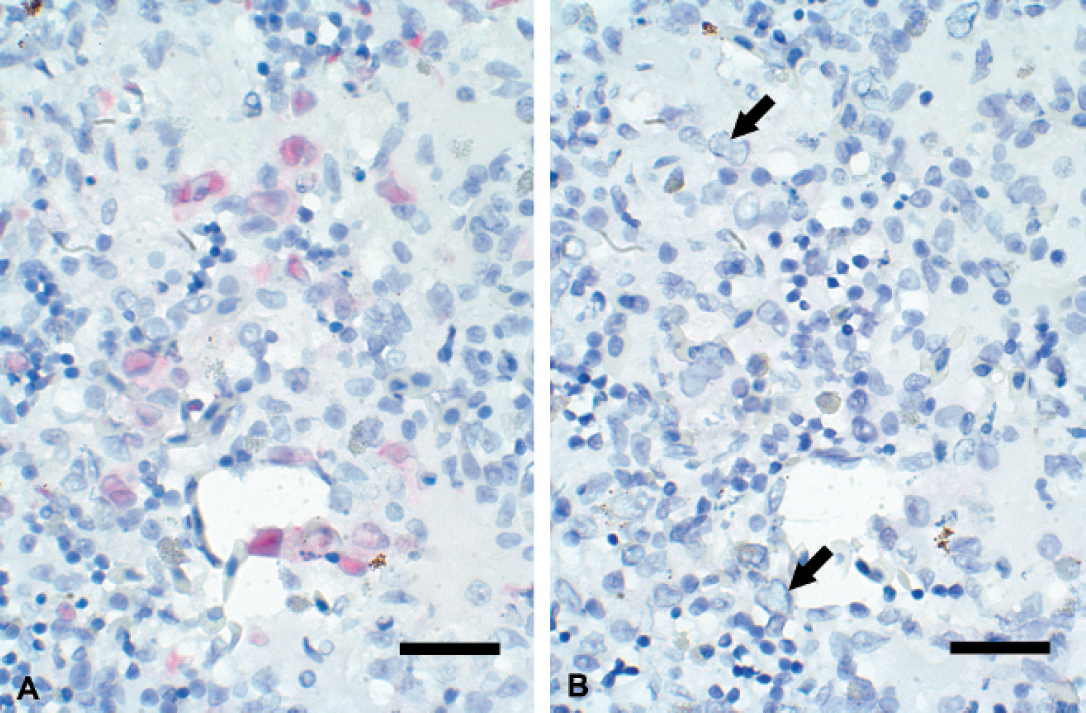
Sulawesi tortoise, spleen; in situ hybridization using riboprobes. A, hybridization signal is present on abnormally sized and shaped nuclei using a digoxigenin-labeled riboprobe transcribed from the cloned 272 base pair (bp) fragment of the adenovirus DNA-dependent DNA polymerase gene. Permanent Red substrate and Gill's hematoxylin counterstain. Bar = 25 μm. B, no hybridization signal is apparent on nuclei (arrows) in a comparable field from a successive tissue section of spleen when the digoxigenin-labeled riboprobe transcribed from the EPICENTRE T7 control template was used. Permanent Red substrate and Gill's hematoxylin counterstain. Bar = 25 μm.
Discussion
To the authors' knowledge, the current study provides the first description of Siadenovirus infection in reptiles, and the first published report of adenoviral infection in chelonians. Adenoviral infections in reptiles have been associated with enteritis, hepatitis and hepatic necrosis, esophagitis, splenitis, and encephalopathy. 24,28,34,41,43,44,52 Adenoviral infections are often opportunistic in many species. Different stressors such as environmental or nutritional inadequacies or underlying disease can play a role in the development of clinical infection. The pathogenicity of reptilian adenoviruses may vary, and concurrent disease and/or immunosuppression may increase the likelihood of severe systemic disease. 27 The Siadenovirus characterized in the present report caused severe systemic disease in Sulawesi tortoises. Intranuclear inclusions were found in most of the organs examined, including the bone marrow. The adenoviral inclusions were endotheliotropic and epitheliotropic, as well as myelotropic. The extensive bone marrow involvement seen in these cases, which is not typical of adenoviral infection, suggests this virus may be immunosuppressive.
The genus Siadenovirus is less well understood than other recognized adenoviral genera. In avian species, Siadenovirus, represented by hemorrhagic enteritis of turkeys and marble spleen disease of pheasants, are consistently more pathogenic than other adenoviral genera. Identification of adenoviral types and species can provide useful diagnostic, prognostic, and epidemiologic information for clinicians. The AdVs are a diverse group; in the best-studied host species (i.e., humans), 52 adenoviral types classified into 6 species are currently recognized. 32 Significant clinical differences may exist between viral types from the same species. This is illustrated by the clinical difference between infection with the more virulent Canine adenovirus 1 (CAdv-1), which often results in serious systemic disease, and the less virulent CAdv-2, which may cause a milder upper respiratory tract disease or diarrhea. 37
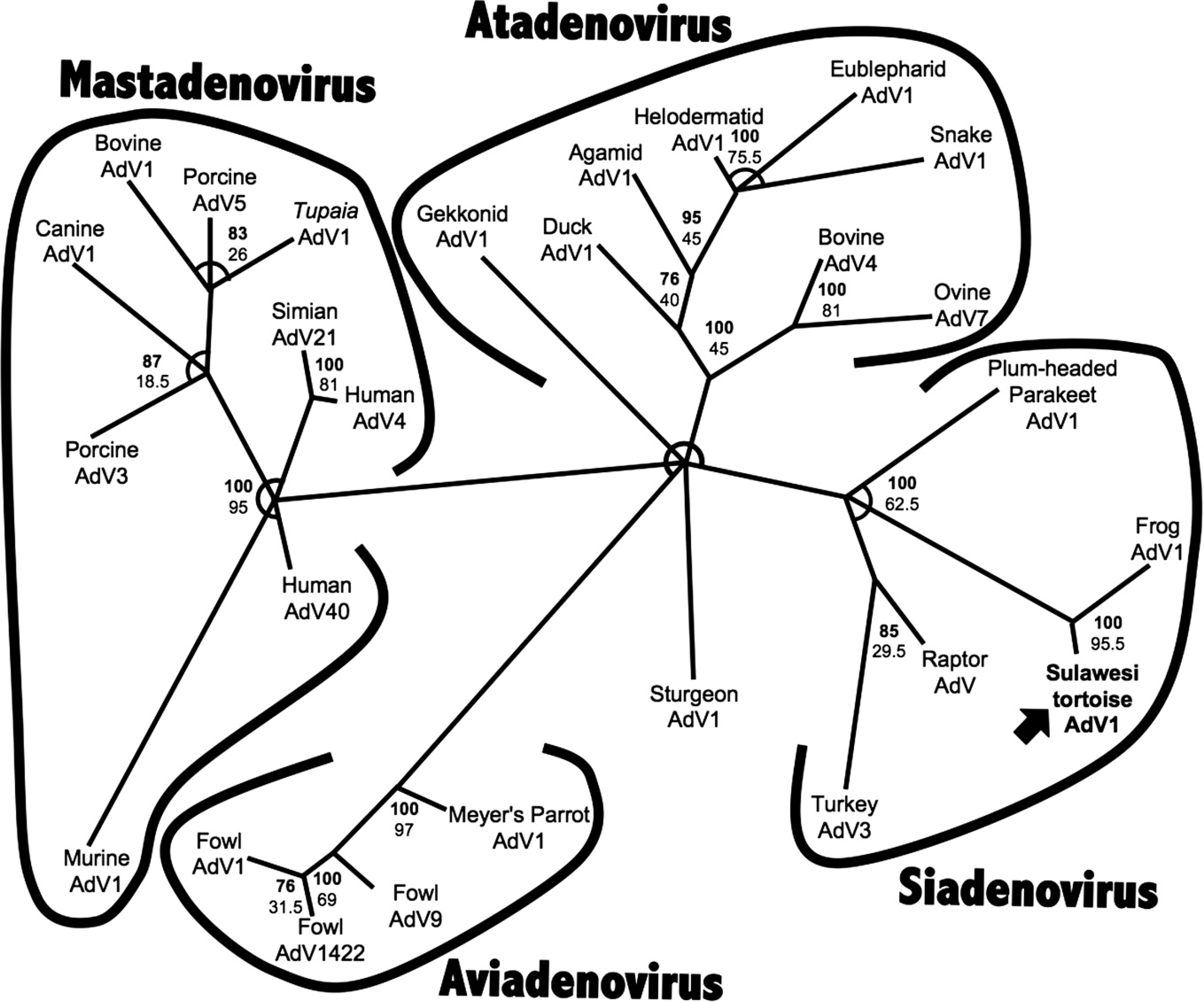
Bayesian phylogenic tree of partial adenoviral DNA polymerase amino acid sequences using T-COFFEE alignment. Sulawesi tortoise adenovirus 1 is in boldface with an arrow. Sturgeon adenovirus 1 was used as an out-group of the unrooted tree. Bayesian posterior probability values for branchings are in boldface, and bootstrap values for the maximum likelihood tree are shown. Branchings with Bayesian posterior probability values <70 are not shown, and areas where multi-furcations occurred are shown as arcs.
The prevalence of adenoviral infection and disease in chelonians is not known. Adenoviruses have been found to be the most common cause of infectious conjunctivitis in humans. 26,53 With the advent of improved antemortem diagnostics, a marked increase in diagnoses of human adenoviral disease has occurred. 45 Few attempts have been made at diagnosis of adenoviral infection in chelonians. It is likely that increased surveillance would result in the recognition of additional disease syndromes and associated adenoviral types.
In situ hybridization experiments utilizing a digoxigenin-labeled riboprobe transcribed from the cloned Sulawesi AdV DNA–dependent DNA polymerase gene PCR product gave hybridization signal from tissues in initial experiments. Among the tissues tested by ISH were sections of spleen from 2 tortoises previously determined by transmission electron microscopy to have adenovirus particles in cells of the splenic parenchyma. Hybridization signal was obtained from cells in sections from these spleens and provided associative evidence that the source of the amplified adenoviral sequence from tissue samples is the adenovirus particle identified by transmission electron microscopy. Further ISH optimization using this riboprobe is necessary because of varying hybridization signal intensity among runs. The adenoviral infection in these tortoises was not detected in separate ISH experiments using the FN-23a/FN-96 oligonucleotide DNA probe cocktail that had been used previously to successfully detect adenoviral infection in squamates. 41 As more sequence data become available, the generation of additional probes may greatly improve the ability to develop sensitive and specific diagnostic ISH techniques useful for the detection of this virus.
In addition to the adenoviral infection, the following other pathogens were identified in the affected tortoises: amoeba, nematodes, E. coli, A. hydrophila, and Chlamydophila sp. Immunosuppression, as a result of multiple factors (malnutrition, stress, illness, and possibly adenoviral infection), may have contributed to the colonization of multiple organs by these diverse pathogens. One of the most obvious clinical signs seen in the sick tortoises was chronic rhinosinusitis with oronasal fistulae and ulceration of the tongue and oral mucous membranes. These lesions have been associated with intranuclear coccidiosis, mycoplasmosis, and Tortoise herpesvirus 1 infections in tortoises. 6,25,40 In the current study, none of these pathogens were recognized as a causative agent of the chronic rhinosinusitis and oronasal fistulae.
Concurrent disease processes (e.g., amoebiasis, nematodiasis, sepsis) and postmortem tissue autolysis precluded detailed microscopic evaluation of cellular architecture in many cases, making it difficult to histologically assess the primary cause(s) of death versus contributory or secondary lesions. Most deaths occurred during the first 2–3 weeks after acquisition. It is possible that extensive tissue damage associated with amoebic enterocolitis exposed deeper tissues and facilitated viral spread to other organs. Medical management appeared to be partially effective in ameliorating these disease processes, as tortoises that died later in the course of treatment only showed evidence of the adenoviral infection.
In the present study, viral DNA was detected in nasal flush, choanal and cloacal samples, and plasma antemortem. Plasma is generally not an ideal sample for the detection of adenoviral DNA. Because of the overwhelming systemic infection seen in these cases, the viral DNA was present at detectable levels in the general circulation. The 3 animals that tested negative for viral DNA in plasma had no other samples submitted for analysis. However, all nasal flush and choanal and choanal/cloacal swabs submitted were positive. Until more is known about Siadenovirus infection in chelonians, it may be prudent to submit both plasma and cloacal samples for testing via PCR.
Owing to the lack of published CBC and plasma biochemistry reference intervals for the Sulawesi tortoise, minimum to maximum values obtained from healthy, captive, Sulawesi tortoises were used for comparison with the laboratory results of ill tortoises. The results of the CBC and plasma biochemical analyses of the ill tortoises were nonspecific and must be interpreted within the constraints of limited reference data for the species. Nonetheless, the observed changes were consistent with chronic illness. The abnormalities seen in the CBC results from many of the sick tortoises may have been an indication of a chronic inflammatory response. 47 The elevated aspartate aminotransferase and creatinine phosphokinase activities were most likely secondary to damage of muscle and possibly liver. The elevated BUN concentrations may have been caused by moderate to severe dehydration. Hyperkalemia also could have been a reflection of dehydration and decreased renal excretion, or it could have been due to massive or widespread tissue necrosis with release of intracellular potassium.
The neoplastic changes seen in several tortoises in the current study merit further investigation. Any potential correlation between adenoviral infection and later development of neoplasia in surviving cases remains to be elucidated. Although siadenoviruses are not considered to be oncogenic, viral-associated immunosuppression (through myeloid necrosis or other mechanisms) could decrease immune surveillance, allowing cells transformed by DNA mutations or retrovirus infection to proliferate.
The international trade of chelonians for the pet trade and Asian food and traditional medicine markets has been well documented. 17,36 The stress of capture and prolonged housing in substandard conditions, accompanied by poor nutrition, greatly contributes to debilitation and disease. Animals in such condition are the most likely to succumb to newly introduced pathogens. However, little is known about the spread of pathogens to naïve populations associated with such chelonian movements. At the time of the current submission, 13 animals survive and are clinically stable, but sporadic deaths without premonitory clinical signs continue. This study extends the known reptilian adenoviruses to the chelonians and extends the known genera of Adenoviridae beyond Atadenovirus to include the genus Siadenovirus.
Acknowledgements
The authors gratefully acknowledge the efforts of the Turtle Survival Alliance, a global nongovernmental organization committed to chelonian conservation, the diligent husbandry care of numerous caretakers, and the skilled technical assistance of laboratory personnel at the participating institutions.
Footnotes
a.
PELCO® 3440, PELCO ColdSpot®, PELCO Prep-Eze™, Ted Pella Inc., Redding, CA.
b.
London Resin Company Ltd., Reading, Berkshire, UK.
c.
Philips 300 electron microscope, Philips Electron Optics, Eindhoven, The Netherlands.
d.
DNeasy® Kit, QIAquick® Gel Extraction Kit, Qiagen, Valencia, CA.
e.
Big-Dye® Terminator v3.1 Cycle Sequencing Kit, Applied Biosystems® 3130 Genetic Analyzer, Applied Biosystems, Branchburg, NJ.
f.
MWG Biotech, High Point, NC.
g.
TOPO TA Cloning Kit, Invitrogen Corp., Carlsbad, CA.
h.
EPICENTRE Biotechnologies, Madison, WI.
i.
Roche Diagnostics Corp., Indianapolis, IN.
j.
ProbeOn Plus™, Fisher Biotech, Pittsburgh, PA.
k.
Ambion®, Applied Biosystems, Foster City, CA.
l.
Sigma-Aldrich, St. Louis, MO.
m.
Dako North America Inc., Carpinteria, CA.
