Abstract
The lipopolysaccharide, also known as the somatic antigen or O-antigen, is an important virulence factor of Pasteurella multocida. In the current study, the genes involved in the biosynthesis of the outer core region of the lipopolysaccharide, which were obtained from somatic type reference strains and field strains of P. multocida, were subjected to polymerase chain reaction–restriction fragment length polymorphism (PCR-RFLP) analysis. The PCR-RFLP analysis classified 11 out of the 16 serotypes into 5 PCR-RFLP types (I–V). Types I and V contain strains belong to serotypes 1 and 13, respectively. The rest of the PCR-RFLP types contain strains belong to certain groups of serotypes. Typing of 38 field strains from poultry using PCR-RFLP analysis and the gel diffusion precipitation test showed consistent results. These results indicate that the PCR-RFLP analysis can be a useful tool for rapid somatic typing of some strains of P. multocida.
Pasteurella multocida is an important pathogen of domestic animals. It is the causative agent of fowl cholera in domestic birds, hemorrhagic septicemia in cattle, and atrophic rhinitis in pigs. 10 Human infections with P. multocida arise largely from the bite of an infected carnivore, but other types of infection are reported occasionally. 9 Of the 5 capsular serogroups (A, B, D, E, and F) 13 and 16 somatic serotypes, 8 fowl cholera is mainly caused by P. multocida of serotypes A:1, A:3, and A:3,4 strains, 5 hemorrhagic septicemia in cattle is caused by serotype B:2 strains, 3 and porcine atrophic rhinitis is frequently caused by capsular serogroups D or A strains that produce dermonecrotic toxin. 4,14 The somatic antigen, also known as the lipopolysaccharide (LPS) or O-antigen, is one of key virulence factors of P. multocida. The method used most commonly to determine the somatic serotype of P. multocida is the gel diffusion precipitation test (GDPT). 8 However, the GDPT involves the preparation of somatic antigens and serotype-specific antisera, and such preparations are time-consuming and labor-intensive.
The LPS consists generally of 3 parts: lipid A, the core-oligosaccharide, and the O-polysaccharide. 11 The core-oligosaccharide is divided further into 2 parts: the inner core region and the outer core region. The LPS of P. multocida does not contain the O-polysaccharide, and is thus called rough LPS. 12 The chemical structures of the core-oligosaccharide expressed by 5 strains of P. multocida have been determined. The names and serotypes of these strains are VP161 (A:1), X-73 (A:1), Pm70 (A:3), M-1404 (B:2), and P-1702 (A:5). 6,15-18 Structural analysis has revealed that the inner core region expressed by these strains is conserved, whereas the outer core region is variable and serotype-specific. The genes required for the assembly of the outer core region of strains VP161, Pm70, M-1404, and P-1702 have been determined. These genes are clustered at a single genetic locus between the conserved priA and fpg genes, which are not involved in the synthesis of LPS. 2,7,15,18 Moreover, the hptE (pm1144) gene, which is adjacent to the fpg gene, is also conserved. In the present study, polymerase chain reaction (PCR) was used to amplify DNA fragments that contained the genes between the priA and hptE genes from somatic serotype reference strains and field strains of P. multocida. A procedure for PCR–restriction fragment length polymorphism (PCR-RFLP) analysis was developed for rapid somatic typing of P. multocida.
Fifty-three strains of P. multocida were used in the current study. The strains included the somatic type reference strains 8 and field strains isolated from cases diagnosed as fowl cholera in Taiwan in 1960–1990 and 1995–2005 (Table 1). Note that reference strains of somatic serotypes 6–9 and 14 were not included in the study owing to loss of bacterial viability during storage. All strains were grown at 37°C in brain–heart infusion broth. a Bacterial DNA was isolated using a commercial kit. b
Two primers (PF28 and 1144R) were designed on the basis of the conserved sequences present in priA and hptE (Fig. 1A). The 2 primers were used to PCR-amplify a DNA fragment that contained the genes required for the assembly of the outer core region of P. multocida. The sequences of PF28 and 1144R were 5’-GCCACACGGATGAGTTGCAT-3’ and 5’-CTTGGCATTGATTAAGCGTA-3’, respectively. The PCR was performed in a thermal cycler. c The PCR mixture (30 µl) contained 21.3 µl of water, 3 µl of 10 × buffer, 3 µl of deoxyribonucleotide triphosphate [dNTP] blend (2.5 mM each dNTP), 2 µl of primers (10 picomole each), 0.5 µl of template DNA, and 0.2 µl of high fidelity Taq XL DNA polymerase. d The PCR conditions were: 1 cycle of 94°C for 2 min, 35 cycles of 94°C for 30 sec, 56°C for 30 sec, and 68°C for 6 min, and 1 cycle of 25°C for 1 min.
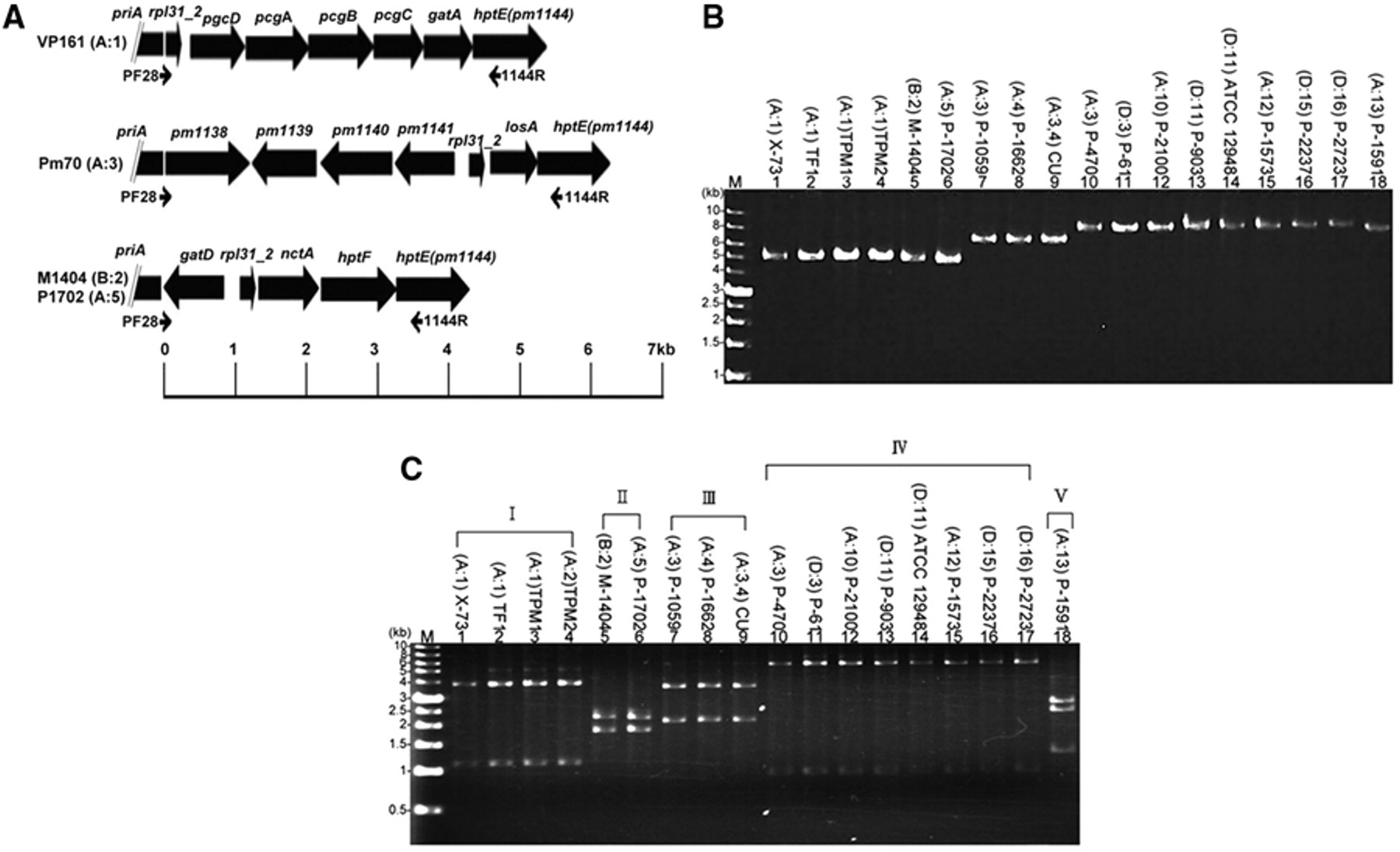
Eighteen strains were initially selected for PCR-RFLP analysis. These strains were selected because they were somatic type reference strains or representatives of field strains isolated during different time periods. The PCR with primers PF28 and 1144R was able to amplify a DNA fragment of 4.2–6.8 kb in size from these strains (Fig. 1B). The PCR product (4 µl) was digested by the addition of 4 µl of water, 1 µl of 10× buffer, e and 1 µl (10 U) of restriction enzyme HincII, e and incubated at 37°C for 2 hr. The digested DNA was assayed in a 0.8% agarose slab gel and then visualized under ultraviolet light after staining with ethidium bromide solution (0.5 pg/ml). The results showed that the 18 strains could be classified into 5 PCR-RFLP types (I–V) according to the patterns observed (Fig. 1C). Type I contained the somatic type reference strain X-73 (A:1) and Taiwan field strains TF1 (A:1), TPM1 (A:1), and TPM2 (A:1). Type II contained somatic serotype reference strains M-1404 (B:2) and P-1702 (A:5). Strains M-1404 and P-1702 are known to contain a similar LPS structure. 12,15 Type III contained the somatic serotype reference strains P-1059 (A:3) and P-1662 (A:4), and the attenuated Clemson University (CU) vaccine strain (A:3,4). Type IV contained strains P-470 (A:3) and P-61 (D:3), and reference strains of somatic serotypes 10–12, 15, and 16. Type V contained a somatic serotype reference strain P-1591 (A:13).
The PCR products shown in Figure 1A were subjected to sequence analysis. Primers PF28 and 1144R, together with a total of 79 “walking primers,” were used to conduct sequence analysis. All sequencing studies were performed using a commercial sequencing kit c in an automatic sequencer. c The sequences were compiled using the Seqman program in the LASERGENE package. f The nucleotide sequences determined in the present study are available in GenBank under the accession numbers HQ162664–HQ162679. Pairwise comparison of the nucleotide sequences showed that the sequences from strains of the same PCR-RFLP type were very similar to each other (96.8–100% identity), whereas the sequences from strains of different PCR-RFLP types were diversified (42.7–72.9% identity). The sequences determined were used to predict the sizes of the restriction fragments generated with HincII digestion. The predicted sizes are 1.1 and 3.8 kb (type I), 0.2, 1.8, and 2.2 kb (type II), 2.1 and 3.6 kb (type III), 0.9 and 5.9 kb (type IV), and 1.2, 2.4, and 2.7 kb (type V). These predicted sizes matched those observed experimentally in RFLP analysis (Fig. 1C). BLAST search showed that the nucleotide sequences of the strains of type I were very similar (99.3–100% sequence identity) to that of VP161 (A:1). 7 This suggested that VP161 was also a strain of type I. The nucleotide sequence of the strains of type II have been reported previously, 15 and thus were not investigated in the current study. The nucleotide sequences of the strains of type III were very similar (96.8–99.3% sequence identity) to that of Pm70. 18 This result suggested that Pm70 is a strain of type III. The nucleotide sequences of the strains of types IV and V showed limited homology to sequences deposited in the current GenBank.
Thirty-eight field strains (TF1–TF12 and TPM1–TPM26) isolated from poultry in Taiwan (Table 1) were subjected to PCR-RFLP typing. The result showed that 37 of the 38 strains showed a PCR-RFLP pattern identical to that of type I, whereas 1 strain (TF3) showed a pattern identical to type III. This result suggested that 37 of the 38 strains belonged somatic serotype 1, whereas TF3 to serotypes 3, 4, or 3,4. Somatic typing with the GDPT showed that the antigen prepared from each of the 37 strains formed a precipitate with antisera against X-73 (serotype 1) but not with antisera against P-1059 (serotype 3), whereas antigen prepared from TF3 showed the reverse. This observation indicated that typing with PCR-RFLP and the GDPT gave a consistent result.
In conclusion, the data presented in the present study shows that somatic type reference strains of P. multocida could be classified into 5 PCR-RFLP types. Although only 11 out of the 16 somatic serotype reference strains were classified, the non-tested 5 serotypes were rarely found in strains associated with diseases of animals. Given that each type showed a unique RFLP pattern, this pattern could be used to identify and differentiate some somatic serotypes of P. multocida. Typing of 38 strains from poultry with PCR-RFLP analysis and GDPT showed a consistent result that 37 of them belonged to serotype 1. This highlights the potential to use PCR-RFLP to identify the strains of serotype 1 isolated from poultry. Other somatic serotypes that could be identified by PCR-RFLP analysis were serotypes 2 and 5 (type II), serotypes 3, 4, and 3,4 from poultry (type III), and serotype 13 (type V). Although a PCR-RFLP method has been reported previously to distinguish P. multocida serotypes A:1, A:3, and B:2, 1 the target gene for this PCR-RFLP analysis is the ompH gene, which is not related to the synthesis of LPS.
Characteristics and sources of Pasteurella multocida strains used in the present study.*
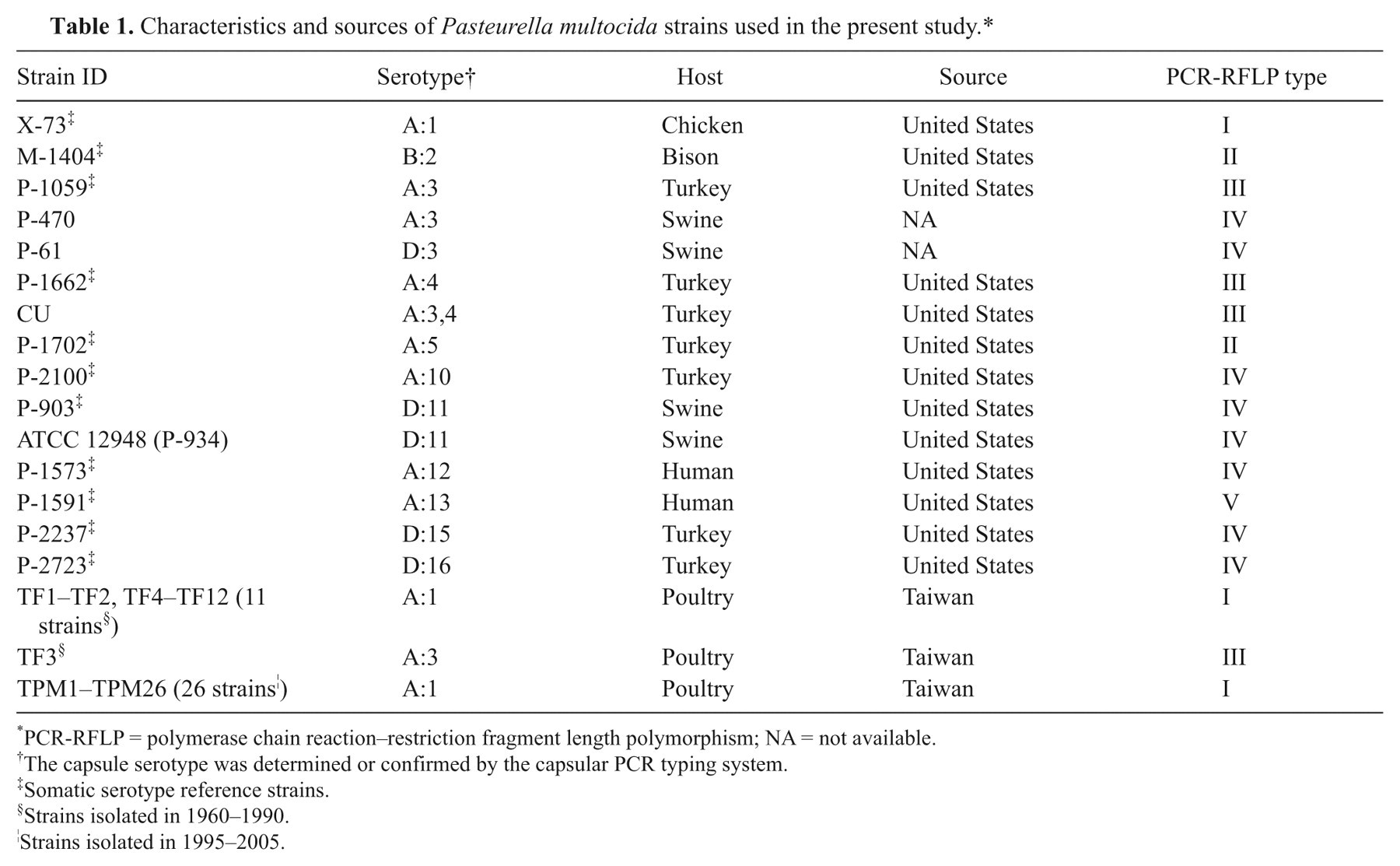
PCR-RFLP = polymerase chain reaction–restriction fragment length polymorphism; NA = not available.
The capsule serotype was determined or confirmed by the capsular PCR typing system.
Somatic serotype reference strains.
Strains isolated in 1960–1990.
Strains isolated in 1995–2005.
Footnotes
a.
Difco, Becton Dickinson, Sparks, MD.
b.
Qiagen Inc., Valencia, CA.
c.
Applied Biosystems, Foster City, CA.
d.
Protech Technology Enterprise Co. Ltd. Taipei, Taiwan
e.
New England Biolabs, Inc. Beverly, MA.
f.
DNASTAR Inc., Madison, WI.
The authors declared that they had no conflicts of interest with respect to their authorship or the publication of this article.
This investigation was supported by grant 97-2313-B-005-008-MY3 from the National Science Council, Taiwan, Republic of China.
