Abstract
A set of polymerase chain reaction (PCR) assays for identification of the most important Pasteurellaceae species encountered in cats and dogs were developed. Primers for Pasteurella multocida were designed to detect a fragment of the kmt, a gene encoding the outer-membrane protein. Primers specific to Pasteurella canis, Pasteurella dagmatis, and Pasteurella stomatis were based on the manganese-dependent superoxide dismutase gene (sodA) and those specific to [Haemophilus] haemoglobinophilus on species-specific sequences of the 16S ribosomal RNA gene. All the primers were tested on respective reference and control strains and applied to the identification of 47 canine and feline field isolates of Pasteurellaceae. The PCR assays were shown to be species specific, providing a valuable supplement to phenotypic identification of species within this group of bacteria.
Bacteria belonging to the family Pasteurellaceae are small Gram-negative bacilli or coccobacilli that inhabit the mucosal membranes of many animals. 1 Some members of this group are known as primary or opportunistic pathogens of their hosts and may account for significant economic losses, especially in cattle and poultry farming. 21 Several species of Pasteurellaceae, mainly Pasteurella multocida, Pasteurella canis, Pasteurella stomatis, Pasteurella dagmatis, and [Haemophilus] haemoglobinophilus, may be involved in a range of pathological conditions in dogs and cats. 13,21 These bacteria are also of epidemiological significance as they may cause human infections, particularly as a result of bite wounds or inhalation 9,12,22 ; thus, a rapid and accurate identification of such isolates may be important from both veterinary and medical points of view.
The conventional identification of Pasteurellaceae remains difficult for many laboratories. Phenotypic characterization requires many biochemical and enzymatic tests, and the results are sometimes inconclusive or misleading. 3,11 DNA-based methods provide a useful alternative for the phenotypic identification of these bacteria. For years, 16S ribosomal RNA gene (rrs) sequence–based identification has been a powerful method in bacterial diagnostics as well as in phylogenetic studies on Pasteurellaceae. 5,6,15 More recently, sequence analysis of the sodA, a housekeeping gene that encodes manganese-dependent superoxide dismutase, has been successfully used in the diagnostics of various bacteria, including Pasteurellaceae. 10 However, sequencing methods are often too expensive for routine use. A combination of both phenotype- and genotype-based methods may offer the optimal solution in cases dealing with a large group of closely related bacteria such as Pasteurellaceae. 4 The already existing polymerase chain reaction (PCR) methods focus mainly on detection and identification of P. multocida, especially in farm animals. 14,23 Considering the economic aspect of bacterial diagnostics and the fact that so far there is no comprehensive PCR-based approach to identifying Pasteurellaceae species encountered in dogs and cats, the present study aimed at developing PCR assays that allow accurate recognition of clinically relevant canine and feline isolates within this group of bacteria. Such PCR-based methods might be a valuable supplement to phenotypic diagnostics.
A total of 47 field strains belonging to the family Pasteurellaceae as well as 12 reference and control strains were investigated (Table 1). The field strains originated from both diseased and healthy dogs and cats from western and southern Poland. All isolates were grown on 5% sheep blood trypticase soy agar, a under aerobic conditions, and subsequently identified phenotypically and genotypically. The following biochemical tests were included in the study: urease test (on Christensen medium supplemented with liver digest and glucose), 8 indole test (in tryptophan broth [beef extract, b 3.0 g, bacto-tryptone, b 10.0 g, NaCl 5.0 g, distilled water 1 liter] with subsequent addition of Ehrlich reagent), ornithine decarboxylase test (ODC), c and production of acid or gas from glucose, sucrose, mannose, maltose, mannitol, sorbitol, and trehalose (in cystine trypticase agar [CTA] medium, d supplemented with 1% of the respective carbohydrate). Results of the indole test were read after overnight incubation, and the remaining tests were observed for up to 3 days. Phenotypic identification was performed as previously described. 7,8
Bacterial strains used in the current study and their characterization.*
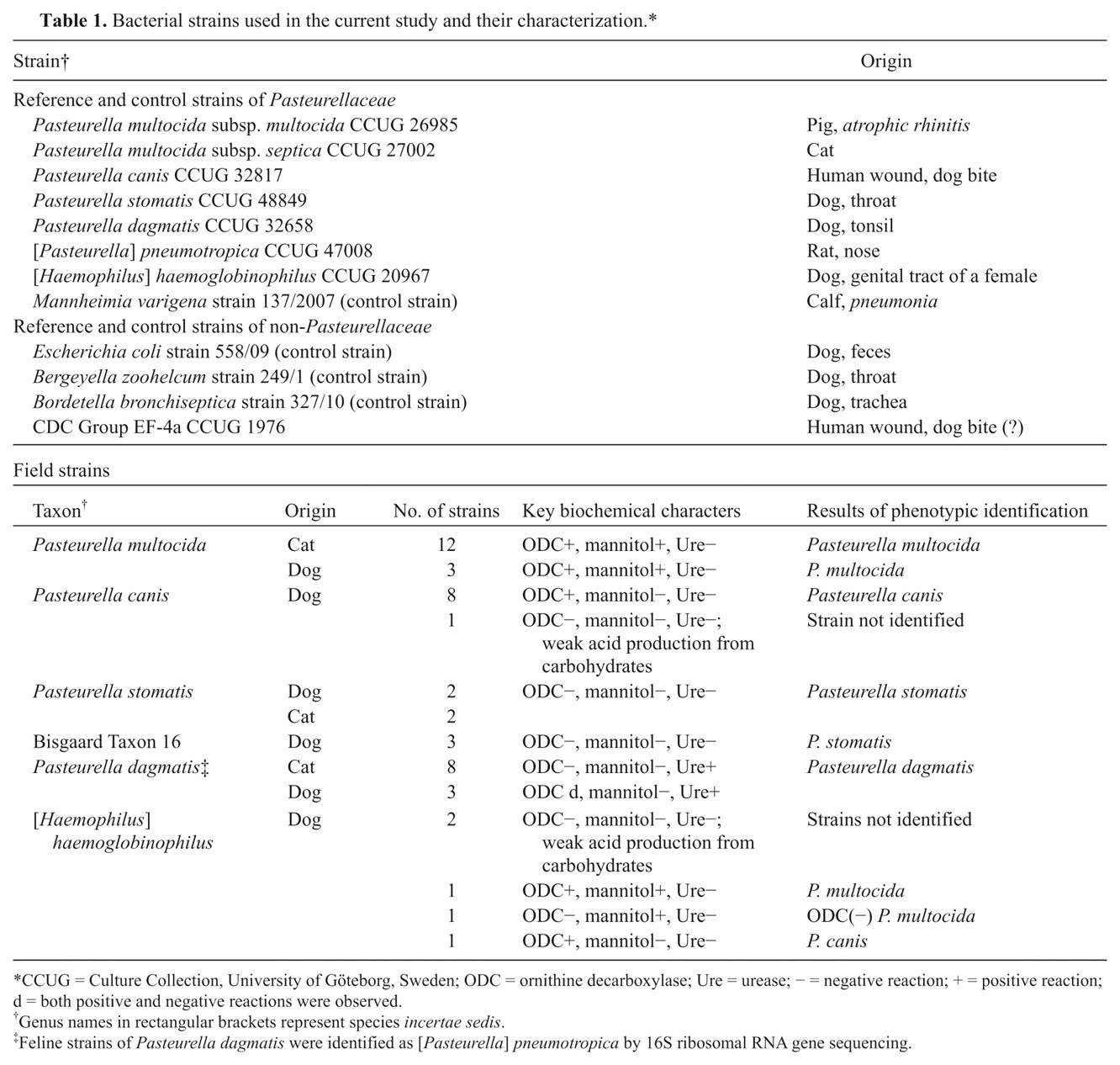
CCUG = Culture Collection, University of Göteborg, Sweden; ODC = ornithine decarboxylase; Ure = urease; − = negative reaction; + = positive reaction; d = both positive and negative reactions were observed.
Genus names in rectangular brackets represent species incertae sedis.
Feline strains of Pasteurella dagmatis were identified as [Pasteurella] pneumotropica by 16S ribosomal RNA gene sequencing.
Bacterial DNA was extracted from overnight blood agar cultures using a genomic DNA kit, e according to the manufacturer’s instructions. All field strains were analyzed by sequencing a 918-bp fragment of the rrs gene using the primers 16S-27f (5’-AGAGTTTGATCMTGGCTCAG-3’) and 16S-907r (5’-CCGTCAATTCMTTTRAGTTT-3’; http://rdna4.ridom.de/static/primer.html). The reaction mixture (25 µl) contained 10 mmol/l of Tris-HCl (pH 8.8), 1.5 mmol/l of MgCl2, 50 mmol/l of KCl, 0.08% Nonidet P40, f 5 pmol of each primer, g 0.2 mmol/l of each deoxyribonucleotide, f 2 U of Taq DNA polymerase, f and 2 µl of DNA. Forty PCR cycles of denaturation at 94°C for 30 sec, annealing at 50°C for 30 sec, and elongation at 72°C for 120 sec were performed. The PCR product was purified by adding 10 U of Escherichia coli–exonuclease I and 2 U of shrimp alkaline phosphatase f to 5 µl of the reaction mixture and subsequently sequenced using a commercial cycle sequencing kit. h
For P. multocida–specific PCR, 2 sets of oligonucleotide primers were used: 1) a pair of primers designated KMT1T7 and KMT1SP6 described previously 23 and 2) a pair of newly designed primers, KMTJB-for and KMTJB-rev, obtained by searching all DNA sequences of P. multocida (available in GenBank) encoding the outer-membrane protein (KMT-1). The conserved KMT-1 regions were selected on the basis of multiple alignment performed with BioEdit software (mbio.ncsu.edu/BioEdit/bioedit.html). The design of primers for the specific detection of P. canis, P. stomatis, P. dagmatis, and [P.] pneumotropica was performed by comparison of available sequences of the sodA gene of these species using BioEdit software. The primers were designed using Beacon Designer 4.0 software. i Primers specific to [Haemophilus] haemoglobinophilus were designed on the basis of rrs gene sequences available in GenBank and of those obtained from the authors’ [H.] haemoglobinophilus isolates. All new primers used in the present study are listed in Table 2. The amplification was performed in a mixture (25 µl) containing 10 mmol/l of Tris-HCl (pH 8.8), 1.5 mmol/l of MgCl2, 50 mmol/l of KCl, 0.08% Nonidet P40, f 20 pmol of each primer, 0.2 mmol/l of each deoxyribonucleotide, f 1 U of Taq DNA polymerase, f and 1 µl of DNA. Thirty-five PCR cycles of denaturation at 94°C for 30 sec, annealing at 52°C (for detection of P. stomatis and [H.] haemoglobinophilus) or 57°C (in the case of remaining species-specific PCR assays) for 30 sec, and elongation at 72°C for 30 sec were performed. The PCR products (7.5 µl) were stained with ethidium bromide, electrophoresed in 2% agarose gel, and visualized on a gel documentation system. j
Primers used for detection of Pasteurella multocida, Pasteurella canis, Pasteurella stomatis, Pasteurella dagmatis, [Pasteurella] pneumotropica, and [Haemophilus] haemoglobinophilus.*
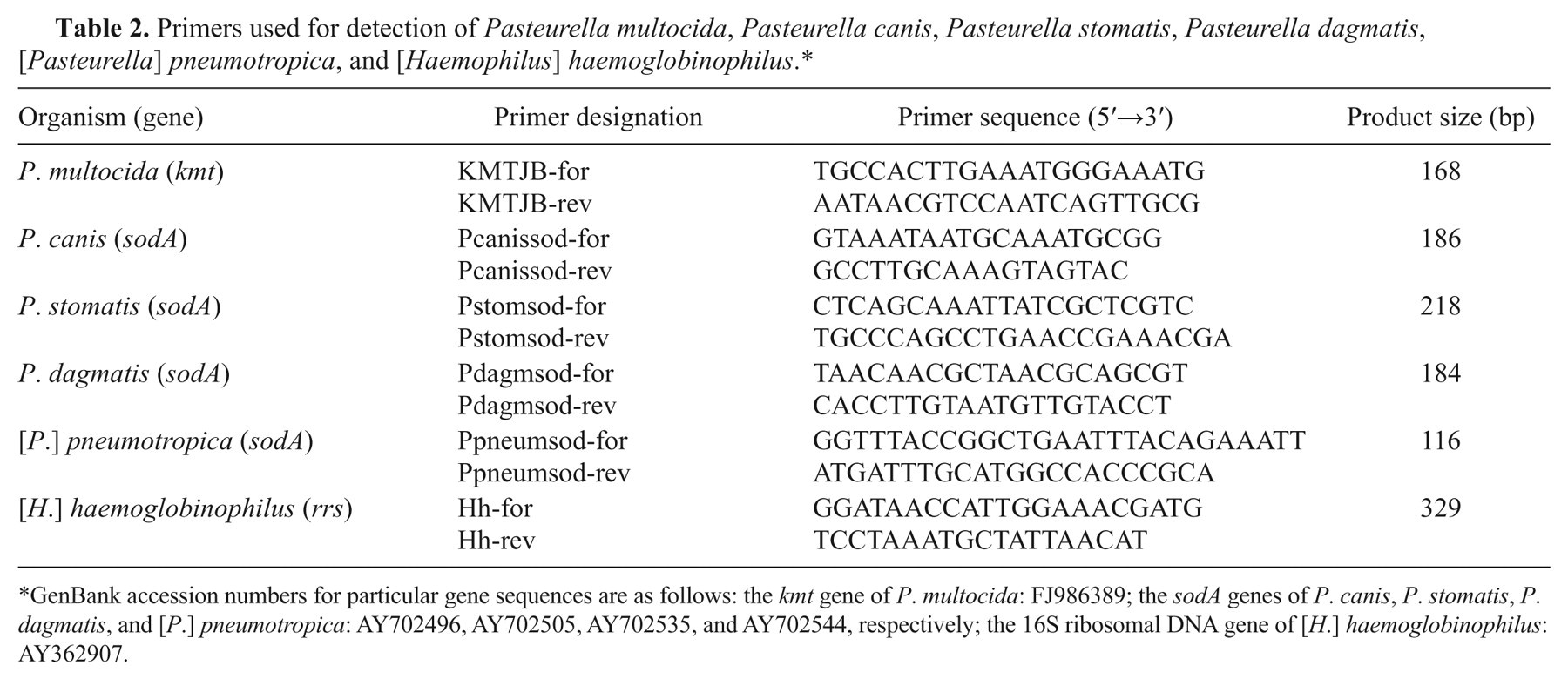

GenBank accession numbers for particular gene sequences are as follows: the kmt gene of P. multocida: FJ986389; the sodA genes of P. canis, P. stomatis, P. dagmatis, and [P.] pneumotropica: AY702496, AY702505, AY702535, and AY702544, respectively; the 16S ribosomal DNA gene of [H.] haemoglobinophilus: AY362907.
Accurate phenotypic identification at the species level, confirmed by sequencing of the rrs gene, was possible only with the strains belonging to P. multocida and canine strains of P. dagmatis. All P. multocida strains were positive in the ODC, mannose, sucrose, and mannitol tests, whereas the ability to produce acid from maltose, sorbitol, and trehalose varied among the strains. The primers KMT1T7–KMT1SP6, previously designed for detection of P. multocida, 23 were able to produce a 460-bp amplicon only in 6 (35.3%) of the P. multocida strains (including the reference strains). This might be attributed to genetic differences among isolates of various geographical and host origin. Using primers KMTJB-for and KMTJB-rev, all strains assigned to P. multocida and the reference P. multocida strains gave a PCR product of 168 bp (Fig. 1). No products were found with the remaining Pasteurellaceae isolates examined.
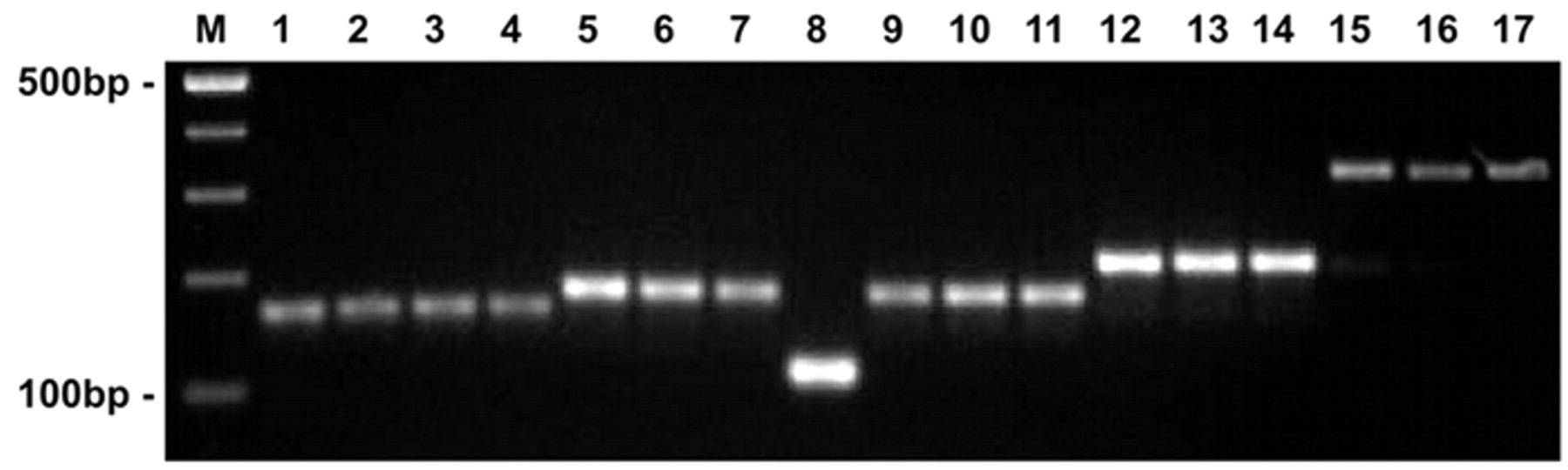
Agarose gel electrophoresis of polymerase chain reaction products amplified from DNA of various Pasteurellaceae strains tested in the present study. Lanes 1–4: Pasteurella multocida, amplicon size of 168 bp (lane 1: P. multocida multocida Culture Collection, University of Göteborg, Sweden [CCUG] 26985; lane 2: P. multocida septica CCUG 27002; lanes 3, 4: field isolates); lanes 5–7: Pasteurella canis, amplicon size of 186 bp (lane 5: P. canis CCUG 32817; lanes 6, 7: field isolates); lane 8: [Pasteurella] pneumotropica CCUG 47008, amplicon size of 116 bp; lanes 9–11: Pasteurella dagmatis, amplicon size of 184 bp (lane 9: P. dagmatis CCUG 32658; lanes 10–11: field isolates); lanes 12–14: Pasteurella stomatis, amplicon size of 218 bp (lane 12: P. stomatis CCUG 48849; lanes 13, 14: field isolates); lanes 15–17: [Haemophilus] haemoglobinophilus, amplicon size of 329 bp (lane 15: [H.] haemoglobinophilus CCUG 20967; lanes 16, 17: field isolates). M = 100-bp size marker. f
Pasteurella dagmatis was identified phenotypically based on its ability to produce urease (positive reaction in all strains investigated). In the case of the feline strains of P. dagmatis, the phenotypic identification was not consistent with the results of rrs gene sequencing. All the feline isolates revealed the partial nucleotide sequence of the rrs gene identical to that of the [P.] pneumotropica National Collection of Type Cultures (NCTC) 10827 strain (GenBank accession no. AF224296) and only 97% homology with the available sequences of P. dagmatis. By contrast, the rrs gene sequence of the canine isolates displayed 100% similarity with the P. dagmatis AY362920 strain. All feline and canine strains tested gave appropriate PCR products with the P. dagmatis–specific primers. This fact might downgrade the usefulness of the method described in the present study. Therefore, to address the discrepancy between sequencing and PCR results, primers targeting the P. pneumotropica sodA gene were additionally designed and employed. A 116-bp PCR product was obtained only with the reference [P.] pneumotropica strain Culture Collection, University of Göteborg, Sweden (CCUG) 47008, and no amplicons were detected in the case of the canine and feline P. dagmatis isolates or in the remaining Pasteurellaceae strains tested. A review of the literature reveals that there are some doubts about [P.] pneumotropica NCTC 10827. It was found 20 that some feline isolates, described as P. dagmatis–like, are very similar (both phenotypically and genetically) to this taxon. [Pasteurella] pneumotropica NCTC 10827 is an unusual strain, even within a very heterogeneous species such as [P.] pneumotropica. 19 The rrs gene sequence of the type strain NCTC 10827 does not match any other sequence of [P.] pneumotropica deposited in GenBank, giving rise to the conclusion that it may be an atypical variant of [P.] pneumotropica or another Pasteurella species. This strain was isolated in 1971 from human blood and identified based on its biochemical properties, mainly a positive urease reaction. 18 However, the reported ability to produce gas from some carbohydrates could indicate that it might belong to P. dagmatis. Similarly, it is believed that the feline strains in the current study are a cat-associated subpopulation of P. dagmatis. The positive reaction with primers targeting a fragment of the sodA gene of this species and negative reaction with primers for P. pneumotropica could support this hypothesis. Further investigation of the strain NCTC 10827, as well as feline isolates of P. dagmatis (or P. dagmatis–like), seems to be necessary to clarify this issue.
In many cases, the phenotypic identification of the P. stomatis, P. canis, and [H]. haemoglobinophilus strains was not reliable or failed (Table 1). Pasteurella stomatis is recognized phenotypically by ruling out other species. 7 It shares many phenotypic properties with Bisgaard Taxon 16, which has also been encountered in dogs and cats. 8,16 The phenotypic differentiation of both taxa is a matter of controversy. According to some authors, 16,17 P. stomatis is maltose negative, whereas in another study, the majority of strains tested gave positive results. 8 Differentiation based on the β-galactosidase (ONPG) test may also be misleading. All Bisgaard Taxon 16 strains investigated by the latter investigators were positive with this test, but others obtained opposite results. 16
Pasteurella canis also presented some difficulties in routine phenotypic identification. Most of the P. canis strains examined herein displayed weak acid production from carbohydrates in CTA medium. The most important test for distinguishing P. canis was the positive reaction for ODC (detected in all but 1 of the isolates examined). Moreover, 1 [H.] haemoglobinophilus strain (ODC-positive and mannitol-negative) was misidentified as P. canis. [Haemophilus] haemoglobinophilus is not usually included in the routine identification of Pasteurellaceae in veterinary laboratories. 7,8 However, it has been isolated by the present authors relatively often. The strains of this species originated from both the upper gastrointestinal tract and the genitourinary tract of dogs and varied greatly in biochemical characteristics. Isolates of [H.] haemoglobinophilus are often mannitol positive 2 and can thus be erroneously classified as P. multocida.
The use of the species-specific primers developed in the present study greatly facilitated the recognition of all the species mentioned. Primers targeting the sodA gene of P. stomatis, P. dagmatis, and P. canis gave the predicted amplicons of 218, 184, and 186 bp, respectively, only with the appropriate species, and no cross-reaction with another strain was observed. Similarly, PCR for the identification of [H.] haemoglobinophilus resulted in a product of 329 bp only with the reference strain of this species and with 5 field isolates of [H]. haemoglobinophilus identified by rrs gene sequencing (Fig. 1). To expedite the laboratory work, PCR assays presented herein can be applied as a set of duplex PCR assays performed after basic biochemical identification (Table 3).
Flow chart for the identification of Pasteurellaceae encountered in cats and dogs in the current study.*
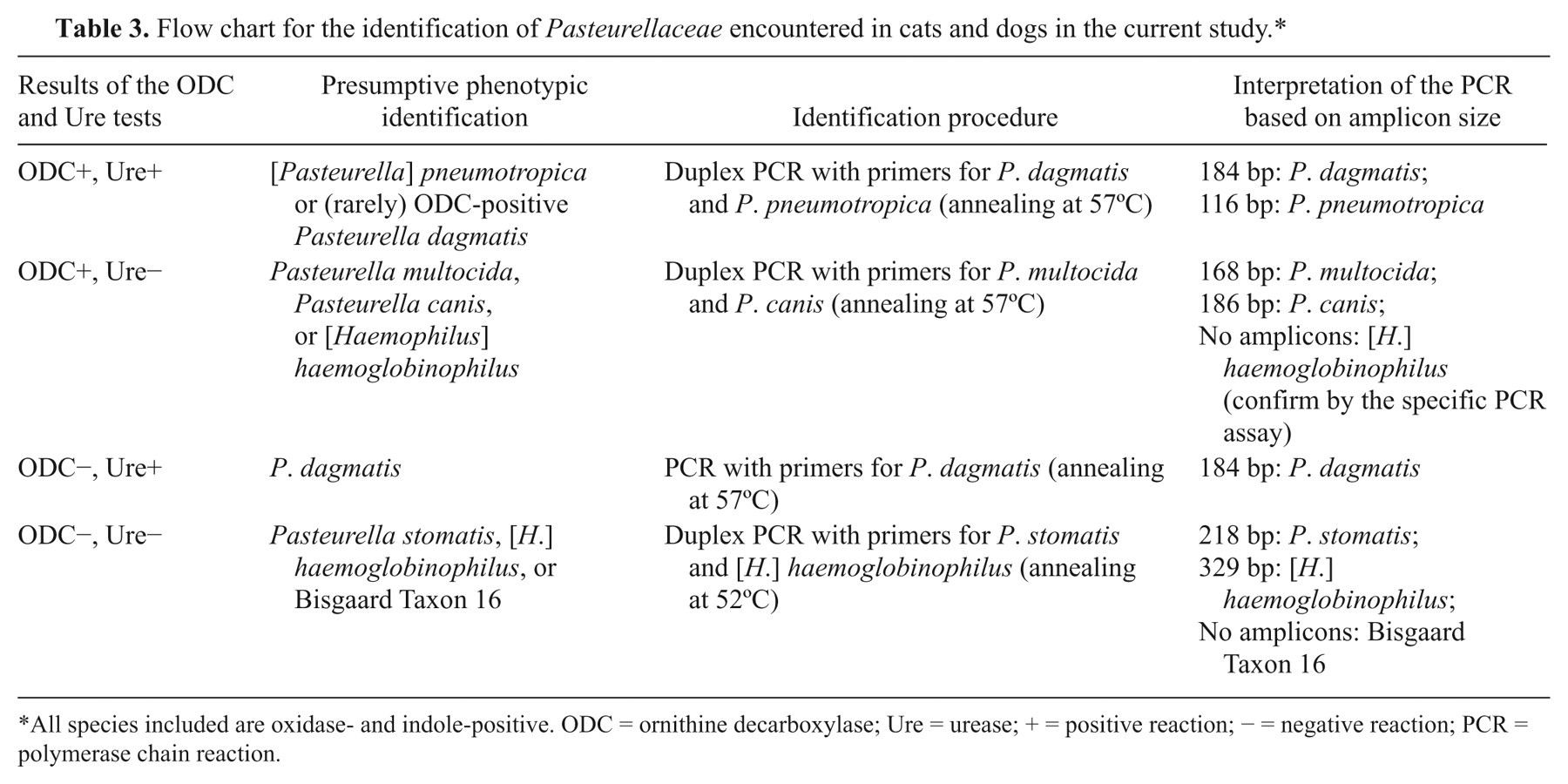

All species included are oxidase- and indole-positive. ODC = ornithine decarboxylase; Ure = urease; + = positive reaction; − = negative reaction; PCR = polymerase chain reaction.
The described method may be employed in both human and veterinary laboratories, especially when dealing with isolates that are difficult to recognize by phenotypic tests. Thorough species identification within this group of bacteria will contribute to a better understanding of the distribution and possible epidemiological significance of Pasteurellaceae in companion animals.
Footnotes
a.
BioMérieux Polska, Warsaw, Poland.
b.
Difco Laboratories, Detroit, MI.
c.
Rosco Diagnostica, Taastrup, Denmark.
d.
Becton Dickinson, Le Pont de Claix, France.
e.
A&A Biotechnology, Gdynia, Poland.
f.
Fermentas, Vilnius, Lithuania
g.
Institute of Biochemistry and Biophysics, Warsaw, Poland.
h.
ABI Prism™ DYEnamic ET terminator cycle sequencing kit, Amersham Biosciences Europe GmbH, Freiburg, Germany.
i.
Premier Biosoft International, Palo Alto, CA.
j.
GelDoc XR gel documentation system, BioRad Polska, Warsaw, Poland.
The authors declared that they had no conflicts of interest with respect to their authorship or the publication of this article.
The authors declared that they received no financial support for their research and/or authorship of this article.
