Abstract
Edema disease is an enterotoxemic disorder of weaned piglets that represents a significant threat to pig husbandry worldwide. The causative Escherichia coli strains are highly adapted to the porcine host and characterized by the production of Shiga toxin type 2e (Stx2e) and adhesive F18 fimbria. The current study assessed the occurrence of F18 fimbrial subtypes in 241 porcine stx2e + fedA + E. coli strains in Germany, including 116 Shiga toxin–encoding E. coli (STEC) and 125 Shiga toxin E. coli/enterotoxigenic E. coli (STEC/ETEC) isolates. In addition, a novel multiplex polymerase chain reaction (PCR) was developed in order to improve the typing system in terms of costs, time, and discriminative power. Utilizing the novel F18 typing PCR, 93 E. coli strains (38.5%) tested positive for the F18ab fimbrial subtype and 147 strains (61.0%) for the F18ac fimbrial subtype, while 1 strain remained nontypeable. Six strains were classified as F18ac using the F18 typing PCR, but were classified as F18ab using the F18-restriction fragment length polymorphism assay. Nucleotide sequencing of the FedA gene revealed that 5 of these strains encoded F18ac fimbriae, while the FedA of 1 strain did not cluster with F18ab or with F18ac amino acid sequences. The F18 fimbrial subtype was significantly associated with the pathovar of the E. coli strains, as 73.2% of the STEC isolates harbored F18ab genes whereas 93.6% of the STEC/ETEC isolates proved F18ac positive. In conclusion, the novel F18 typing PCR allows a specific identification of the F18 fimbrial subtype. The genetic and phenotypic heterogeneity of F18 fimbriae in porcine E. coli strains should be considered in the development of new vaccines and diagnostic tools.
Introduction
Enteric diseases in piglets induced by representatives of certain Escherichia coli pathovars are some of the most important causes for economic losses in piggeries. Shiga toxin–encoding E. coli (STEC) strains encoding the Shiga toxin type 2e (Stx2e) represent the causative agents of edema disease in weaning piglets, 23 while enterotoxigenic E. coli strains (ETEC) encoding for heat stable (STa, STb, EAST1) and/or heat labile (LT) enterotoxins cause secretory diarrhea in newborn and weaned piglets. 16,25 Interestingly, some strains harbor both the Stx2e genes and enterotoxin genes and therefore may be capable of causing symptoms of edema disease and those of diarrhea in the same animal (STEC/ETEC). 3 A common feature of porcine STEC and many porcine ETEC strains is the expression of F18 fimbriae, which facilitates bacterial colonization of the mucosal surface of the intestine. 13 After oral uptake of the bacteria, F18 fimbriae mediate adhesion of the bacterial cell to the ECF18R receptor exposed by enterocytes of the small intestine on their apical cytoplasmic membrane. 38,39 Some porcine STEC and STEC/ETEC strains can express other cytoadhesive fimbriae or nonfimbrial adhesins in addition to the F18 fimbria, in particular F4 or F5 fimbriae or the “adhesin involved in diffuse adherence” (AIDA). 3,12,16,27 While the AIDA adhesin occurs frequently in both pathovars, F4 or F5 fimbriae are rare in those strains. 15
Two subtypes of the F18 fimbria are distinguishable by serologic methods: F18ab (formerly, F107 fimbria) and F18ac (formerly, 2134P, 8813, or Av24 fimbria). 18,29,31 Both F18 fimbrial subtypes appear correlated with different pathovars and O antigens. O antigens are constituted by certain sugar residues of lipopolysaccharides present in the outer membrane of Gram-negative bacteria. Although O antigens appear not directly involved in bacterial virulence, serotyping is often used in routine diagnostics for presumptive identification of virulent strains among porcine E. coli isolates. 16 While F18ab fimbriae have been found predominantly in STEC exposing the O138 or O139 antigen on the bacterial surface, F18ac fimbriae were reported dominant in ETEC strains of serogroups O141, O147, and O157. 1,17,25,40 The F18 fimbriae are encoded in the fed gene locus, which is localized on a plasmid that often additionally harbors the nonfimbrial adhesin AIDA and the α-hemolysin gene hlyA. 24,41 The fed gene locus comprises 5 genes. The fedA gene encodes the major subunit of the fimbria, fedB the usher protein, fedC the periplasmatic chaperone, fedE a small linker protein between FedA and FedF, and fedF the fimbrial adhesin. 34,35 Knowledge about the distribution and prevalences of the fimbrial subtypes is important for the development of vaccines and diagnostic tools. In particular, it was shown that pigs vaccinated with one F18 fimbrial subtype were not or only partially protected during challenge experiments with ETEC strains expressing the other fimbrial subtype. 6,20,32
Several methods for the differentiation of the F18 subtypes have been published. The first method published was a direct agglutination assay utilizing absorbed test sera and monoclonal antibodies, respectively. 11,29 However, many strains do not or poorly express the F18 fimbriae during growth in vitro even if cultures are supplemented with alizarin yellow and eosine in a 10 vol.% CO2 atmosphere as suggested by some authors. 41
Currently, a method based on the restriction fragment length polymorphism (RFLP) of a polymerase chain reaction (PCR)-generated amplicon of the fedA gene is widely accepted for subtyping of F18 fimbriae (hereafter, F18 PCR-RFLP assay). 19 This assay takes advantage of a 3-bp insertion at positions 364–366 in the fedAac gene. This insertion constitutes an NgoMIV restriction site in fedAac, which is absent from fedAab. In addition to this F18 PCR-RFLP assay, a single-strand conformational polymorphism (SSCP) assay for analysis of the fedA genes was generated. 7 Due to a significant slower migration of 1 single strand, a group of investigators were able to differentiate between F18ab and F18ac encoding strains, and identified 19 different SSCP profiles among porcine E. coli isolates. 7 However, such method requires the use of radioactive nucleotides, which harbor health risks for laboratory workers in addition to the necessity of implementing additional precautions for the removal of waste. In 2005, a PCR typing method was published using 2 primer pairs located in the fedA gene. 8 However, such method failed to differentiate reliably between F18ab and F18ac. In another study, DNA sequencing of the fedA gene was performed. 12 But, sequencing still is too expensive to be utilized during routine diagnostic procedures.
To date, the distribution of F18 fimbrial subtypes among porcine STEC and STEC/ETEC strains in Germany has only been assessed with slide agglutination assay. 41 Contemporary molecular methods have not yet been applied. In the current study, a novel duplex PCR was developed in order to improve the F18 fimbria typing procedure in terms of costs, time, and discriminative power. Subsequently, the new PCR was used to determine the F18 fimbrial subtypes encoded in stx2e+ E. coli isolates from pigs in Germany. The results were utilized to examine possible correlations between F18 fimbrial subtypes with certain virulence genes and O antigens.
Material and methods
Bacterial strains
Well-characterized E. coli isolates (n = 244) were selected from a field strain collection originating from 2,443 pigs in Germany. 3 Thereof, 241 strains were positive for stx2e and fedA genes (116 STEC and 125 STEC/ETEC isolates). Virulence genes and O antigens in those isolates have been determined previously. 3 For nucleotide analysis of the fedA gene, 3 additional stx2e-negative strains were included: 1 enterotoxigenic E. coli strain (P6006/01-3) and 2 nontoxigenic strains that were fedA positive (P1748/03-1 and 326). The E. coli strains 107/86 a (O139, stx2e, fedAab) and 2134 b (O157, estb, estap, fedAac) were used as controls. All strains were stored at –70°C in lysogeny broth (LB) with 30% (v/v) glycerin. Prior to any test, the E. coli strains were freshly cultivated on sheep blood agar.
F18 fimbriae agglutination assay
Strains were tested for expression of F18 fimbriae with a slide agglutination assay utilizing a polyclonal F107 test serum. c Briefly, bacteria were cultivated on Iso-Sensitest alizarin agar d at 37°C (10 vol.% CO2). After a 16–20 hr incubation period, 3–4 single colonies were mixed with the test serum. 41 The bacterial suspension was examined macroscopically for agglutination. To exclude spontaneous agglutination, each strain was also tested with 0.89% sodium chloride solution instead of test serum.
F18 typing polymerase chain reaction assay
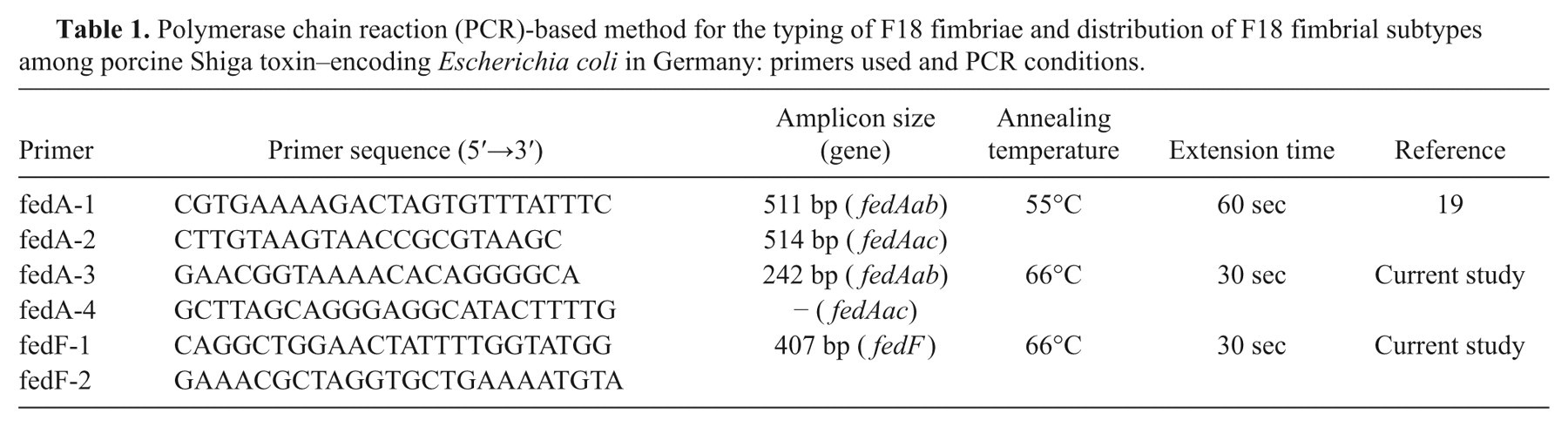
Prior to PCR analysis, the E. coli strains were grown under aerobic conditions for 14–16 hr at 37°C in LB. For typing of the fedA gene, a duplex PCR using primers fedA-3, fedA-4, fedF-1, and fedF-2 (Table 1) was established. The PCR mix (30 µl) contained 1× buffer (ammonium mix e ), 2 mM of magnesium chloride, e 1 U of PanScript polymerase, e 133 µM of each nucleotide, e 0.5 µM of each oligonucleotide, f and 3 µl of the respective bacterial culture (approximately 3 × 105 CFU bacterial cells). Cycling conditions g were as follows: 1 denaturation cycle (94°C, 5 min), 30 cycles with denaturation (94°C, 30 sec), annealing (temperature as listed in Table 1, 30 sec), and extension (72°C, time as listed in Table 1) as well as 1 final extension cycle (72°C, 5 min). All PCR products were analyzed after electrophoresis in 2% agarose gels.
Polymerase chain reaction (PCR)-based method for the typing of F18 fimbriae and distribution of F18 fimbrial subtypes among porcine Shiga toxin–encoding Escherichia coli in Germany: primers used and PCR conditions.
F18 restriction fragment length polymorphism assay
Analysis of the PCR-RFLP assay–generated fedA gene fragments was performed as previously described. 19 With the exception of primers fedA-1 and fedA-2 (Table 1), the reaction mix composition and PCR conditions were identical to the F18 typing PCR described above. Subsequently, 500 ng of the PCR product was purified using a commercial PCR purification kit. h Purified DNA was digested with 3 U of NgoMIV i (formerly, NgoMI). After 3 hr of incubation at 37°C, fragments were analyzed by electrophoresis in 2% agarose gels.
DNA sequencing
Purified PCR products generated with primers fedA-1 and fedA-2 (see above) were used as templates for DNA sequencing. Sequencing was performed by a commercial sequencing service. j The nucleotide sequences were deposited in the GenBank database under accession numbers GQ325621–GQ325633. The software package Lasergene k was applied for analysis of the DNA sequences. Basic local alignment research tool for nucleotide (BLASTn) 2.2.21 and Basic local alignment research tool for proteins (BLASTp) 2.2.21 were used for comparison with database entries. 2
Statistical analysis
Data were analyzed using the program SPSS Statistics. l Significance levels were defined as follows: P ≤ 0.001, strongly significant; 0.01 ≥ P > 0.001, significant; 0.05 ≥ P > 0.01, weakly significant; P > 0.05, not significant.
Results
Primer design and optimization of the F18 typing polymerase chain reaction assay
A novel PCR assay was designed in order to determine the F18 fimbrial subtypes in E. coli field isolates. The forward primer (fedA-3) matched exactly to positions 349–369 in the fedAab sequence. However, the 3’-terminus of this primer could not bind exactly to the fedAac gene due to a proline-encoding triplet inserted at positions 364–366 in this gene. The conserved binding site of the reverse primer (fedA-4) was localized approximately 50 bp downstream of the fedA stop codon. As strains with the F18ac fimbrial subtype would give no amplification signal, another pair of primers was added as an internal control system. The target sequences of these primers were localized in the fedF gene, encoding the F18 fimbrial adhesin FedF, 34 which was previously shown to be highly conserved in both F18ab and F18ac fimbrial subtypes. 37 The fedF primer pair matched all but one published fedF sequences exactly; 1 fedF entry (accession no. DQ914286) contained a mismatch to the fedF-1 forward primer. At the adjusted annealing temperature of 66°C, the F18 typing PCR yielded both expected amplicons of 407 bp and 242 bp for the fimbrial subtype F18ab and 1 amplicon of 407 bp for the subtype F18ac, respectively, with the corresponding control strains (Fig. 1).
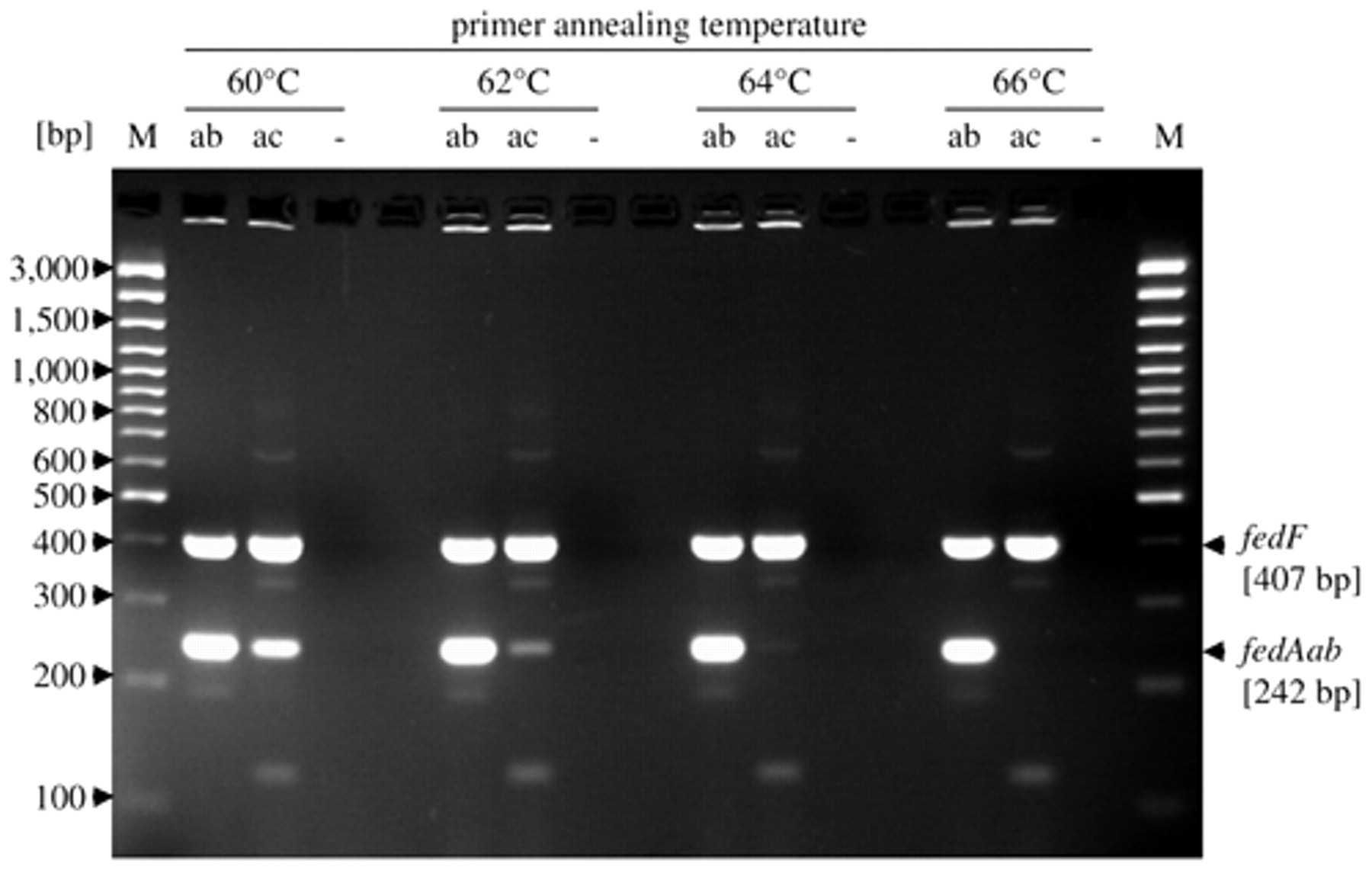
Polymerase chain reaction (PCR)-based method for the typing of F18 fimbriae and distribution of F18 fimbrial subtypes among porcine Shiga toxin–encoding Escherichia coli in Germany: detection of genes fedF and fedA in 2 E. coli strains encoding F18ab or F18ac fimbriae, respectively, with the new F18 typing polymerase chain reaction (PCR) at different annealing temperatures. At 66°C annealing temperature fedAab gene positive strains yielded two amplicons (242 bp and 407 bp) and fedAac positive strains only one (407 bp). Two percent (2%) Tris–acetate–ethylenediamine tetra-acetic acid agarose gel after ethidium bromide staining. Bacterial lysates were used as PCR templates. ab = E. coli 107/86 (F18ab); ac = E. coli P6006/01-3 (F18ac); – = negative control (lysogeny broth); M = molecular size marker, Gene Ruler Plus. m
Comparison of the F18 typing PCR and the F18 PCR-RFLP-assays
All 241 fedA + STEC strains were tested with both the new F18 typing PCR and the F18 PCR-RFLP assay as previously described, 19 which served as the reference assay. Both tests typed 234 E. coli strains (97.1%) in an identical manner (Table 2). Six strains were shown to possess the fedAac gene when using the F18 typing PCR while the F18 PCR-RFLP assay classified them as fedAab positive. One strain (strain 85), which harbored a fedAac gene according to the results in the F18 PCR-RFLP assay, produced neither fedAab nor fedF amplicons in the F18 typing PCR.
Polymerase chain reaction (PCR)-based method for the typing of F18 fimbriae and distribution of F18 fimbrial subtypes among porcine Shiga toxin–encoding Escherichia coli in Germany: comparison of the fedA typing results of 241 porcine E. coli strains in 2 assays.*
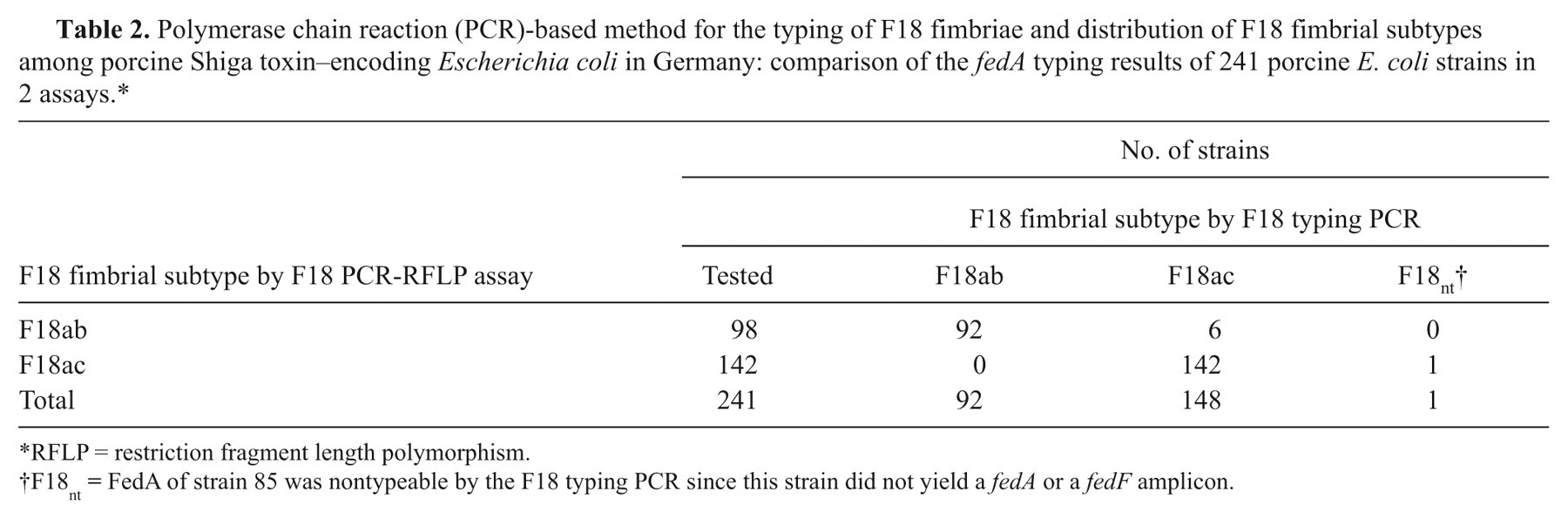
RFLP = restriction fragment length polymorphism.
F18nt = FedA of strain 85 was nontypeable by the F18 typing PCR since this strain did not yield a fedA or a fedF amplicon.
DNA and deduced amino acid sequence analysis
The 6 strains named above, which showed different results in both assays (F18 typing PCR and F18 PCR-RFLP assay) as well as strain 85 (nontypeable in the F18 typing PCR), were submitted to nucleotide sequence analysis of their fedA genes. By using the primer pair fedA-1 and fedA-2, the fedA gene was amplified from the start codon through the triplet encoding the C-terminal amino acid of the premature FedA protein. In order to elucidate the sequence variability of fedA genes in porcine E. coli isolates, 6 additional fedA + strains of the institutional strain collection were included in this analysis. These consisted of 3 fedAab STEC strains and 1 fedAac ETEC strain (all of which were typed identically in both assays) as well as 2 strains without Stx2e and enterotoxin genes that showed different typing results (i.e., strains that were fedAac-positive by F18 typing PCR and fedAab-positive by F18 PCR-RFLP assay [strains P1748/03-1 and 326]).
Comparison of all 13 sequences elaborated in the present study as well as of 10 sequences available in GenBank revealed polymorphisms at 40 sites of the fedA gene. Such polymorphisms at the DNA level corresponded to 27 polymorphic sites at the amino acid level, taking also into consideration both previously published FedA sequences. 7 Table 5 depicts the polymorphic sites at the amino acid level in these 25 FedA sequences. Apart from such variability, all strains encoded for identical putative signal peptides comprising the first 21 amino acids. Only minor differences occurred between nucleotide sequences of the 4 strains that had been typed in an identical manner in both assays. In contrast, fedA sequences of the 9 strains exhibiting inconclusive results in both assays (7 strains from the typing study plus 2 additional strains) showed major differences to previously published data. 7,19,21 All 9 strains had been classified as fedAab by the F18 PCR-RFLP assay; but sequencing revealed that the triplet insertion typical for fedAac was in fact present in all these strains at positions 364–366 of the fedA gene. However, in 8 strains, this insertion did not constitute an NgoMIV recognition site. In 7 of these strains, the inserted triplet differed from that in the fedAac reference gene (accession no. L26107) by only 1 nucleotide, while the triplet of strain 6196/01-1 was different at all 3 sites. Due to such substitutions, the encoded amino acid at site 122 was not proline but leucine (4 strains), alanine (1 strain), serine (1 strain), and methionine (1 strain), respectively (Table 5). In strain P1748/03-1, the proline-encoding triplet was present, but a substitution in the following triplet made this fedA gene resistant to digestion with NgoMIV as well. These results strongly suggest that the F18 PCR-RFLP assay misclassified these 8 E. coli strains as fedAab due to its missing the NgoMIV recognition site. In contrast, the fimbrial subtype of strain 85 was typed correctly as fedAac since the fedA gene of this strain harbored the proline-encoding triplet and a functional NgoMIV recognition site at codon 122.
In order to determine the homology between primary structures of mature FedA proteins, their predicted amino acid sequences were submitted to ClustalW sequence analysis, and their relatedness was illustrated by a dendrogram (Fig. 3). 36 Such analysis revealed that the FedA sequences of 8 of the 9 strains with inconclusive PCR typing results (F18 typing PCR: fedAac gene, F18 PCR-RFLP assay: fedAab) showed greater homology to published FedAac sequences than to FedAab sequences (Fig. 3). Thus, the mature FedA protein sequence of all strains except strain 6196/01-1 displayed 94–98.7% similarity to published F18ac fimbrial sequences. The FedA of strain 6196/01-1 (GQ 325633) could not be allocated to a particular subtype, as the similarity to F18ab subtypes was not greater than 91.3%, and to F18ac subtypes, was not greater than 92.7% (Fig. 3).
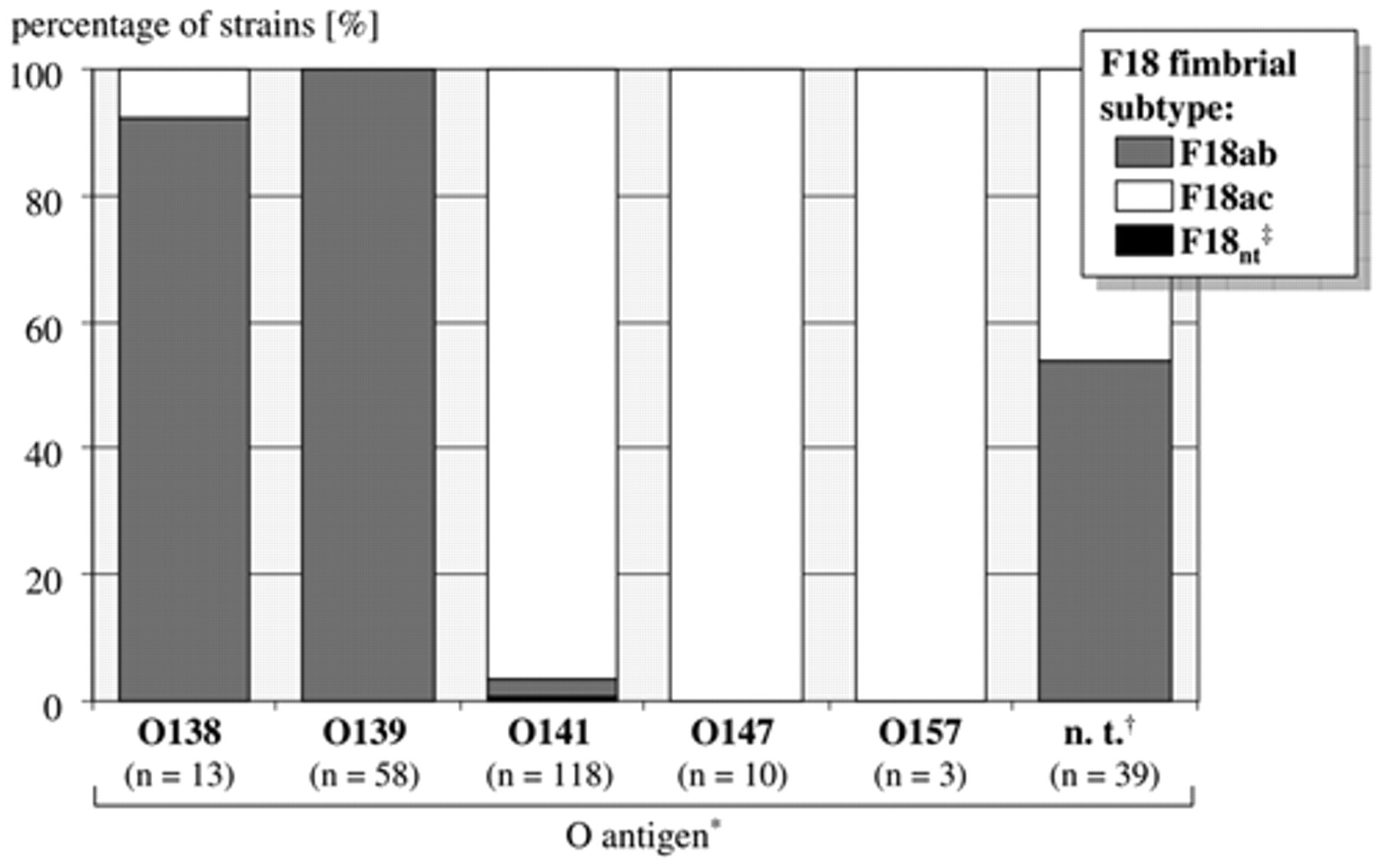
Polymerase chain reaction (PCR)-based method for the typing of F18 fimbriae and distribution of F18 fimbrial subtypes among porcine Shiga toxin–encoding Escherichia coli in Germany: distribution of F18 fimbrial subtypes in 241 porcine stx2e +, fedA + E. coli strains depending on the O antigen of the strains (F18 typing PCR results).
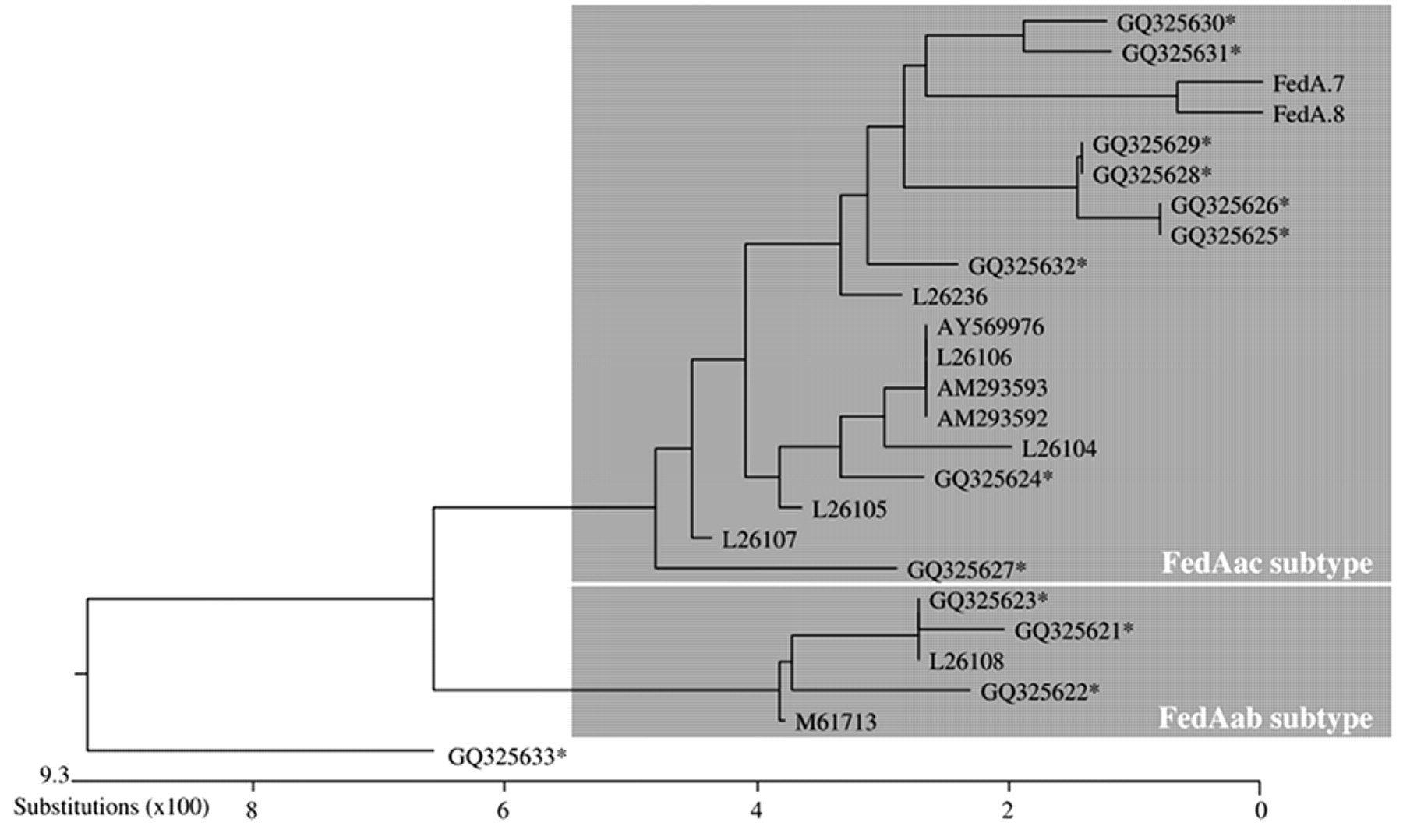
Polymerase chain reaction (PCR)-based method for the typing of F18 fimbriae and distribution of F18 fimbrial subtypes among porcine Shiga toxin–encoding Escherichia coli in Germany: rooted phylogenetic tree derived from ClustalW sequence analysis of mature FedA proteins encoded by 25 porcine E. coli strains (amino acid positions 22-156; deduced amino acid sequences without the putative signal sequence). Cluster of the same fimbrial subtype (more than 94% similarity) are marked by grey squares. Sequences were designated with their respective GenBank accession numbers, with the exception of sequences FedA.7 and FedA.8 that were taken from Bosworth et al., 1998 7 ; ClustalW multiple sequence alignment (modus: slow/accurate; PAM 35 ). k Asterisk (*) indicates strains of the current study.
Association of F18 fimbrial subtypes with other virulence genes
In total, 116 STEC and 125 STEC/ETEC strains from pigs in Germany were tested for their F18 subtype using the F18 typing PCR (Table 3). Among these strains, there was a significant but not exclusive correlation between the F18 subtype and the presence of genes for ST-Ip, ST-II, LT-I, and/or EAST-1 enterotoxins (χ2: P < 0.001). The majority (73.3%) of enterotoxin-negative STEC strains encoded for the F18ab fimbrial subtype. In contrast, 93.6% of the STEC/ETEC strains encoded for the F18ac subtype (χ2: P < 0.001). Furthermore, F18ab was significantly associated with AIDA genes since nearly all of those strains (94.6%) possessed these genes. This was only the case in 71.6% of the fedAac strains (χ2: P < 0.001). None of the F18 fimbrial subtypes was significantly correlated with faeG (F4 fimbrial subunit gene; χ2: P > 0.05).
Polymerase chain reaction (PCR)-based method for the typing of F18 fimbriae and distribution of F18 fimbrial subtypes among porcine Shiga toxin–encoding Escherichia coli in Germany: encoded F18 subtypes in porcine E. coli strains carrying genes for different combinations of virulence factors.*
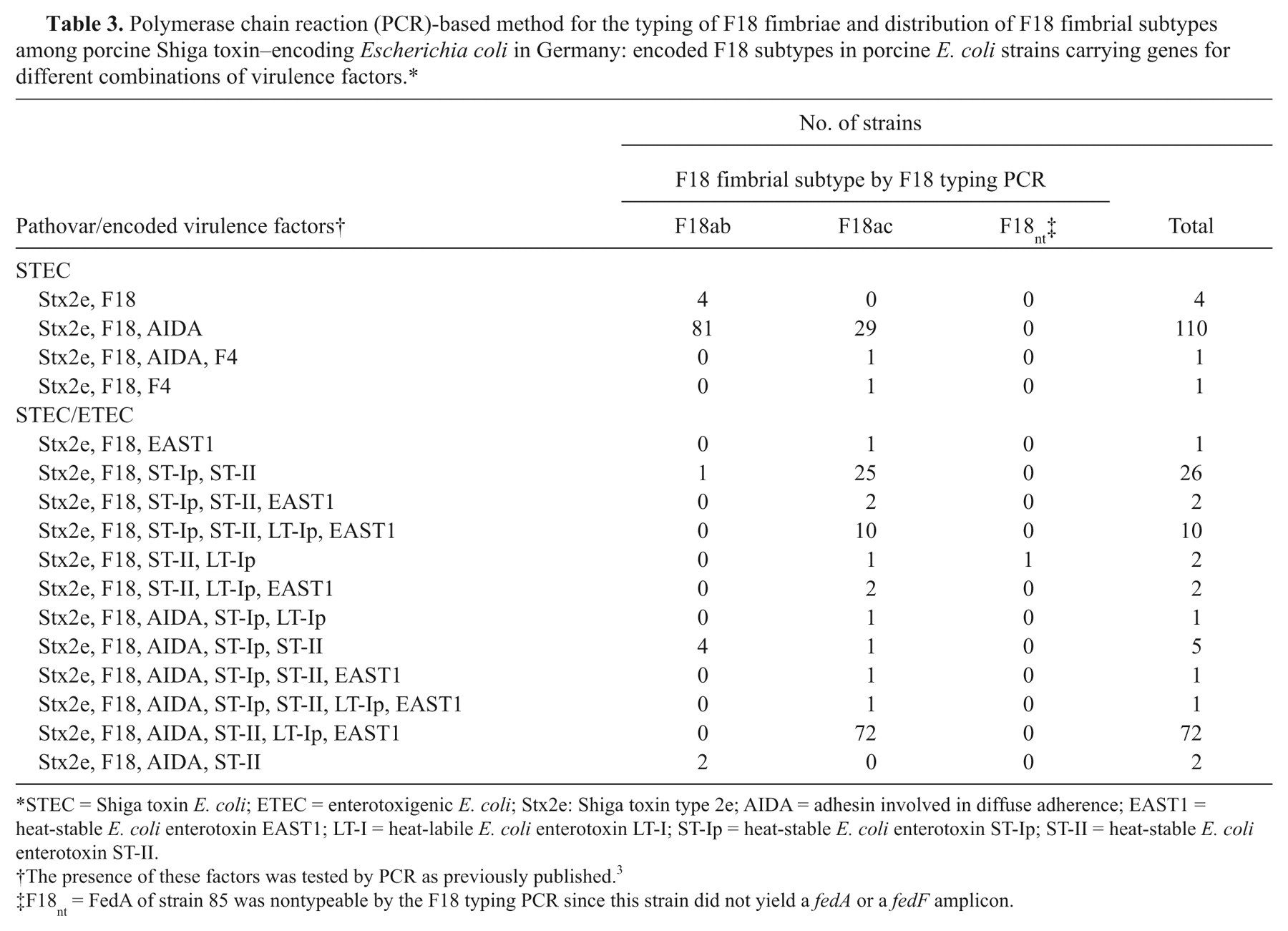
STEC = Shiga toxin E. coli; ETEC = enterotoxigenic E. coli; Stx2e: Shiga toxin type 2e; AIDA = adhesin involved in diffuse adherence; EAST1 = heat-stable E. coli enterotoxin EAST1; LT-I = heat-labile E. coli enterotoxin LT-I; ST-Ip = heat-stable E. coli enterotoxin ST-Ip; ST-II = heat-stable E. coli enterotoxin ST-II.
The presence of these factors was tested by PCR as previously published. 3
F18nt = FedA of strain 85 was nontypeable by the F18 typing PCR since this strain did not yield a fedA or a fedF amplicon.
Association of the F18 fimbrial subtype with F18 fimbriae expression and O antigens
Based on earlier observations that some O antigens are dominantly present within certain pathovars, 3 it was assessed whether F18 fimbrial subtypes were associated with the respective O antigen. In the present study, almost all strains (97.1%) expressing O antigens 138 or 139 encoded for F18ab fimbria. In contrast, almost all those strains (96.9%) that expressed O antigens 141, 147, or 157 on their bacterial surface also encoded for F18ac fimbriae (Fig. 2). However, both subtypes were present at almost equal numbers among 39 strains that proved nontypeable with the test sera used in the current study (F18ab, 20 strains; F18ac, 19 strains).
All 241 (stx2e+ , fedA+ ) strains were tested for the expression of their F18 fimbrial genes by slide agglutination assay (Table 4). Specific agglutination was detectable in 40 strains only (16.6%) although the strains had been cultured under conditions (Iso-Sensitest alizarin agar, d 10 vol.% CO2 atmosphere) that should have induced the expression of F18 fimbriae according to the literature. 41 A positive result was yielded by 23.8% of all F18ac strains and by 5.4% of the F18ab strains (χ2: P < 0.001).
Polymerase chain reaction (PCR)-based method for the typing of F18 fimbriae and distribution of F18 fimbrial subtypes among porcine Shiga toxin–encoding Escherichia coli in Germany: phenotypic detection of F18 fimbriae in 241 porcine E. coli strains in comparison to their fimbrial subtype.
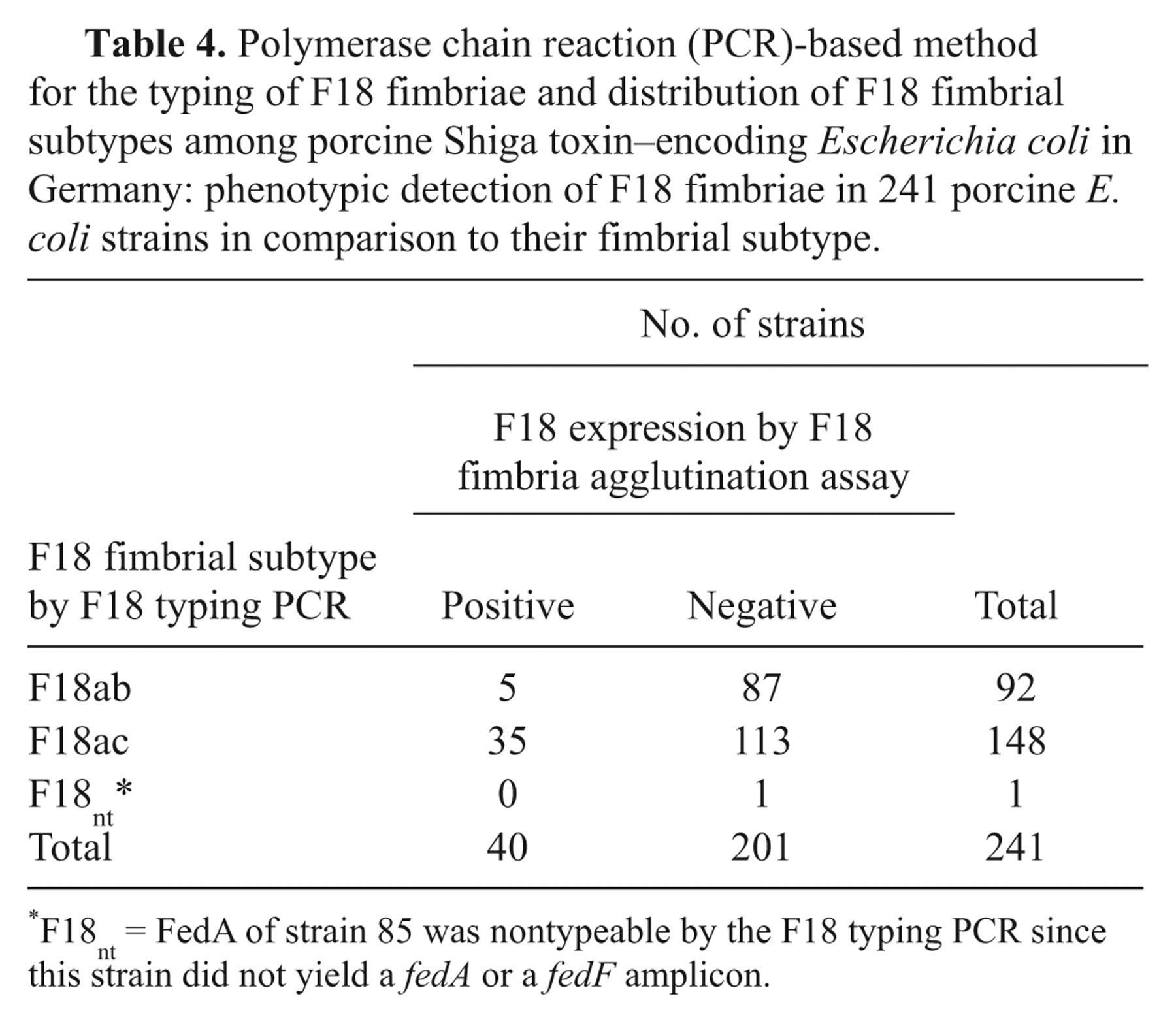
F18nt = FedA of strain 85 was nontypeable by the F18 typing PCR since this strain did not yield a fedA or a fedF amplicon.
Discussion
In the current study a new F18 typing PCR was established targeting the fedA gene for differentiation and the fedF gene for control. Using such typing system, a significant correlation of the F18 fimbrial subtype with the presence of enterotoxin genes and certain O antigens was similarly detected, as described by some investigators. 10,25,40
The original procedure for the differentiation of F18 fimbrial subtypes in E. coli isolates utilized polyclonal antisera able to agglutinate bacteria expressing one of the respective fimbrial antigens. 41 In addition to the slide agglutination assay, enzyme-linked immunosorbent assay (ELISA) systems or immunodot assays were developed for this phenotypic approach of fimbrial antigen typing. 4,22 Later, genotypic characterization was preferred, as many strains did not express the fimbriae even when they were cultured under conditions described to enhance the expression of F18 fimbria. 12,25,30,41 Genetic characterization allows distinguishing between strains that fail to express detectable amounts of F18 fimbrial antigen and strains that lack the fed gene locus. Both types of E. coli strains have been repeatedly found in pigs. 3,9,31,40 The present study confirms those observations due to the fact that over 80% of the fedA gene–positive strains tested failed to agglutinate with the specific antisera (Table 4).
The F18 typing PCR established in the current study is a fast and simple assay that was able to identify the respective F18 fimbrial subtype based on the nucleotide sequence of the fedA gene. In contrast to previous studies, 8 nonspecific DNA amplification with fedAab-specific primers from the fedAac-positive control strain was eliminated by adjusting the annealing temperature to 66°C (Fig. 1). However, 9 strains were typed differently by the new F18 typing PCR in comparison to the previously established F18 PCR-RFLP assay. 19 In order to reassess these typing results, nucleotide sequencing of the fedA genes of these 9 strains and 4 strains with identical results in both assays was performed. Sequencing revealed that the fedA gene was more heterogeneous than suspected. Five amino acids at positions 31, 57, 59, 83, and 122 of the premature FedA, which represent a specific signature either for the F18ab (N31A57S59T83–122) or the F18ac subtype (S31T57N59N83P122), were previously identified. 7 Furthermore, the authors predicted 2 antigenic regions (amino acid positions 116–130 and 152–160) by computer analysis. 7 Interestingly, the amino acid at position 122 was the only signature residue that was localized in such predicted antigenic regions and, therefore, seemed to be crucial both for genetic and phenotypic differentiation between FedA subtypes. The triplet encoding proline at position 122 in FedAac introduces also the NgoMIV recognition site that is used for differentiation between fedAab and fedAac genes by the F18 PCR-RFLP assay. However, when the 13 FedA sequences of the present study were determined, only 5 sequences classified by F18 PCR-RFLP, as FedAab (3 strains) and FedAac (2 strains), respectively, matched the criteria of these subtypes (Table 5). Eight other FedA sequences were similar to the F18ac subtype but exhibited leucine (4 strains), alanine (1), serine (1), or methionine (1) at position 122 instead of proline at position 122. Recently, significant amino acid variability at such position among North American strains had been demonstrated as well. 12 In line with the present findings, investigators detected, besides proline, the amino acids alanine (15 strains), cysteine (1), glutamine (1), and glycine (1). 12
Polymerase chain reaction (PCR)-based method for the typing of F18 fimbriae and distribution of F18 fimbrial subtypes among porcine Shiga toxin–encoding Escherichia coli in Germany: polymorphic sites in the deduced FedA amino acid sequences of 25 E. coli strains.*
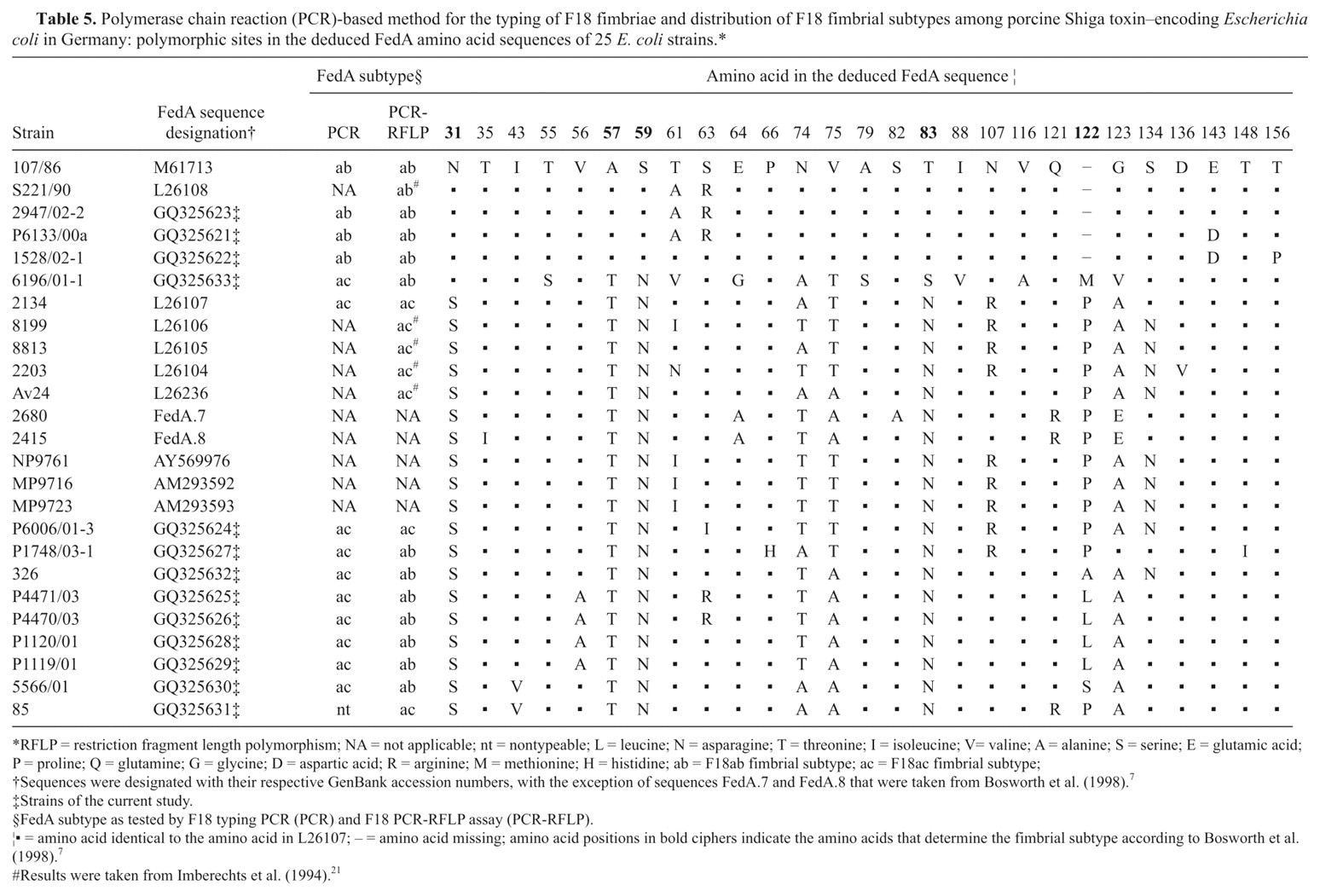
RFLP = restriction fragment length polymorphism; NA = not applicable; nt = nontypeable; L = leucine; N = asparagine; T = threonine; I = isoleucine; V= valine; A = alanine; S = serine; E = glutamic acid; P = proline; Q = glutamine; G = glycine; D = aspartic acid; R = arginine; M = methionine; H = histidine; ab = F18ab fimbrial subtype; ac = F18ac fimbrial subtype;
Sequences were designated with their respective GenBank accession numbers, with the exception of sequences FedA.7 and FedA.8 that were taken from Bosworth et al. (1998). 7
Strains of the current study.
FedA subtype as tested by F18 typing PCR (PCR) and F18 PCR-RFLP assay (PCR-RFLP).
▪ = amino acid identical to the amino acid in L26107; – = amino acid missing; amino acid positions in bold ciphers indicate the amino acids that determine the fimbrial subtype according to Bosworth et al. (1998). 7
Results were taken from Imberechts et al. (1994). 21
In the current study, 11 of 13 FedA amino acid sequences could be assigned to one specific subtype beyond doubt, as they showed more than 94.0% similarity during pairwise alignment in the ClustalW analysis and possessed the specific signature for the respective subtype with no or only one substitution (Table 5; Fig. 3). The deduced FedA sequence of strain 6196/01-1 took up an intermediate position and may represent a novel subtype of the F18 fimbria (Fig. 3). Such FedA sequence contained characteristics from both the F18ab (N31) and the F18ac sequence signatures (T57N59) as well as novel signature residues (S83M122). In the phylogenetic tree, the FedA of strain 6196/01-1 is localized outside the FedAac and FedAab clusters. It is possible that strain 6196/01-1 expresses an ancestor F18 type, but it cannot be excluded that this intermediate type has newly evolved from one of both currently known subtypes F18ab and F18ac.
Strain 85 generated no amplification signal in the F18 typing PCR and was, therefore, assumed fedAab and fedF negative, but was fedAac positive in the F18 PCR-RFLP assay. Such result suggests an incomplete fed gene locus to be present in this particular strain due to the fact that fedF is considered to be a highly conserved gene in the fed locus of E. coli. 37 However, the fed locus of this aberrant strain remains to be elucidated in the future.
Taken together, the F18 typing PCR provided a higher accuracy than the F18 PCR-RFLP assay. Using the F18 typing PCR, only 1 of 241 F18+ strains (0.8 %) was typed incorrectly, while 6 strains (2.5%) were typed incorrectly as fedAab + by use of the F18 PCR-RFLP assay. The second primer pair in the F18 typing PCR made it possible to exclude false-positive typing for fedAac, which could have happened in the case of an incomplete fed locus, as it is possibly present in strain 85. The NgoMIV recognition site used in the F18 PCR-RFLP assay does not seem highly conserved among F18ac strains, and, therefore, should not be regarded as an absolutely reliable marker for this subtype. Such conclusion is in concordance with previous studies, where the F18ac sequences FedA.7 and FedA.8 were identified by SSCP analysis. 7 Both sequences would not have been allocated correctly by the F18 PCR-RFLP method although those sequences exhibited a proline at position 122.
In the current study, F18ab was the predominant fimbrial subtype in E. coli strains exposing the O138 or the O139 antigen, while most, if not all, strains carrying O141, O147, or O157 antigens encoded for F18ac fimbriae. With the exception of a recent study, 12 where both F18 fimbrial subtypes were detected at almost equal rates in porcine O147 strains isolated in the United States, other investigators observed similar associations in E. coli isolates from pigs in the Czech Republic, Australia, and Hungary. 1,17,26 Such findings suggest that F18 fimbriae are genetically linked to some O types or that the same dominant STEC or STEC/ETEC strains are distributed in those countries. Also, similar to what was previously observed in other countries 8,26 and in Germany, 40 the F18ac subtype and the presence of genes for ST-Ip, ST-II, LT-I, and/or EAST-1 enterotoxins were significantly associated with each other in the current study. An explanation for such phenomenon has not been determined. Since the fed locus as well as enterotoxin genes is localized on plasmids, it could be assumed that plasmids encoding the F18ab subtype are less compatible with those encoding for E. coli enterotoxins. However, in a previous study, it was determined in a collection of ETEC, STEC, and STEC/ETEC strains that regardless of some size differences, both types of F18 plasmids were unireplicon plasmids belonging to the same replicon family RepFIc of the incompatibility complex F. 14 Therefore, it is hypothesized that F18 encoding plasmids may have potentially emerged from a common ancestor plasmid and that an undetermined process must have co-selected enterotoxin and F18ac encoding plasmids.
Several authors have demonstrated that a high percentage of porcine STEC and ETEC strains encodes for AIDA, in addition to F18 fimbriae, which are encoded on the same plasmid in those strains. 24,27 The AIDA is an autotransporter protein that mediates bacterial autoaggregation, biofilm formation, and bacterial attachment to epithelial cells by a diffuse adherence pattern in vitro. 5,28,33 However, the precise role of AIDA in the pathogenesis of edema disease is undetermined. Experimental infection of colostrum-deprived newborn piglets revealed that AIDA can be essential for ETEC strains to colonize the intestinal mucosa and to cause diarrhea. 28 It appears plausible to assume that AIDA acts similar during STEC and STEC/ETEC infections, where it presumably fosters intestinal colonization by these pathogens. In the present study, the F18ab subtype was positively correlated with the presence of AIDA genes and negatively correlated with the expression of the F18 fimbriae as demonstrated by negative results in the slide agglutination assay. Similar, both strains encoding for F4 fimbriae did not express their F18ab fimbrial genes. Such findings suggest that AIDA and adhesive F4 (and possibly F5 12 ) fimbriae may act as alternative or complement attachment factors to the F18 fimbrial adhesin in those strains that cannot produce these fimbriae at rates sufficient for bacterial attachment to the mucosa.
The knowledge about which F18 fimbrial subtypes are present in the field is important for the development of new vaccines and diagnostic assays. In the last few years, several experimental studies with piglets orally vaccinated against post weaning diarrhea and/or edema disease have been performed. 6,32 Results suggested that protection of the piglets was only achieved by oral vaccination with a live E. coli strain expressing the same F18 fimbrial subtype as the challenge strain. 6 Pigs vaccinated with a F18ac strain were only partially protected during oral challenge with a F18ab strain, as measured by reduced fecal shedding of challenge bacteria. 32 Similar, F18-specific immunoglobulin Y (IgY) yielded better protection in passive immunization experiments if these IgY antibodies were generated with the same F18 fimbrial antigen as carried by the challenge strain. 20 Results of the present study suggest that a fimbrial vaccine directed against porcine STEC/ETEC infection in Germany would have to cover both subtypes of F18 fimbriae in order to provide significant protection to the susceptible population.
Footnotes
Acknowledgements
The authors are grateful to Prof. Dr. Hans U. Bertschinger, Zürich, Switzerland, and Prof. Dr. Béla Nagy, Budapest, for generously supplying the reference strains and to Philip S. Bridger, Ph.D., for his suggestions on the manuscript.
a.
Strain 107/86 was kindly provided by Prof. Dr. Hans U. Bertschinger, Zürich, Switzerland.
b.
Strain 2134 was kindly provided by Prof. Dr. Béla Nagy, Budapest, Hungary.
c.
BfR, Federal Institute for Risk Assessment, Dessau, Germany.
d.
Oxoid Deutschland GmbH, Wesel, Germany.
e.
PAN Biotech GmbH, Aidenbach, Germany.
f.
MWG-Biotech AG, Ebersberg, Germany.
g.
Model T9, Biometra GmbH, Göttingen, Germany.
h.
QIAquick®, Qiagen GmbH, Hilden, Germany.
i.
New England Biolabs GmbH, Frankfurt/Main, Germany.
j.
GENterprise GmbH, Mainz, Germany.
k.
DNAStar Inc., Madison, WI, via GATC Biotech, Konstanz, Germany.
l.
Version 17.0.0, SPSS Inc., Chicago, IL.
m.
Fermentas, St. Leon-Rot, Germany
The authors declared that they had no conflicts of interest with respect to their authorship or the publication of this article.
The authors declared that they received no financial support for their research and/or authorship of this article.
