Abstract
Aquabirnaviruses (ABVs) cause serious diseases in a variety of fish species used worldwide in aquaculture and have been isolated from a variety of healthy fish and shellfish species. The type species of ABV is Infectious pancreatic necrosis virus (IPNV), which is the causative agent of a highly contagious disease in juvenile salmonid fish. Marine birnaviruses (MABVs) have been isolated from various marine fish and shellfish. In Korea, ABV infection has been identified in several fish and shellfish. The current study presents sequence data from nested polymerase chain reaction products of 3 ABV strains obtained from different species of asymptomatic aquarium fish collected from a private commercial aquarium in Korea. Phylogenetic analysis of these strains, based on the partial nucleotide sequence of the VP2/NS junction, placed them within the genogroup VII (95–99% bootstrap confidence), which also contains MABV. The subclinically infected fish may be a source of MABV infection for other susceptible fish species inside the aquarium and potentially represent a serious challenge for the management of MABV infections. Additionally, the presence of MABV in these subclinically infected aquarium fish imported from other countries indicates that there is a need for the establishment of appropriate quarantine practices.
The Aquabirnavirus genus of the family Birnaviridae infects many species of fish and shell fish. 4 Different aquabirnavirus (ABV) strains can infect many species of both freshwater and marine fish, causing a variety of clinical signs. The type species of ABV is Infectious pancreatic necrosis virus (IPNV), which is the causative agent of a highly contagious disease in juvenile salmonid fish worldwide. 9 Other aquabirnaviruses have been isolated from various marine fish and shellfish and are tentatively named marine birnaviruses (MABV). 11 MABV, first isolated from yellowtail or Japanese amberjack (Seriola quinqueradiata) in Japan, was designated as Yellowtail ascites virus because of the disease's characteristic signs. 23
The ABV genome consists of 2 segments (large and small RNA segments, designated A and B, respectively) of double-stranded RNA, packaged in a nonenveloped, icosahedral shell that is 60 nm in diameter. 3 Segment A contains 2 partially overlapping open reading frames (ORFs). The large ORF encodes a 106-kD polyprotein that is cleaved by a viral nonstructural (NS) protease to generate the structural viral proteins (VPs) 2 and 3. 6 The junction region of the VP2/NS was previously used for the phylogenetic analysis of the ABV isolates. 20 The ABV can be divided in to 7 genogroups; more specifically, IPNVs belong to the genogroups I–VI, and MABV belongs to the genogroup VII. 20 Segment B encodes VP1, which is the RNA-dependent RNA polymerase associated with virions. 5
In Korea, IPNV has been isolated from cultured chum salmon (Oncorhynchus keta) and goldfish (Carassius auratus). 8 In addition, MABV infection has been reported in Japanese flounder (or olive flounder; Paralichthys olivaceus), Korean rockfish (Sebastes schlegeli), Suzuki (Lateolabrax japonicus), Mediterranean mussel (Mytilus galloprovincialis), and sea squirt (Halocynthia roretzi). 1,13,17,21 Korea imports a large variety of aquarium (marine or freshwater) fish from Southeast Asian countries, and the numbers of such imported species increase annually. 15 The importation of aquarium fish is a potentially important factor in the transmission of fish diseases. Recently, a viral pathogen (betanodavirus) was detected from apparently healthy aquarium fish and invertebrates that had been imported by a commercial aquarium in Seoul, Korea. 7 In the present study, tissue samples from clinically healthy fish collected from a commercial aquarium in Korea were evaluated for the presence of ABV, using reverse transcription polymerase chain reaction (RT-PCR). Phylogenetic analysis was performed on the obtained viral sequences to determine whether the imported aquarium fishes could be a source of ABV.
From February to September 2009, 78 tissue samples (from 44 species) of clinically healthy marine and freshwater fish were collected from a commercial aquarium in Seoul, Korea. Sixty-two tissue samples were from 39 imported species and 16 tissue samples originated from 5 local species. The tissue samples, which consisted of brain, liver, kidney, spleen, intestine, and gill, were collected aseptically, pooled, and maintained at −80°C until use.
Total RNA was extracted from pooled tissues (one pool per fish) using an RNA extraction kit with TRIzol reagent, a in accordance with the manufacturer's instructions. The RT-PCR and nested RT-PCR were performed as previously described, 24 and the products were analyzed using 2.0% agarose gel electrophoresis in 1% Tris–borate–ethylenediamine tetra-acetic acid buffer. The gels were stained with ethidium bromide (0.5 μg/ml), visualized, and photographed under ultraviolet illumination.
Nested RT-PCR products were recovered from agarose gels and purified using a power gel extraction kit, b as described in the manufacturer's instructions. The purified nested RT-PCR amplification products were cloned into a plasmid vector using a TOPO cloner PCR cloning kit, c following the manufacturer's instructions. The cloned products were sequenced using universal primers (M13F: GTAAAACGACGGCCAGT; M13RpUC: CAGGAAACAGCTATGAC) and a terminator cycle sequencing kit. d Electrophoresis of the sequencing reactions was performed using an automatic DNA analyzer. e The nested RT-PCR products were identified as the junction region of the VP2/NS of ABV using a BLAST search (http://www.ncbi.nlm.nih.gov/blast/Blast.cgi), and these products were used for the phylogenetic analysis of the ABV isolates because of the diversity present among the isolates. 13,20 The variable region of the VP2/NS (134 nucleotides [nt], from nt 1,517 to nt 1,650) was aligned and compared with previously reported data for the ABV isolates. Multiple alignment algorithms from the ClustalX f method were analyzed with the Molecular Evolutionary Genetics Analysis (MEGA) program (version 4.0) g using the neighbor-joining method. Bootstrap confidence levels were determined by analysis of 1,000 replicate trees. The nucleotide sequences used for the phylogenetic tree are available from the GenBank databases, and the corresponding countries, host species, and accession numbers are provided in Table 1.
Virus isolation from nested RT-PCR–positive fish organs was performed. All viscera were homogenized in Hank balanced salt solution. h The homogenate was centrifuged at 2,450 × g for 10 min, and supernatant was filtered through a 0.45-μm pore-size filter. One hundred microliters of filtrates were inoculated to monolayers of Chinook salmon embryo–214 (CHSE-214) cells in culture flasks, followed by incubation at 20°C. The CHSE-214 cells were cultured in Eagle minimum essential medium, supplemented with 10% fetal bovine serum, penicillin (50 IU/ml), and streptomycin (50 μg/ml). The flasks were observed for 2 weeks for cytopathic effects (CPE).
Aquabirnavirus was detected by nested RT-PCR in 3 of the 78 fish (3.85%), with products (168 base pair [bp]) obtained from a Japanese eagle ray (Myliobatis tobijei), a lyretail anthias (or sea goldie; Pseudanthias squamipinnis), and a yellow boxfish (Ostracion cubicus). All 3 fish were imported from aquaria in Japan. Partial VP2/NS junction nucleotide sequences for the 3 ABV strains were determined using a BLAST (megablast) search for highly similar sequences in the National Center for Biotechnology Information (NCBI) nonredundant nucleotide (nr/nt) database. The determined nucleotide sequences were registered with GenBank under accessions GU590860–GU590862. Based on the phylogenetic tree, all 3 isolates from the aquarium fishes in the present study belonged to genogroup VII (Fig. 1). The isolates showed a high homology of 96.8–98.9% among the 3 strains and exhibited 95.2–99.5% similarity with other genogroup VII strains. However, these nested RT-PCR–positive fish samples did not produce any CPE in CHSE-214 cells.
In the present study, the presence of ABV in subclinically infected aquarium fish was investigated using RT-PCR and nested RT-PCR primers to amplify the ABV VP2/NS junction region, which is a highly variable domain of the viral genome. 2 The nested RT-PCR–positive samples were detected from 3 different clinically healthy marine fish species imported from Japan. The fact that virus was not isolated from nested RT-PCR–positive fish samples may be due to the presence of a low virus titer in the carrier fish. The phylogenetic analysis of nested RT-PCR products obtained from these samples placed them within the ABV genogroup VII, which also contains the MABV subgroup. 20,27 Based on previous reports, isolates of MABV have been limited only to Korea and Japan. 13,20,23 Isolates of ABV described to date are summarized in Table 1.
Phylogenetic analysis of the obtained MABV sequences placed them within the ABV genogroup VII, clustering most closely with isolates from Japan forming the MABV subgroup. The type isolate of MABV, originally isolated from cultured S. quinqueradiata, was first reported in Japan. 23 Since then, MABV has been reported from several cultured and wild marine fish in Japan, such as ayu sweetfish (Plecoglossus altivelis), Japanese flounder, snowy sculpin (Myoxocephalus brandti), saffron cod (Eleginus gracilis), Arctic rainbow smelt (Osmerus mordax dentex), and Japanese dace (or big-scaled redfin; Tribolodon hakonensi), all of which cluster genetically in the MABV subgroup within the ABV genogroup VII. 14,18,25 Although there have been ABV isolates from Korea that belong to the genogroup VII cluster, those isolates are different from the ones that cluster within the MABV subgroup. Because the 3 sequences obtained in the current study were most similar to strains isolated from Japan, rather than those isolated from Korea, it seems likely that these samples acquired the infection before their introduction to the Korean aquarium.
At present, there are no specific regulations regarding quarantine practices on the importation of aquarium fish into Korea. Consequently, the imported samples may also be infected after transport via horizontal transmission because the fish are usually comingled in one aquarium with other potentially subclinically infected wild fish collected from the Korean sea. Marine birnaviruses have been isolated from wild marine fish in Korea, such as belted beard grunt (Hapalogenys mucronatus), blackthroat sea-perch (Doederleinia berycoides), whitespotted conger (Conger myriaster), cubed snailfish (Liparis tessellatus), ridged-eye flounder (Pleuronichthys cornutus), largehead hairtail (Trichiurus lepturus), daggertooth pike conger (Muraenesox cinereus), Korean rockfish, scorpionfish (Scorpaena neglecta), yellowback seabream (Dentex tumifrons), and yellow croaker (Larimichthys polyactis). 26 Moreover, outbreaks have been recorded at several Japanese flounder farms in Korea. 22 The commercial aquarium involved in the present study uses multiple-use aquarium tanks, equipped with heating and filtered recirculating systems for rearing fish. It is possible that the MABV was horizontally transmitted to the fish via contaminated water or zooplankton 16,19 because most of the imported fish in the aquariums are comingled together with other fish, which may be subclinically infected with MABV.
Sources of aquabirnavirus strains or isolates whose sequence data were used in the genetic comparison with strains detected in the present study.
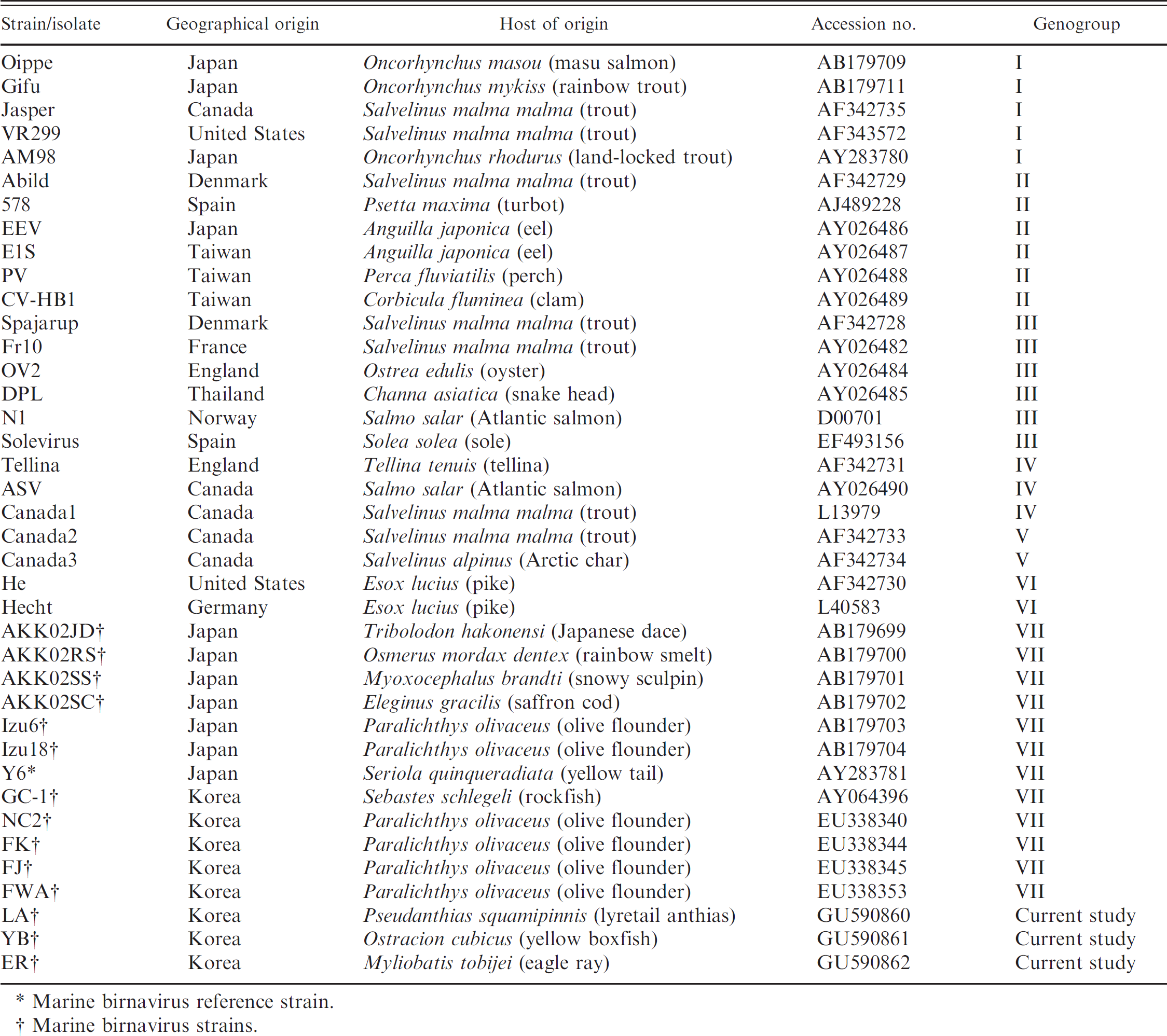
Marine birnavirus reference strain.
Marine birnavirus strains.
Despite the broad host specificity of MABV, the virulence of MABV varies from asymptomatic to lethal, according to the strain and host fish species. 10 The presence of subclinically infected fish inside the aquarium can lead to epizootics, suggesting that some of these carrier fish can shed infectious MABV particles in the water and that susceptible, healthy fish in the aquarium are at risk in this virus-contaminated environment, although the exact mechanism of infection has not been established. 19 Because any fish in the aquarium can potentially carry pathogens, preventive steps should be taken. Thus, it is recommended that aquariums quarantine all new arrivals with specific pathogen–free (SPF) sentinel fish, such as makogarei (or marbled flounder, Pseudopleuronectes yokohamae) or yellowtail, which are susceptible to MABV, 12 before transferring the new arrivals to the display. Based on a previous report, many strains of MABV were isolated from makogarei with isolation rates of>92% after experimental infection, and yellowtail was an appropriate, susceptible fish species to check the virulence of MABV strains. 12 Such a quarantine system could be implemented in conjunction with RT-PCR testing of the sentinel fish or of some of the new arrivals, similar to what was done in the current study. Only after the batch of fish has been determined to be free of ABV infection, would the fish be released into the general population.
Another possible source for potential horizontal transmission is via the use of contaminated aquarium tools and materials, associated with the staff's handling of fish inside the aquarium. Additionally, poor water quality and poor filtration system conditions in the aquarium may contribute to disease transmission. Disinfecting tools and materials that are used in the aquarium can help eliminate pathogen transmission to other fish in other aquaria. In addition, used water should be disinfected to prevent contamination of the aquaculture system or other aquatic environments.
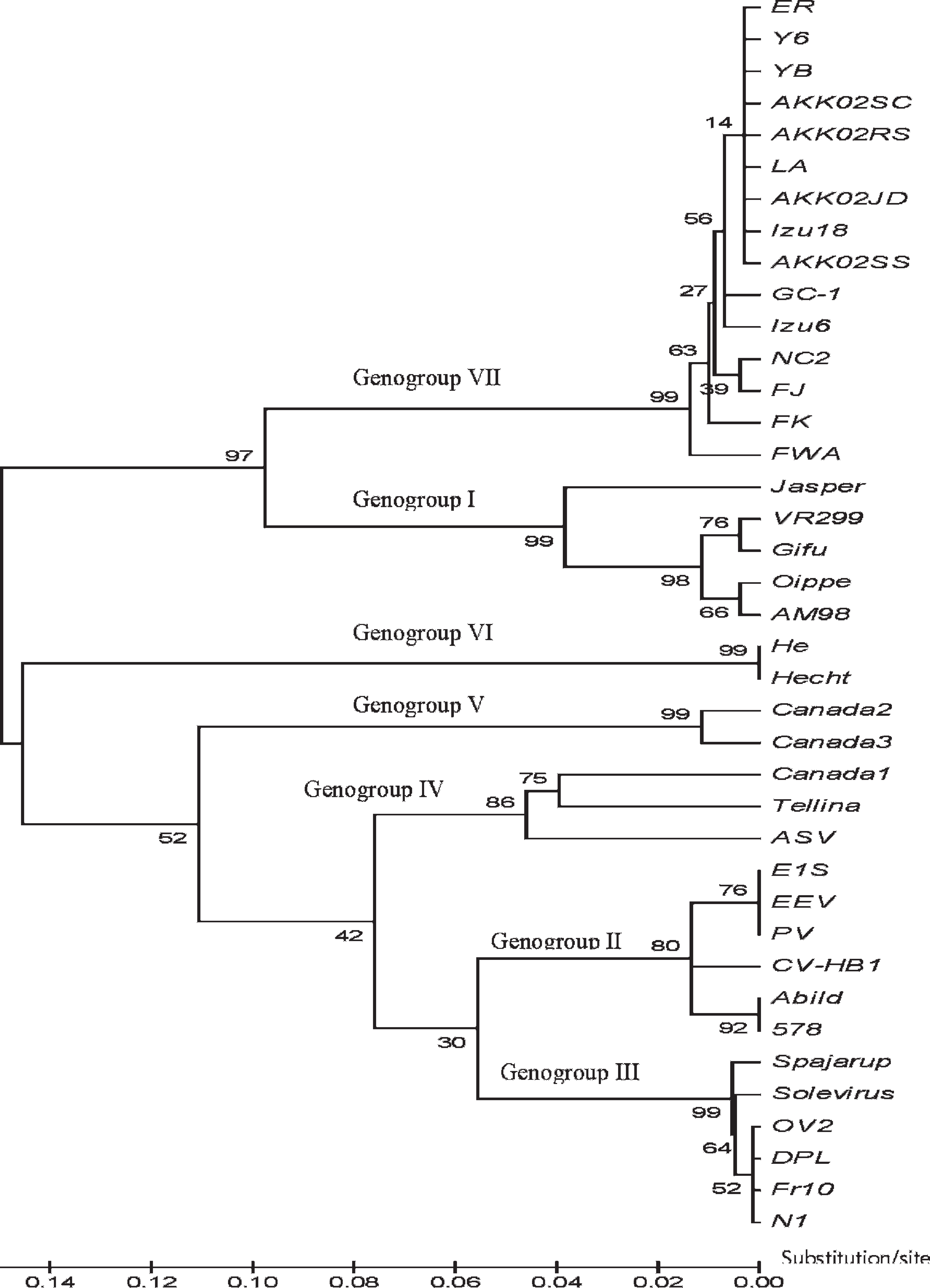
Molecular phylogenetic tree based on the nucleotide sequences of the VP2/NS junction region among marine birnavirus and Infectious pancreatic necrosis virus. The statistical confidence for each group was evaluated by bootstrap analysis with 1,000 replicates.
Based on the current study results, it is suggested that subclinically infected samples may constitute a potential source of MABV; therefore, there is a need for the establishment of a quarantine system in Korea. Until such system has been implemented, imported aquarium fishes should be randomly assessed by PCR-based techniques on a routine basis to ensure that they are free of specific pathogens.
Acknowledgements. The study was financially supported by a Korean Research Foundation Grant (KRF-2008–331-E00385) and by the Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education, Science and Technology (2009–0074437).
Footnotes
a.
Invitrogen Corp., Carlsbad, CA.
b.
Dyne Bio Inc., Gyonggi-do, Korea.
c.
Enzynomics, Seoul, Korea.
d.
BigDye™, Applied Biosystems, Foster City, CA.
e.
3730x1, Applied Biosystems, Foster City, CA.
f.
Institut Génétique Biologie Moléculaire Cellulaire, Illkirch Graffenstaden, France.
g.
Biodesign Institute, Tempe, AZ.
h.
Invitrogen Corp., Carlsbad, CA.
