Abstract
Although the causative agent of bovine viral diarrhea was initially categorized as 1 species, phylogenetic analysis revealed that these viruses belong to 2 different species, Bovine viral diarrhea virus 1 (BVDV-1) and BVDV-2, with 2–11 subgenotypes within each species. Distribution of species and subgenotypes has been shown to vary with geographic region. Whether distribution shifts over time is not known. Surveys conducted between 1994 and 2008 reported 3 subgenotypes circulating among cattle in the United States: BVDV-1a, BVDV-1b, and BVDV-2a. The average percent prevalence of BVDV-1a, BVDV-1b, and BVDV-2a strains reported in surveys before 2001 were 21%, 43%, and 36%, respectively. Surveys conducted on viruses isolated after 2001 reported decreasing percentages of BVDV-1a and BVDV-2a strains, with BVDV-1b strains accounting for 75–100% of samples. Comparison of these surveys is confounded by differences in geographic location, collection methods, and sample type used in the survey. The purpose of the present study was to determine whether the prevalence of BVDV subgenotypes shifted in samples collected from the same geographic region and by the same laboratory over time. BVDV strains isolated in years 1988, 1998, and 2008, at the Texas Veterinary Medical Diagnostic Laboratory, Amarillo, Texas, were genotyped, and the prevalence of BVDV-1a, BVDV-1b, and BVDV-2a strains were determined. Typing, on the basis of phylogenetic analysis, was done on 148 samples. The strongest trend detected among these samples was a pronounced decrease in the number of BVDV-1a strains over time.
Introduction
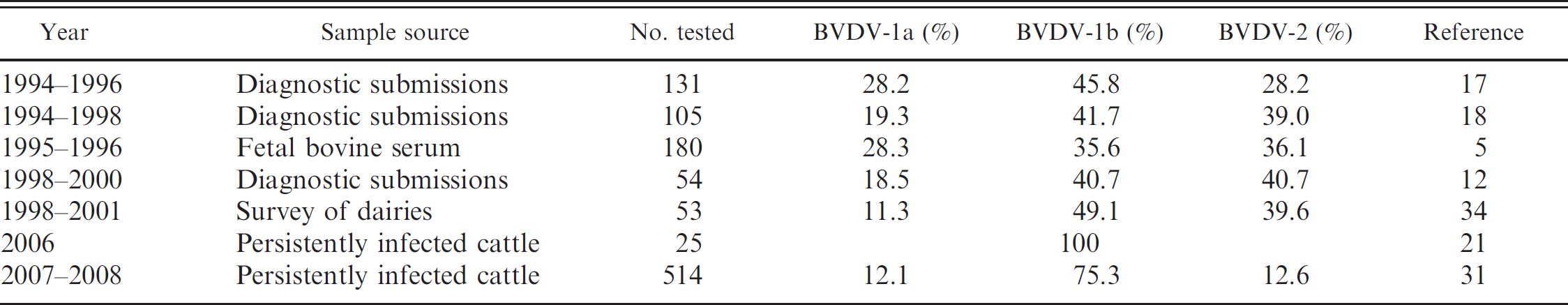
The viral agent associated with acute bovine viral diarrhea was first isolated in cell culture in 1954 2 and was named Bovine viral diarrhea virus (BVDV). Strains isolated in the next 10 years, such as NY-1, 2 C24V, 22 Singer, 25 and NADL 23 are used as reference and vaccine strains to this day. Although initially identified as a non-insect-borne togavirus, taxonomists later placed these viruses in the pestivirus genus of the Flaviviridae family along with Hog cholera virus (later renamed Classical swine fever virus) and Border disease virus. 24 Originally viral isolates were designated as BVDV on the basis of host origin so that any pestivirus isolated from cattle was referred to as BVDV. This approach proved to be untenable because it was discovered that the different pestivirus species could infect more than 1 host species. 13 Attempts to segregate the pestivirus species on the basis of serology were more successful 3,6,11,36,37,41 but were hampered by antigenic heterogeneity within species. 3,7,9,10,40 Phylogenetic analysis led to the division of BVDV strains into 2 species (BVDV-1 and BVDV-2) 26,30 and several subgenotypes within species (BVDV-1a–k and BVDV-2a and -2b). 14,35 Surveys conducted between 1994 and 2008 reported 3 BVDV subgenotypes circulating among cattle in the United States: BVDV-1a, BVDV-1b, and BVDV-2a (Table 1). The average percent prevalence of BVDV-1a, BVDV-1b, and BVDV-2a strains reported in surveys before 2001 were 21%, 43%, and 36%, respectively. 5,12,16,17,34 Surveys conducted on viruses isolated after 2001 reported lower percentages of BVDV-1a and BVDV-2a strains, with BVDV-1b strains accounting for 75–100% of samples. 15,20,31 Although these results suggest an increase in the proportion of BVDV-1b isolates with time, comparison of these surveys is confounded by differences in geographic location, method of collection, and type of sample used in the surveys. The presence and prevalence of BVDV species and subgenotypes affects vaccine design. One of the observations leading to the discovery of the BVDV-2 species was the birth of calves persistently infected with BVDV-2 strains to dams that had been vaccinated with BVDV-1–based vaccines. 30 Recognition of antigenic differences between BVDV-1 and BVDV-2 strains led to the development of bivalent BVDV vaccines that contained antigens derived from both BVDV-1a and BVDV-2a strains. 16 Antigenic differences have also been demonstrated among BVDV-1a, BVDV-1b, and BVDV-2a strains. 31 Although the BVDV-1b subgenotype is prevalent in the United States, most BVDV vaccines do not contain BVDV-1b–specific antigens.
Prevalence of Bovine viral diarrhea virus (BVDV) species and subgenotypes reported in various surveys conducted between 1994 and 2008.
The purpose of the current study was to determine whether the prevalence of BVDV subgenotypes shifted in the same geographic region in samples collected by the same laboratory over time. To this end, submissions to the Texas Veterinary Medical Diagnostic Laboratory, Amarillo, Texas, were analyzed over a 20-year period. The same methods of sample collection, storage, and virus isolation were used over the span of the study. BVDV strains isolated in years 1988, 1998, and 2008 were geno-typed, and the relative prevalence of BVDV-1a, BVDV-1b, and BVDV-2a strains was determined.
Materials and methods
Virus isolation and storage
All viruses were recovered on primary bovine fetal kidney (BFK) cell cultures by using the second to sixth subculture. All passages of BFK cells used for virus isolation were evaluated and confirmed negative for BVDV by direct fluorescent antibody test using commercially available BVDV polyclonal antiserum conjugated with fluorescein isothiocyanate. a Noncytopathic BVDV (BVDV1b-NY-1) was included as a positive control, along with all diagnostic samples and BVDV samples being propagated for testing. Negative cell culture controls were included with all virus isolation attempts and virus propagation work. The BFK cells were grown in Leighton tubes or 25-cm 2 flasks with Minimum Essential Medium and Earle salts, L-glutamine, nonessential amino acids, and sodium bicarbonate with 10% fetal bovine serum (FBS). All media contained 100 IU/ml penicillin, 100 μg/ml streptomycin, 100 μg/ml gentamicin, 10 μg/ml ciprofloxacin, and 1.0 μg/ml Amphotericin B. Before inoculation of virus samples, the medium was changed to contain only 2% FBS. The FBS used for cell culture was obtained from commercial sources and was gamma irradiated at 4.5 megarads. The FBS tested negative for BVDV by virus isolation with primary BFK cells and negative for BVDV antibody at 1:4 final serum dilution. All virus isolates were identified by direct fluorescent antibody testing with commercially available BVDV polyclonal antiserum conjugated with fluorescein isothiocyanate. a Virus isolates were stored frozen at −20°C to −70°C.
Selection of viruses for analysis
All isolates analyzed were isolated from bovine samples submitted to the Texas Veterinary Medical Diagnostic Laboratory, Amarillo, Texas. Accession numbers were assigned per submission rather than per animal, so that all animals submitted at one time from the same production unit had the same accession number. Thus, 1 positive accession can represent more than 1 animal. These isolations were made from tissues of animals submitted as a result of clinical presentation. No samples were generated as the result of BVDV surveillance efforts. The type of production unit in the area serviced by the diagnostic laboratory remained similar throughout the sampling period. The preponderance of cases were from cow/calf, feeder calf, and feedlot operations. However, it should not be assumed that only beef breeds were included. A significant proportion of animals in feeder calf and feedlot operations originate as dairy bull calves and in some lots could actually make up the majority population.
In 1988, virus isolations were performed for 829 accessions. Of these, 145 were positive (17%) for BVDV and no other virus. Similarly, in 1998, of 671 accessions, 57 (8.5%) were positive for BVDV and no other virus, and in 2008, of 666 accessions, 44 (6.6%) were positive for BVDV and no other virus. All viruses that were still viable in 2008 were sent to the NADC for phylogenetic analysis—a total of 66 viruses from 1988, 44 viruses from 1998, and 44 isolates from 2008.
The reasons for loss of viability could not be determined, and it is not known if any 1 BVDV species and subtype were more or less fit under the storage conditions used. The NADC BVDV strain repository consists of more than 5,000 strains collected between the mid-1960s to the present. No difference in viability associated with species or subgenotype has been noted in viruses in that repository, although this does not rule out a possible difference in viability; however, no reports of this appear in the literature, and no observed phenotypic differences result in reduced viability in storage.
Polymerase chain reaction amplification and sequencing
A total of 154 strains (66 isolated in 1988, 44 in 1998, and 44 in 2008) were analyzed. References strains BVDV1a-Singer, BVDV1a-NADL, BVDV1a-C24V, BVDV1b-NY1, BVDV1b-TGAN, BVDV2a-296c, and BVDV2a-5912 were also included. Genotyping was based on comparison of sequences from the 5′ untranslated region (5′-UTR). A 140-μl aliquot of each viral sample was used to prepare RNA with a viral RNA mini-kit b per the manufacturer's directions. The final volume of RNA template solution was 100 μl. A 10-μl aliquot of this solution was used to amplify a 280-nucleotide region of the 5′-UTR by reverse transcription polymerase chain reaction (RT-PCR) as described previously. 29 All sequencing reactions were done in duplicate, and all sequences were confirmed by sequencing both strands. The PCR products were quantitated with the Pico Green assay for dsDNA. c The appropriate quantity of dsDNA PCR product was labeled in both directions with terminator chemistries d according to manufacturer's instructions. The labeled products were sequenced with the use of a genetic analyzer. d Sequences were aligned and compared on the basis of the Higgins–Sharp algorithm (CLUSTAL4) by the MacDNASIS program. e This program takes as input a dendrogram produced by applying the unweighted pair group method using arithmetic average (UPGMA) 32 to a matrix of similarity scores for all the aligned sequences. The similarity scores are calculated as the number of exactly matched residues in a Wilbur and Lipman 38 alignment minus a fixed penalty for every gap.
Results
The number of accessions and animals positive for BVDV and the percentage of accessions positive for BVDV in the years 1984–2008, are shown in Figure 1. During a 4-year period between 1986 and 1989, an increased number of BVDV isolations were made (Fig. 1a). The average of BVDV-positive isolations during that 4-year period was 132, compared with an average 61 isolates for all other years. The number of accessions submitted for virus isolation was also higher between 1986 and 1989 (Fig. 1b). However, less variation was seen in the percentage of accessions positive for BVDV (Fig. 1c). Breaking down the data by decade, the average percentage of BVDV-positive accessions in the years 1984–1989 was 12.5%. The average for years 1990–1999 was 11.1%, and the average for years 2000–2008 was 11.8%. According to the least significant difference method 33 for analyzing variance, there was no overall difference among the averages by decade (P > 0.08).
The phylogenetic analysis of isolates from 1988, 1998, and 2008 are shown in Figure 2. Sequences generated from isolates were compared with vaccine strains C24V (BVDV-1a), NADL (BVDV-1a), Singer (BVDV-1a), NY-1 (BVDV-1b), 296c (BVDV-2a), and 5912 (BVDV-2a) and laboratory reference strain TGAN (BVDV-1b). Samples from each of the 3 years analyzed had sequences identical to vaccine strains. This outcome was interpreted as a reisolation of a vaccine strain. In 1988 (Fig. 2a), 2 isolates were identical to C24V and 2 were identical to Singer. In 1998, 4 isolates were identical to C24V (Fig. 2b). In 2008, 2 isolates were identical to C24V and 10 were identical to Singer (Fig. 2c). The number of samples containing multiple BVDV strains and the number of samples that were reisolations of vaccine strains were subtracted from the total number of isolates received to determine the number of field strains analyzed (Table 2). In addition to isolates that were identical to vaccines strains, groups of isolates were identical to each other (groups A–R in Fig. 2a–c). In 9 of these groups, the isolates were derived from animals submitted by the same owner but at different times. No obvious connection could be made on the basis of accession records for the remaining 9 groups. Isolates that contained more than 1 BVDV strain, as determined by mixed sequence chromatograph results, were not included in the phylogenetic analysis (Table 2), and groups of identical field strains were only counted once in the determination of prevalence. The prevalence of BVDV species and subgenotypes among unique field isolates in each of the years 1988, 1998, and 2008 is summarized in Table 2. Although the percentage of BVDV-1b and BVDV-2a strains increased from 1988 to 2008, the number of isolates segregated to these 2 subgenotypes did not show large year-to-year variation. In contrast, both the number and percentage of BVDV-1a strains showed a decline from 1988 to 2008.
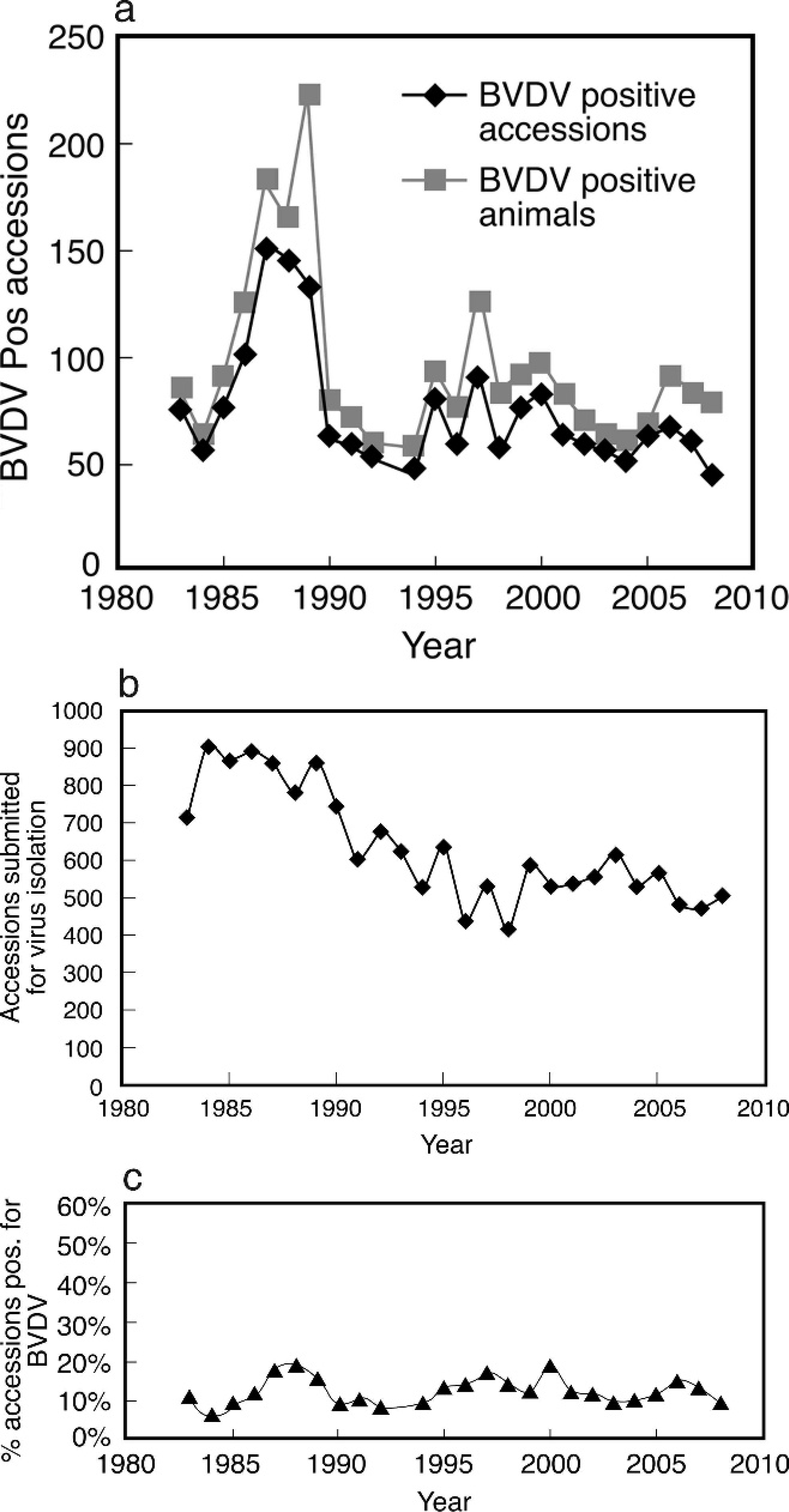
Submissions to the Texas Veterinary Medical Diagnostic Laboratory, Amarillo, Texas, made between 1983 and 2008, broken down by number of accessions, total number of accessions positive for Bovine viral diarrhea virus (BVDV), and percentage of submissions positive for BVDV.
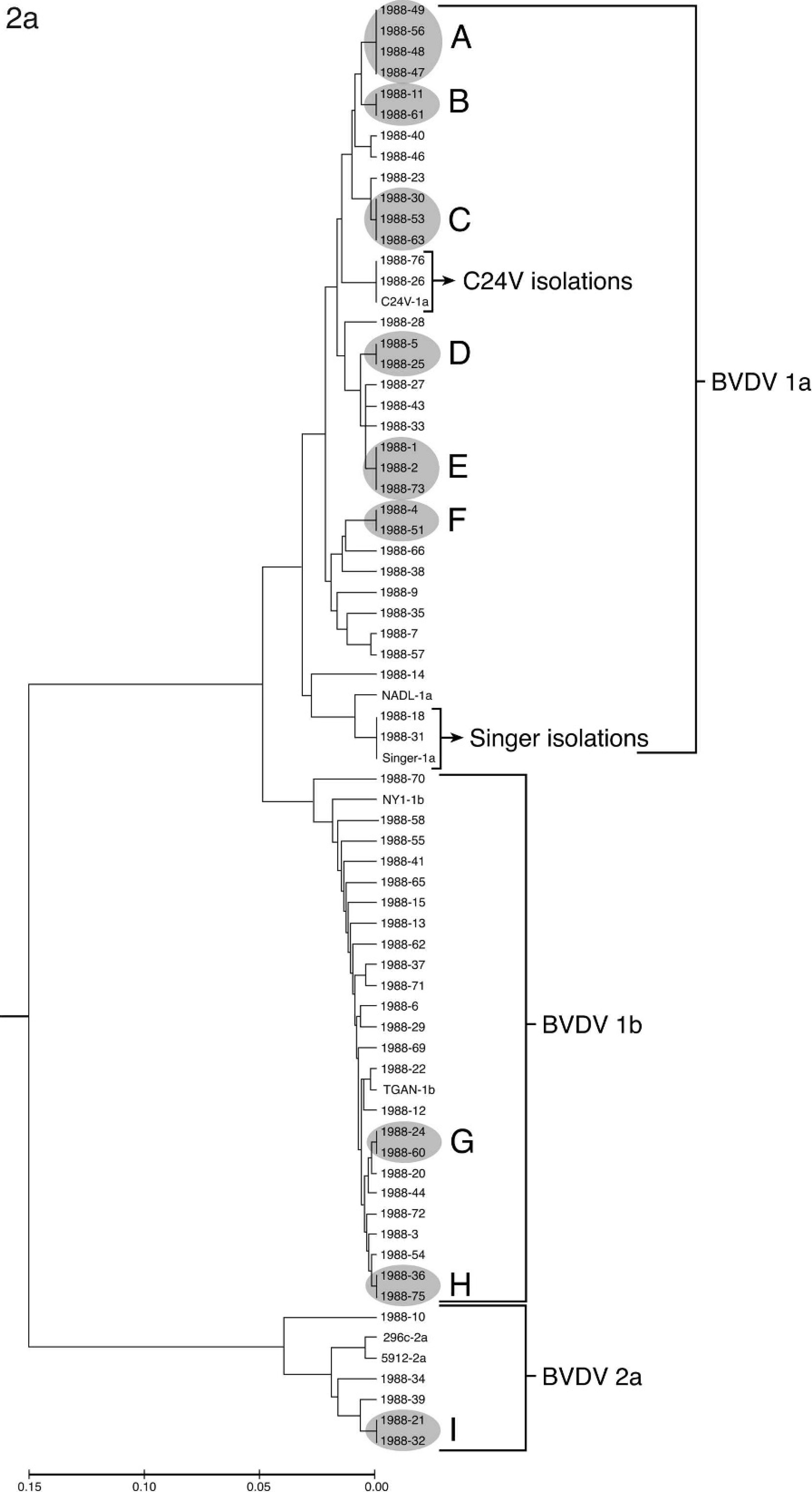
Phylogenetic analysis of Bovine viral diarrhea virus (BVDV)–positive accessions from the years 1988 (
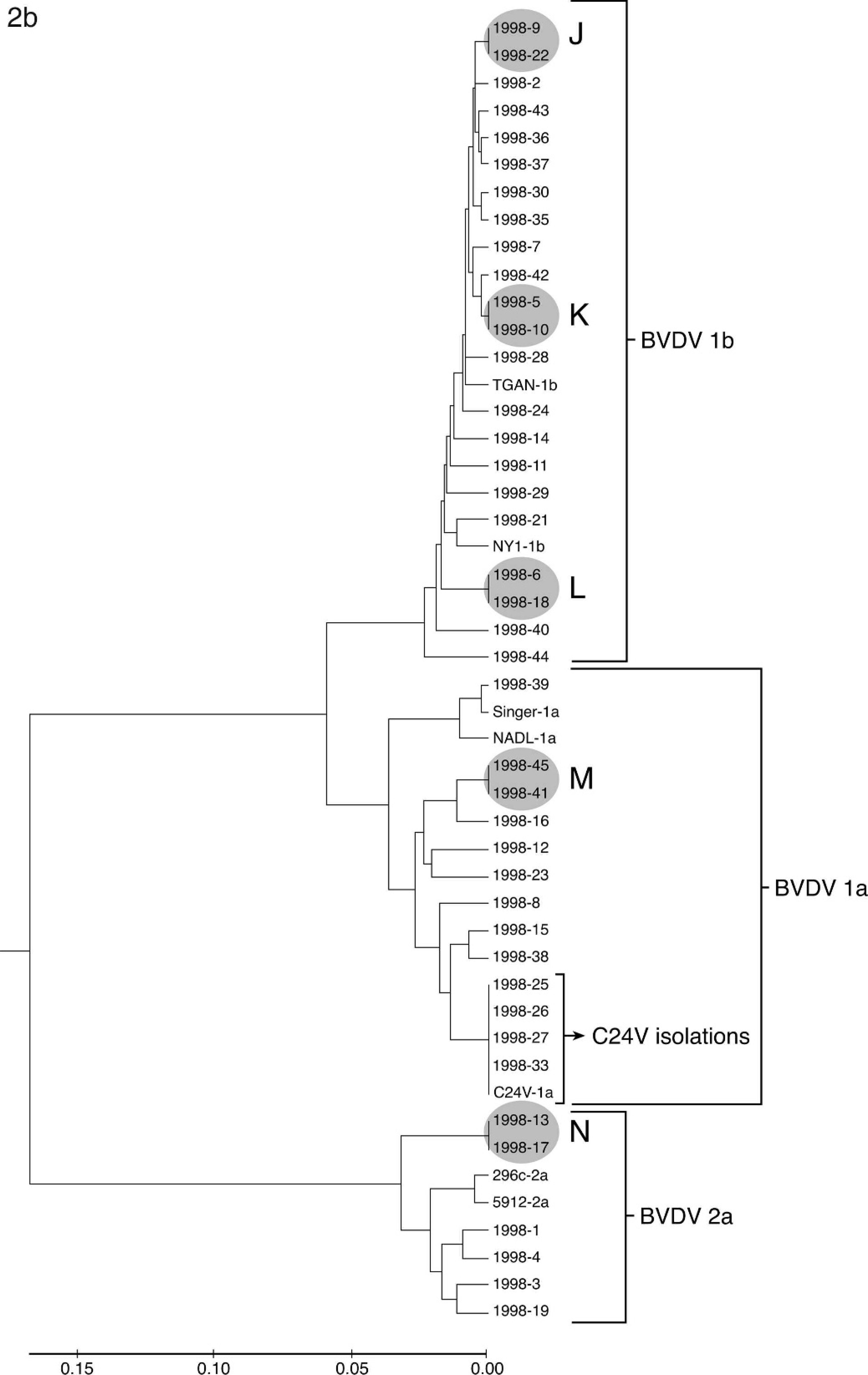
Continued.
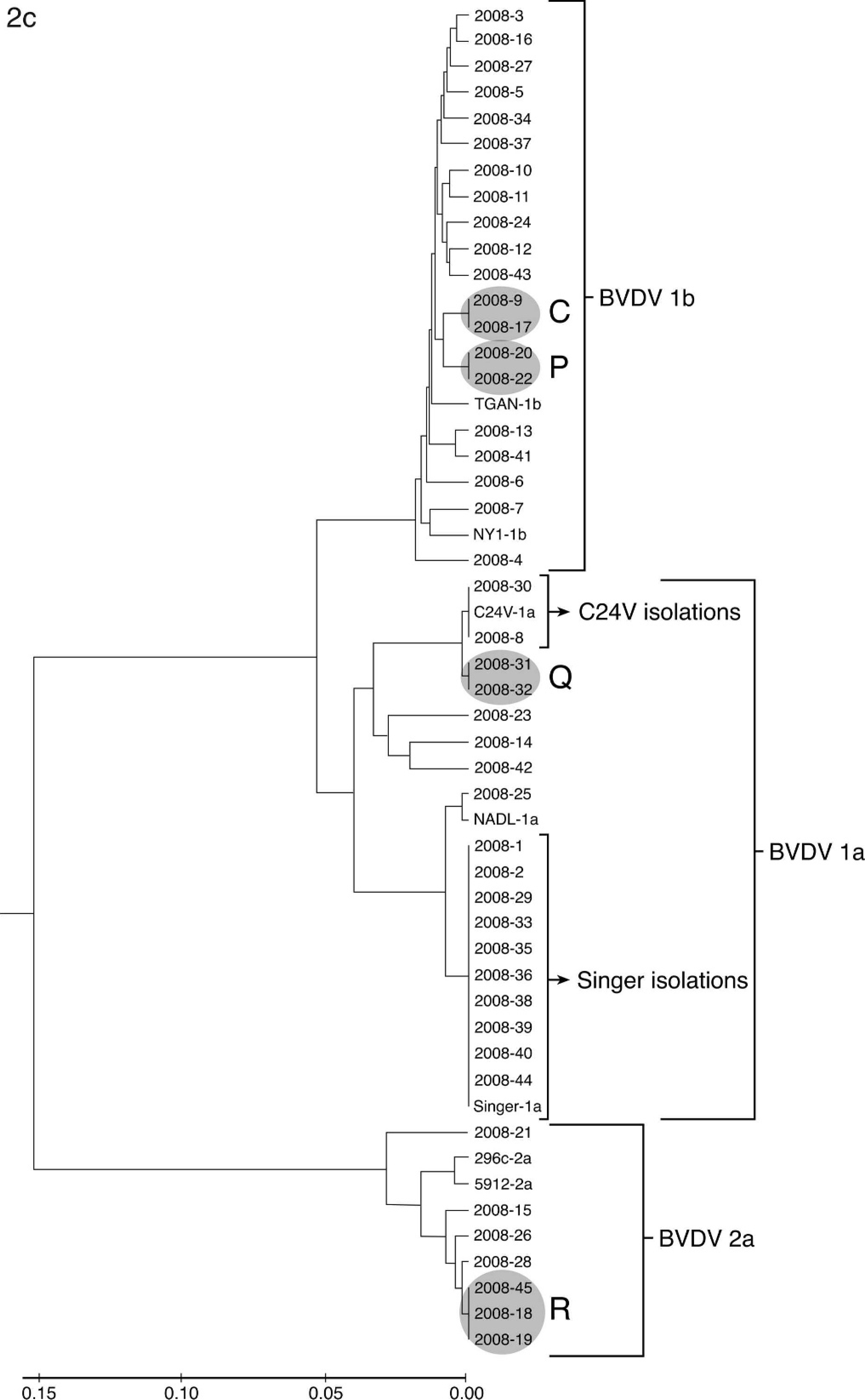
Continued.
Summation of genotyping results from Bovine viral diarrhea virus (BVDV)–positive samples submitted in 1988, 1998, and 2008.*
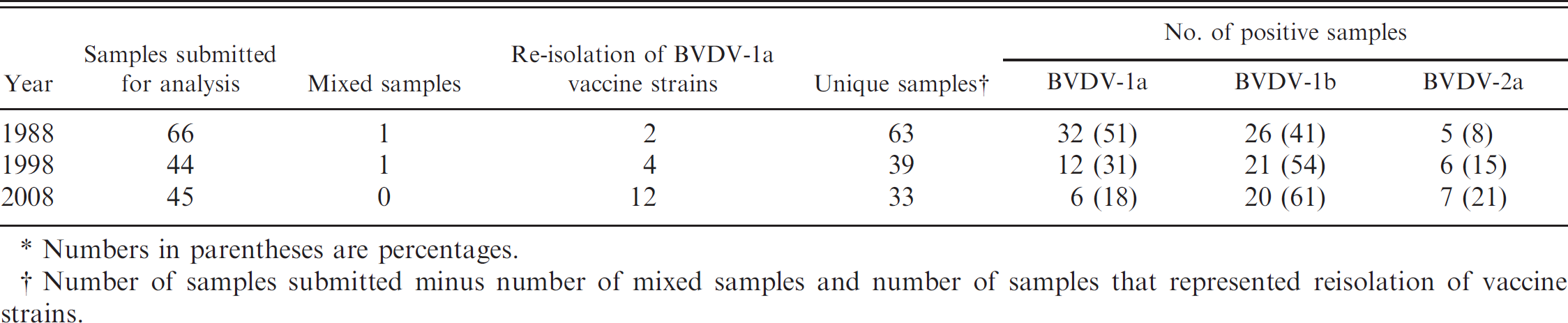

Numbers in parentheses are percentages.
Number of samples submitted minus number of mixed samples and number of samples that represented reisolation of vaccine strains.
Discussion
The present study does not reflect incidence of acute or persistent BVDV infection but instead looks at the predominance of BVDV species and subgenotypes identified in diagnostic samples at 3, 10-year intervals. A decline in both the number and relative percentage of BVDV-1a strains detected among samples submitted to a regional laboratory was observed. This finding is consistent with a shift in the predominance of BVDV-1a strains reported in published surveys. 5,12,16 – 18,21,31,34 Although the reason for the decline in BVDV-1a strains cannot be extrapolated from this data, one hypothesis is the effectiveness of vaccines based on BVDV-1a strains in preventing infection with BVDV-1a strains. The largest selling BVDV vaccines all contain BVDV-1a antigens, a portion include BVDV-2a strains, and a minority include BVDV-1b strains. 16 The first cytopathic BVDV isolated, 22 Oregon C24V (also known as C24V), was a BVDV-1a strain, 16 and it was the basis for the first mass-produced modified live vaccine. 27 Two other cytopathic strains incorporated in early vaccines, the Singer and NADL strains, also belong to the BVDV-1a subgenotype. 16 These 3 virus strains are still used in vaccines in 2010. Of the 9 largest selling modified live BVDV vaccine lines, 8 contain 1 of these 3 virus strains. 16 The ninth line contains a different BVDV-1a strain. In addition to a BVDV-1a strain, 4 of the 9 lines also include BVDV-2a antigens. Similarly 7 of 8 of the largest selling killed vaccine lines contain antigens derived from the C24V, Singer, or NADL strains. Four of these 7 also contain BVDV-2a antigens. The only killed vaccine line that does not contain C24V, Singer, or NADL contains BVDV-1a and BVDV-2a antigens from strains unique to that company. One of the killed vaccine lines contains Singer antigen and NY-1 antigen (a BVDV-1b strain) but no BVDV-2a antigen. Thus, none of the 17 top-selling modified live and killed vaccine lines contain antigens from all 3 of the BVDV subgenotypes circulating in U.S. cattle populations, and only 1 vaccine line contains antigens from the predominant subgenotype, BVDV-1b.
Although the predominance of BVDV-1a strains decreased in this study from 1988 to 2008, studies conducted from 1985 to 2008 indicate the incidence of BVD, based on percentage of herds harboring animals persistently infected (PI) with BVDV, did not. A 1985 study reported a herd incidence of 9.0%. 4 A 1996 study, published in 2001, reported a herd incidence of 19% in BVDV-suspect herds and a 4% incidence in randomly selected herds. 39 The National Animal Health Monitoring System of Beef 2007–2008 study reported a herd incidence of 8.8%. 8 It should be noted however, that incidence of PI does not reflect the true incidence of BVDV because it ignores acute infections. Thus, although a lower incidence of BVDV-1a isolations has been reported, there does not seem to be a reduction in the incidence of BVDV.
The advent of PCR and antigen capture enzyme-linked immunosorbent assay (ACE) testing has promoted increased testing, but this is a double-edged sword. As can be seen from the present data, the number of samples being submitted for virus isolation has decreased. Although more animals might be tested by PCR and ACE, because of speed and economy, these 2 tests do not generally yield information on species and subgenotypes or allow the comparison of viruses. In the present study, 18 groups of identical viruses were identified. Of these, 9 groups of viruses could be traced to animals belonging to 1 owner. The available records reveal no known connection between the infected animals identified in the other 9 groups.
The practical implications of dividing BVDV strains into subgenotypes lies in the differences observed in antigenicity. Studies have shown antigenic differences between subgenotypes based on differences in cross-neutralization, 1,28,31 monoclonal antibody binding, 5 and response of persistently infected animals to vaccination. 19 The widespread use of BVDV-1a–based vaccines and the decline in the prevalence of BVDV-1a strains, while incidence of BVDV-1b strains remains unchanged, suggests that control of BVDV might be improved by the use of vaccines that contain BVDV-1b antigens in addition to BVDV-1a antigens. Although further research is required, including antigens from all BVDV subgenotypes circulating in the region in which the vaccine is to be used is an idea worth exploring. It should be noted that improved vaccines are only part of the solution to improved BVDV control. Successful control programs must also provide surveillance to remove PI animals and biosecurity measures that prevent the introduction of BVDV into herds.
Acknowledgements
The authors thank Patricia Federico and Kathryn Fulk for excellent technical support, Harold Ridpath for statistical consultation, Michael Marti for preparing figures, and Sandy Johnson for manuscript preparation.
Footnotes
a.
VMRD, Pullman, WA.
b.
Qiagen Inc., Valencia, CA.
c.
Invitrogen Corp., Carlsbad, CA.
d.
Applied Biosystems Inc., Foster City, CA.
e.
Hitachi Software, San Bruno, CA.
