Abstract
The sequences of short fiber genes of the
Introduction
Hydropericardium syndrome (HPS) is a disease of chickens, first reported
5
in Pakistan in 1987. Since then, HPS has been reported in Iraq, Kuwait, India, Mexico, Ecuador, Peru, Chile, Korea, and Japan.
1–4,8,11,14
From these cases, an adenovirus, which was either isolated from, or visualized electron microscopically, in the liver of affected chickens, has been implicated in the syndrome.
6
The syndrome has been reproduced by inoculation of isolated
Functionally, the fiber of the adenovirus is important in virus infectivity. 28 Briefly, type-specific neutralizing, subgenus-specific neutralizing, and type-specific nonneutralizing epitopes all exist within the fiber. Furthermore, the adenovirus fiber has been shown to be responsible for binding to the cell surface and to be related to the tissue tropism. 10,16,17,19,20 However, the analyses of fiber genes of HPS–FAdV-4 strains have not been fully performed.
The purposes of the current study were to compare the fiber genes and pathogenicity to chickens of FAdV-4 isolates from HPS in chickens in Japan, India, and Pakistan with conventional FAdV-4 strains to increase understanding of the molecular characteristics of highly pathogenic HPS–FAdV-4 strains and to develop a simple and easy method of differentiation between HPS–FAdV-4 isolate and non-HPS–FAdV-4 isolates.
Materials and methods
Viruses
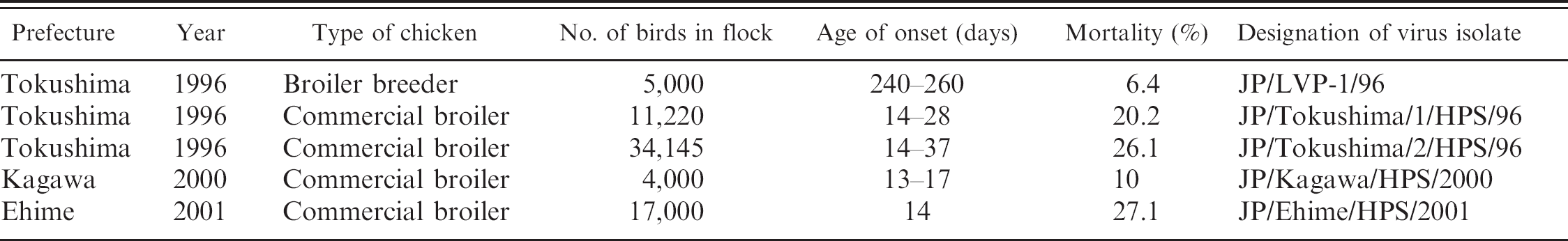
The case history of the HPS outbreaks in chickens in Japan used in the present study is summarized in Table 1. Primary chicken kidney cell (CKC) cultures were routinely used for isolation and propagation of the adenovirus. Livers from chickens naturally affected by HPS between 1996 and 2001 were ground with sterile sand and diluted with maintenance medium to make a 10% (wt/vol) suspension. The homogenates were then centrifuged at low speed (3,000 ×
The conventional FAdV-4 strains (KR-5, 828-3, 1078-5, 1081-2, and 1262-4) used in the current study were previously collected. 12 Among them, the KR-5 strain is the reference strain of FAdV-4. The following FAdV isolate from the HPS were used: H-1 strain a (India); PARC-1/96, PARC-1/97, and PARC-1/98 (Pakistan). b
Summary of hydropericardium syndrome cases from Japan used in the current study.
Polymerase chain reaction and phylogenetic analysis
To determine the genetic relationship between the HPS–FAdV-4 strains and other viruses, complete nucleotide sequences of the short fiber gene of the isolates were determined. Viral DNA from infected culture fluids was extracted using a commercial kit. a Primers were designed based on a comparison of the sequences of the short fiber gene of FAdV-4 and FAdV-10 (GenBank accession AY340863 and AF007579, respectively) because these strains were demonstrated to have a close genetic relationship with each other. 9 The following primer set was used in the current study: FibF1 (5'-CAGGGTTACGTCTACTCCCC-3'; sense) and FibR1 (5'-TTTGTCACGCGGTGGGGAGG-3'; antisense).
Polymerase chain reaction (PCR) was performed using 35 cycles of 94°C for 30 sec, 55°C for 30 sec, and 72°C for 30 sec. The predicted size of the PCR products was approximately 1,500 base pairs (bp) of the entire short fiber gene. The derived nucleotide sequences were analyzed using commercial software b and through GenBank searches. The phylogenetic analysis was conducted using the ClustalX program, 27 and the tree was constructed by the neighbor-joining (NJ) method. 24
Polymerase chain reaction–restriction fragment length polymorphism
The generated PCR products were used in restriction endonuclease analysis for the development of a simple and rapid differentiation of FAdV-4 strains. The enzyme
Pathogenicity to 1-day-old chicks of FAdV-4
The pathogenicity to 1-day-old chicks of selected FAdV-4 isolates of each cluster in the phylogenetic analysis based on the short fiber gene was examined. The 1-day-old chicks used in the current study were obtained from a specific-pathogen–free White Leghorn flock (line PDL-1). Throughout the experimental period, the chicks were kept in isolators that were placed in a negatively pressured house with high-efficiency particulate air-filtered intakes and exhaust. Ten chickens for the selected FAdV-4 strains from each genetic cluster were inoculated orally with 0.1 ml of diluted infectious culture fluids containing 106 50% tissue culture infective dose (TCID50) of test virus. All chicks were observed daily for 14 days, and clinical signs and/or deaths were recorded. All experimental procedures and animal care were performed in compliance with the guidelines of the National Institute of Animal Health for the humane use of laboratory animals.
Results
Molecular characterization and phylogenetic analysis based on short fiber gene
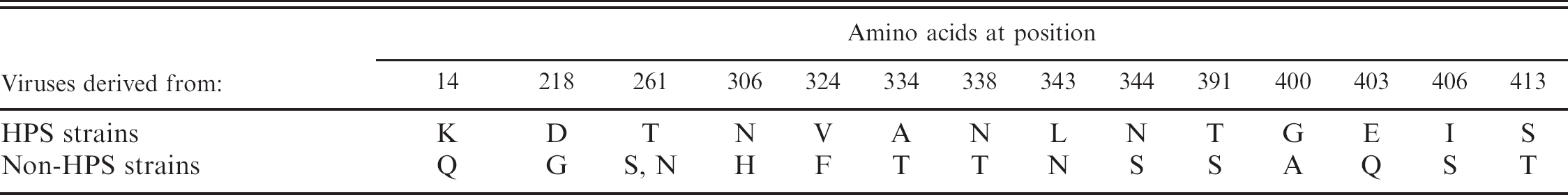
A previous study indicated that the fiber gene is responsible for variations of virulence in the case of FAdV-8 23 ; therefore, the amino acid sequences of this protein in the FAdV-4 isolates used in the present study were compared. When predicted amino acid sequences for the HPS–FAdV-4 strains and those not derived from FAdV-4 were compared, certain amino acids appeared to be specific to HPS–FAdV-4 viruses (Fig. 1; Table 2). Typical knob sequences of all FAdV-4 strains examined in the current study were not identified like FAdV-1 and FAdV-10. 7,9 By phylogenetic analysis based on the short fiber gene with available sequences from GenBank, HPS–FAdV-4 strains were divided into a different cluster from non-HPS–FAdV-4 strains (Fig. 2).
PCR-RFLP analysis
All HPS–FAdV-4 strains showed identical
Pathogenicity to 1-day-old chicks
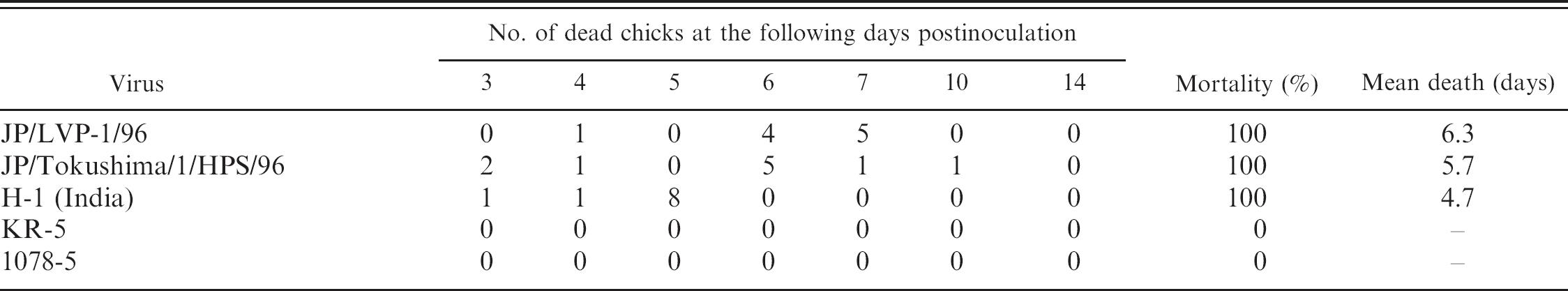
Only the HPS–FAdV4 strains showed 100% mortality in 1-day-old chicks (Table 4). The chicks infected with HPS–FAdV-4 (JP/LVP-1/96, JP/To-kushima/1/HPS/96, and H-1 strains) died in 3 to 8 days. None of the chickens inoculated with conventional FAdV-4 strains (KR-5 and 1078-5) died, and no clinical signs were observed.
Discussion

Multiple alignment of deduced amino acids of short fiber protein of
Differences in amino acids between
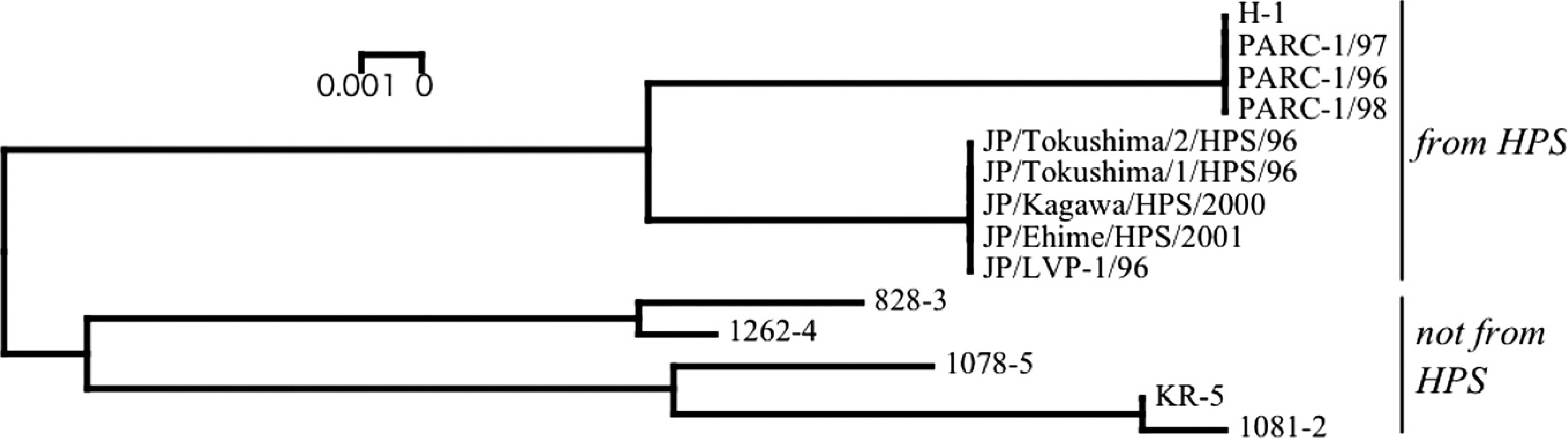
Phylogenetic tree based on the short fiber gene of
In the present study, the PCR primer pair (FibF1 and FibR1) successfully amplified approximately 1,500-bp products from DNA samples of all FAdV-4 isolates employed in the current study. The nucleotide sequences of all HPS–FAdV-4 isolates in Japan were identical. Isolates from India and Pakistan were also identical, but different from those of Japan with a homology of 98.5% at the nucleotide sequence level. The predicted amino acid sequences were compared by determining the complete nucleo-tide sequences of short fiber genes of HPS–FAdV-4 strains and those not derived from HPS–FAdV-4. Although the function of the short fiber protein is not fully understood, several amino acids specific to HPS–FAdV-4 used in the current study were observed and could prove to be useful as molecular markers of HPS–FAdV-4.
Predicted size of polymerase chain reaction–restriction fragment length polymorphism analysis by
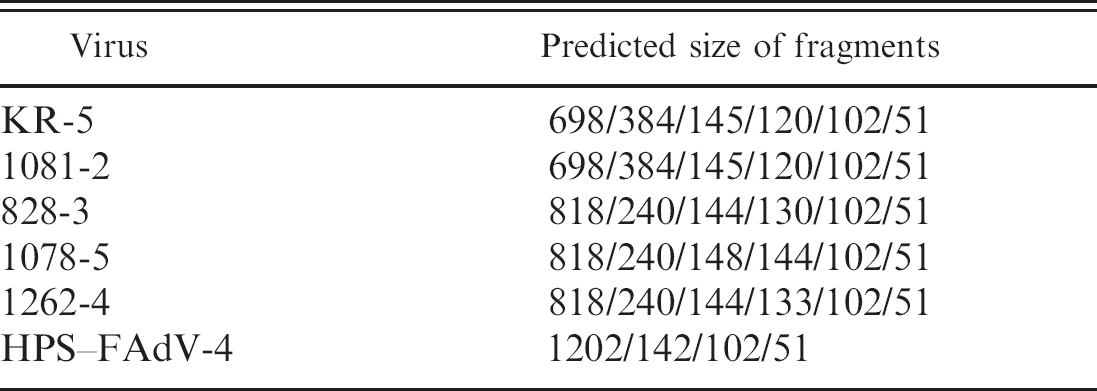
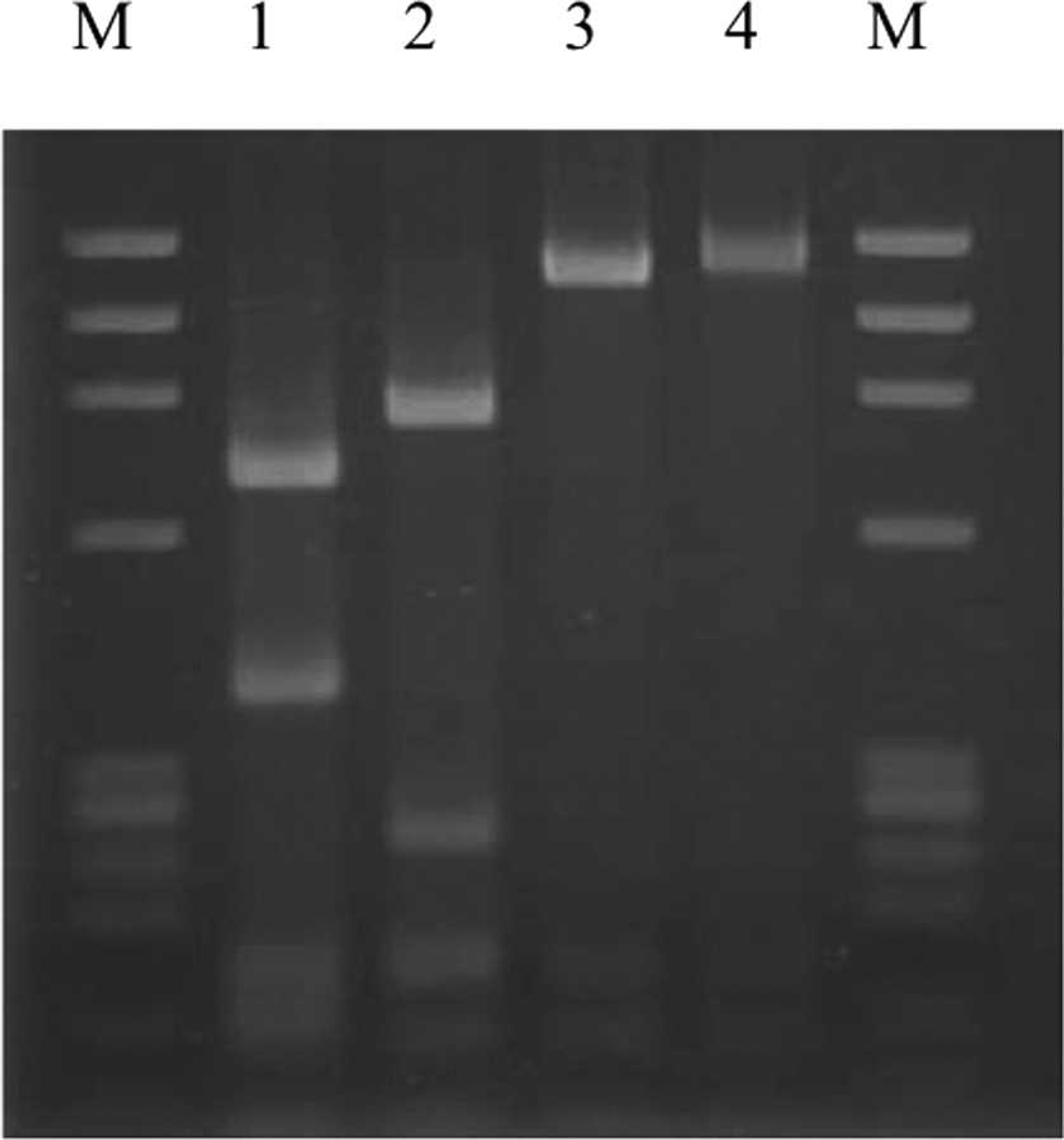
Restriction endonuclease analysis of the polymerase chain reaction–amplified short fiber genes from the four representative genetic groups of
Mortality in 1-day-old chicks inoculated with
By phylogenetic analysis based on the short fiber gene, the FAdV-4 strains used in the present study were classified into 2 clusters, namely the HPS–FAdV-4 group and non-HPS–FAdV-4. The origin of Japanese HPS–FAdV-4 seemed to be different from the Indian strains; these 2 groups were easily differentiated by RFLP pattern using enzyme
The typical knob sequences of all FAdV-4 examined herein were not evident, as in the FAdV-10 previously reported, 25 which has a close genetic and antigenic relationship with FAdV-4. 9 The lack of knob regions responsible for binding to the cell surface in the short fiber gene of FAdV-4 is interesting for its importance in the analysis of other genes. Particularly, it was discovered that the long fiber was important for the in vivo biology in the case of the FAdV-1 CELO strain. 26 However, though the complete genomes of some fowl adenoviruses have been determined, 7,21 no complete FAdV-4 strain genome has been determined. To identify the critical sites responsible for high pathogenicity of HPS–FAdV-4, further molecular analyses of each gene would be required.
Acknowledgements
The authors would like to thank the veterinary officials of Tokushima, Kagawa, and Ehime prefectures, Japan, for their cooperation in the collection of viral samples. Furthermore, the authors thank Dr. M. Qasim Khan, Animal Health Institute, National Agricultural Research Centre, Islamabad, Pakistan, and Dr. R. Manickam, Tamil Nadu Veterinary and Animal Sciences University, Chennai, India, for providing viruses.
Footnotes
a.
Kindly provided by Dr. R. Manickam, Tamil Nadu Veterinary and Animal Sciences University, Madhavaram Milk Colony Campus, Chennai, India.
b.
Kindly provided by Dr. M. Qasim Khan, Animal Health Institute, National Agricultural Research Centre, Islamabad, Pakistan.
c.
QIAamp® DNA micro kit, Qiagen Inc., Valencia, CA.
d.
GENETYX®-Mac version 13.0, Software Development Corp., Tokyo, Japan.
e.
Takara Co. Ltd., Tokyo, Japan.
