Abstract
In November 2013, a fatal encephalomyocarditis virus (EMCV) case in a captive African elephant (Loxodonta africana) occurred at the Réserve Africaine de Sigean, a zoo in the south of France. Here we report the molecular characterization of the EMCV strains isolated from samples collected from the dead elephant and from 3 rats (Rattus rattus) captured in the zoo at the same time. The EMCV infection was confirmed by reverse-transcription real-time PCR (RT-rtPCR) and genome sequencing. Complete genome sequencing and sequence alignment indicated that the elephant’s EMCV strain was 98.1–99.9% identical to the rat EMCV isolates at the nucleotide sequence level. Phylogenetic analysis of the ORF, P1, VP1, and 3D sequences revealed that the elephant and rat strains clustered into lineage A of the EMCV 1 group. To our knowledge, molecular characterization of EMCV in France and Europe has not been reported previously in a captive elephant. The full genome analyses of EMCV isolated from an elephant and rats in the same outbreak emphasizes the role of rodents in EMCV introduction and circulation in zoos.
Introduction
Encephalomyocarditis virus (EMCV; Picornaviridae, Cardiovirus, Cardiovirus A) 20 is an RNA virus that was first isolated in 1945 in Florida from a captive male gibbon that died suddenly of pulmonary edema and myocarditis. 15 Since that time, EMCV has been recognized worldwide as a pathogen infecting many domestic and wild animals. Depending on the strain and host, EMCV can induce myocarditis, reproductive failures, diabetes, or nervous disorders. 6 Although disease transmission is poorly understood, rodents are considered to be natural hosts of EMCV and are thought to be the primary reservoir and vectors of the virus. 34 The primary route of transmission for EMCV is believed to be fecal–oral, through ingestion of food or water contaminated with the virus. 46 Pigs are considered the most commonly and severely infected domestic animal. The virus can cause acute myocarditis and sudden death in pre-weaned piglets, or severe reproductive failure in breeding sows. In Europe, severe EMCV outbreaks have been reported in Italy, Greece, Belgium, and Cyprus.10,21,24,29,30,36,42 An investigation in France of pig sera for antibodies against EMCV, carried out in 2000 and 2001, revealed an overall seroprevalence of EMCV infection of 2.5%. 1
As well as outbreaks in pigs, several cases of fatal EMCV infections involving a variety of exotic mammals have been described in zoos in Australia, the United States, Italy, Russia. and Singapore.4,13,31,43,44,45 Sudden death was often the first indication of infection and was associated with acute fatal myocarditis. Primates appear to be particularly susceptible, and the disease has been reported in chimpanzees, orangutans, semi-wild bonobos, baboons, lemurs, and rhesus macaques.4,9,17,19,25,31,45 EMCV infections have also been repeatedly implicated worldwide in the deaths of elephants. In 1976–1977, 4 African elephants in 2 zoological gardens in Florida died of a fulminant illness caused by EMCV. 37 In 1984, the virus was incriminated in the death by myocarditis of a female African elephant in an Australian zoo. 35 A spontaneous outbreak of fatal EMC was also reported in free-ranging African elephants in the Kruger National Park, South Africa, in 1993–1994. 14
We report here the isolation and molecular characterization of EMCV from a captive African elephant (Loxodonta africana) that died suddenly in a French zoo. 22 After postmortem investigations, EMCV was isolated from the elephant’s heart, spleen, liver, and blood. It was also isolated from the stools and ileum of rats captured on the zoo grounds at the same time. Full genome sequence analyses revealed a high homology between the elephant and rat strains, suggesting transmission of EMCV between the 2 species and thus strengthening the hypothesis that EMCV was introduced by rats.
Materials and methods
Case presentation
In October 2013, 2 female African elephants (A and B) from the Réserve Africaine de Sigean, a zoo in the south of France, were exchanged with 2 female elephants (C and D) from the Thuringian Zoo Park in Erfurt, Germany. In November 2013, 1 mo after the transfer, elephant D—an 18-y-old wild-born female African elephant—was found dead in the morning without any preliminary clinical signs. No clinical signs were noted in the other 2 African elephants living at the same zoo. 22
Samples
After autopsy of the dead elephant, blood and tissue samples from the heart, spleen, and liver were collected and stored at –80°C before shipment to the laboratory for analyses. Mesenteric lymph nodes, ileum, and/or fecal samples were collected from 21 rats captured on the zoo grounds in the weeks following the elephant’s death and stored at –80°C. Five rats (2, 3, 10, 11, and 14) were captured in the elephant barn. Serum samples were collected from a further 3 captured rats (9, 20, and 21) and stored at –20°C. Serum samples were collected from elephants before and after their transfer and from 99 animals representing 28 species between March 2014 and February 2016; sera were stored at –20°C.
Virus isolation and titration
Ten percent homogenates of tissue samples and feces were prepared in phosphate-buffered saline (pH 7.2) and the debris cleared by centrifugation at 3,000 × g for 10 min. To isolate the virus, 150-µL aliquots of clarified supernatant fluids were inoculated onto monolayers of baby hamster kidney (BHK-21) cells prepared in 24-well, flat-bottom culture plates using minimal essential medium (MEM; Invitrogen), 2% fetal bovine serum (Eurobio), 1% nonessential amino acids (Invitrogen), 1% sodium pyruvate, and antibiotics (1 U/mL of penicillin and 1 μg/mL of streptomycin; Invitrogen). The plate was incubated at 37°C under 5% CO2 atmosphere. Cell cultures were monitored daily for cytopathic effects (CPE) over 72 h. Monolayers negative for CPE were passaged once more to confirm negative virus isolation results.
Virus was quantified in samples by endpoint dilution using 8 wells per virus dilution as described previously. 5 CPE were evaluated 3 d after infection. The titers were expressed as the median tissue culture infective doses per mL (TCID50/mL).
Reverse-transcription real-time PCR
Total RNA was extracted from tissue samples and feces homogenate (QIAamp viral RNA mini kit; Qiagen), according to the manufacturer’s recommendation. RNA was eluted in 60 µL of elution buffer. The EMCV genome was detected by an in-house one-step reverse-transcription real-time PCR (RT-rtPCR; AgPath-ID one-step RT-PCR kit; Life Technologies), using EMCV-specific primers, and a TaqMan probe located in a conserved region of the 3D gene of the viral genome (EMC-TMF1: 5′-GGGATCAGCTTTTACGGCTTT-3′, EMC-TMR1: 5′-TGCATCCGATAGAGAACTTAAT GTCT-3′, EMC-TMP: 5′-FAM-CGATGCCAACGAGGACGCCC-MGB-3′). The RT-rtPCR was multiplexed for simultaneous detection of β-actin as an endogenous control with a VIC reporter dye at the 5’-end of the probe. 41 The RT-rtPCR was performed in reaction mix containing 5 µL of RNA, 12.5 µL of buffer (2×), 1 µL of enzyme mix (25×), 1 µL of EMC-TMF1 (10 µM), 1 µL of EMC-TMR1 (10 µM), 0.4 µL of EMC-TMP (10 µM), 1 µL of β-actin-F (10 µM), 1 µL of β-actin-R (10 µM), 0.6 µL of β-actin-P (5 µM), and H2O to make 25 µL. The reaction mixture was placed in a thermocycler (ABI 7300; Life Technologies) to perform the reverse transcription for 10 min at 45°C, followed by 10 min of denaturation at 95°C, 45 denaturation cycles of 15 s each at 95°C, and 1 min annealing-extension at 60°C. RNA that was extracted from EMCV BEL-2887A/91–infected BHK-21 cells was used as positive control. Fluorescence emissions were measured during the annealing-extension step. The reporter dye (FAM and VIC) signal was measured relative to the internal reference dye (ROX) signal to normalize for non–PCR-related fluorescence fluctuations occurring from well to well.
Virus neutralization test
The sera were tested for antibodies to EMCV by the microtiter virus neutralization test (VNT). The test was carried out using a 2-fold dilution series of the heat-inactivated sera in 96-well, flat-bottom culture plates, starting at 1:8. The virus was diluted in MEM to deliver 100 TCID50 per well in 50 μL. Eight wells were used in duplicate for each serum (2 wells per serum dilution). The virus and 50 μL of diluted serum solution were incubated for 1 h at 37°C. Finally, 100 μL of BHK-21 cell suspension containing 3 × 105 cells/mL was added. The cultures were examined for CPE after incubation for 72 h at 37°C. The sera titers were expressed as the initial dilution of the sera at a 50% endpoint. The sera were considered positive if the titer was equal to or higher than 1:30.
Sequencing
Library preparation and NGS sequencing
Total RNA was extracted (Trizol LS; Life Technologies) from supernatants of infected BHK-21 cells. Samples were DNase and RNase treated prior to RNA extraction. The total RNA extract (20–70 ng) was treated with DNase (Turbo DNA-free kit; Ambion) then depleted of ribosomal RNA (rRNA; Low Input RiboMinus eukaryote system kit; Ambion). The DNA and rRNA-depleted RNA solution was then converted into representative complementary DNA (cDNA) libraries (Ion Total RNA-Seq kit; Life Technologies) according to the supplier’s instructions, except for 2 modifications: 1) RNA was fragmented with RNase III for 1 min at 37°C, and 2) the fragmented RNA was hybridized and ligated to adaptors overnight at 16°C. The resulting cDNA were amplified by 18 cycles of PCR (Ion Xpress RNA-Seq barcode primers; Life Technologies). The size distribution of the library was assessed for quality control (High Sensitivity DNA kit; Agilent). The library’s quality was validated prior to sequencing given that 30–60% of fragments were >160 bp.
The cDNA library sequencing started with an enrichment of 10 pM of the amplified library (Ion OneTouch 2 system; Life Technologies). Sequencing was performed (Ion Proton sequencer, Ion PI Chip v2; Life Technologies).
Bioinformatic analysis
Sample reads were cleaned with Trimmomatic software and aligned to the EMCV KM269482 ATCC VR-129B complete genome using TMAP (Torrent Suite v.4.0.2; https://github.com/iontorrent/TMAP) to evaluate coverage depth for each sample. 3 The MIRA assembler was run on uncleaned, down-sampled reads to a mean coverage depth of 80 after best k-mer size estimation by KmerGenie.7,8 The SPAdes assembler was used after Trimmomatic cleaning. 2
Contigs were aligned and reordered with Mauve. 32 Unrelated contigs were removed. Viral contigs not matching the reference were kept. Ambiguous nucleotides in the de novo assembly contigs were manually curated by visualizing read alignments in Tablet. 27
Sequence analysis and phylogenetic study
The assembled sequencing reads have been deposited in the NCBI database under BioProject accession PRJNA614001 and the consensus sequences under GenBank accessions MT250508, MT250509, MT250510, and MT250511. ORF, P1, VP1, and 3D sequences obtained were compared with EMCV sequences available in GenBank using the BLASTn online tool (https://blast.ncbi.nlm.nih.gov). Multiple sequence alignment and phylogenetic analyses based on the sequences obtained in our study and homologous EMCV sequences available in GenBank were conducted using MEGA 6 software.16,40 EMCV sequences were aligned using the Muscle program with default parameters. 12 Only one representative of redundant sequences and of strains showing 100% nucleotide sequence homology and isolated in the same country, from the same species, and in the same year was included in the analyses. Phylogenetic analyses were conducted by the neighbor-joining (NJ) method using the optimal substitution model giving the lowest Bayesian Information Criterion score in a model test implemented in MEGA 6.33,39 The confidence of the NJ tree was assessed by bootstrap analysis with 1,000 replicates.
Results
EMCV detection and isolation
The postmortem examination and histologic observations conducted after the death of elephant D oriented the diagnosis toward viral infection, and particularly a cardiovirus. 22 The EMCV genome was detected by RT-rtPCR in heart, kidney, and liver samples (Table 1). EMCV infection was further confirmed by isolation of the virus from the same organs and from blood (Table 1). The RT-rtPCR cycle threshold (Ct) value obtained from the heart sample (9.6) was lower than that obtained from the liver (21.3) and kidney (23.3) samples, indicating a higher quantity of viral RNA in the heart than in the other samples. This was consistent with the virus titer observed from the homogenate of samples in which it was much higher in the heart (108.69 TCID50/g) than in the liver (105.18 TCID50/g) or the kidney (104.52 TCID50/g; Table 1).
Results of virologic tests on tissue and blood samples collected from elephant D that died of encephalomyocarditis virus (EMCV) infection.
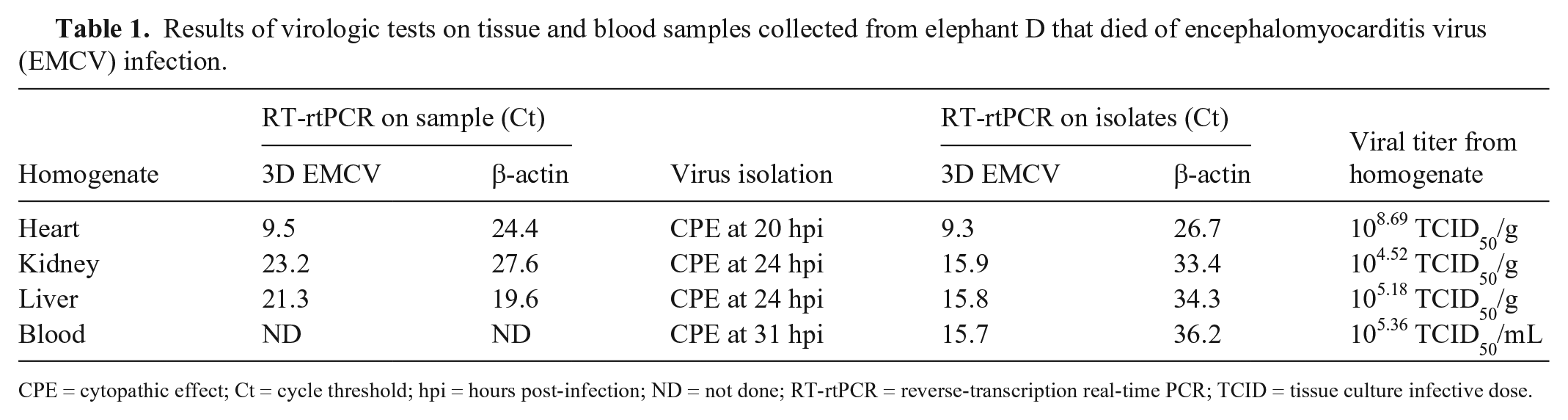
CPE = cytopathic effect; Ct = cycle threshold; hpi = hours post-infection; ND = not done; RT-rtPCR = reverse-transcription real-time PCR; TCID = tissue culture infective dose.
Origin of the EMCV infection
To trace the origin of the EMCV infection in elephant D, an investigation was conducted on sera collected from elephants present at Sigean and Erfurt zoos before and after the death of elephant D. EMCV antibodies were detected in sera collected from elephants A and B in June and February 2013, respectively, before the mutual transfer of animals in October 2013 (Table 2). The antibody titer was higher in the serum of elephant A (1:1,024) than that of elephant B (1:64). Sera collected in February 2013 from 3 elephants housed at Erfurt zoo were found negative. This includes elephants C and D that were later transferred to Sigean. Archived sera collected in September 2004 and February 2009 from 2 elephants housed at Sigean zoo were also found negative. After the exchange of animals, elephant A—the elephant transferred from Sigean to Erfurt zoo—showed signs of illness and was euthanized in December 2013. Serum collected from elephant A at this date was found positive for EMCV antibodies (Table 2). However, it is not known if the animal’s illness was the result of EMCV infection. The serum samples collected at Erfurt in April and November 2014 from elephant B, the second elephant transferred from Sigean, revealed a slight decrease in EMCV antibody titer (1:32 and 1:22) compared to the sample collected in February 2013 at Sigean. Both elephant E, an elephant that remained at Erfurt, and elephant C, an elephant from Erfurt that was transferred to Sigean, were negative for EMCV antibodies. Interestingly, the serum sample collected from elephant D on the day the elephant died was also negative (Table 2).
Results of serologic tests on samples collected from elephants at Sigean and Erfurt zoos.
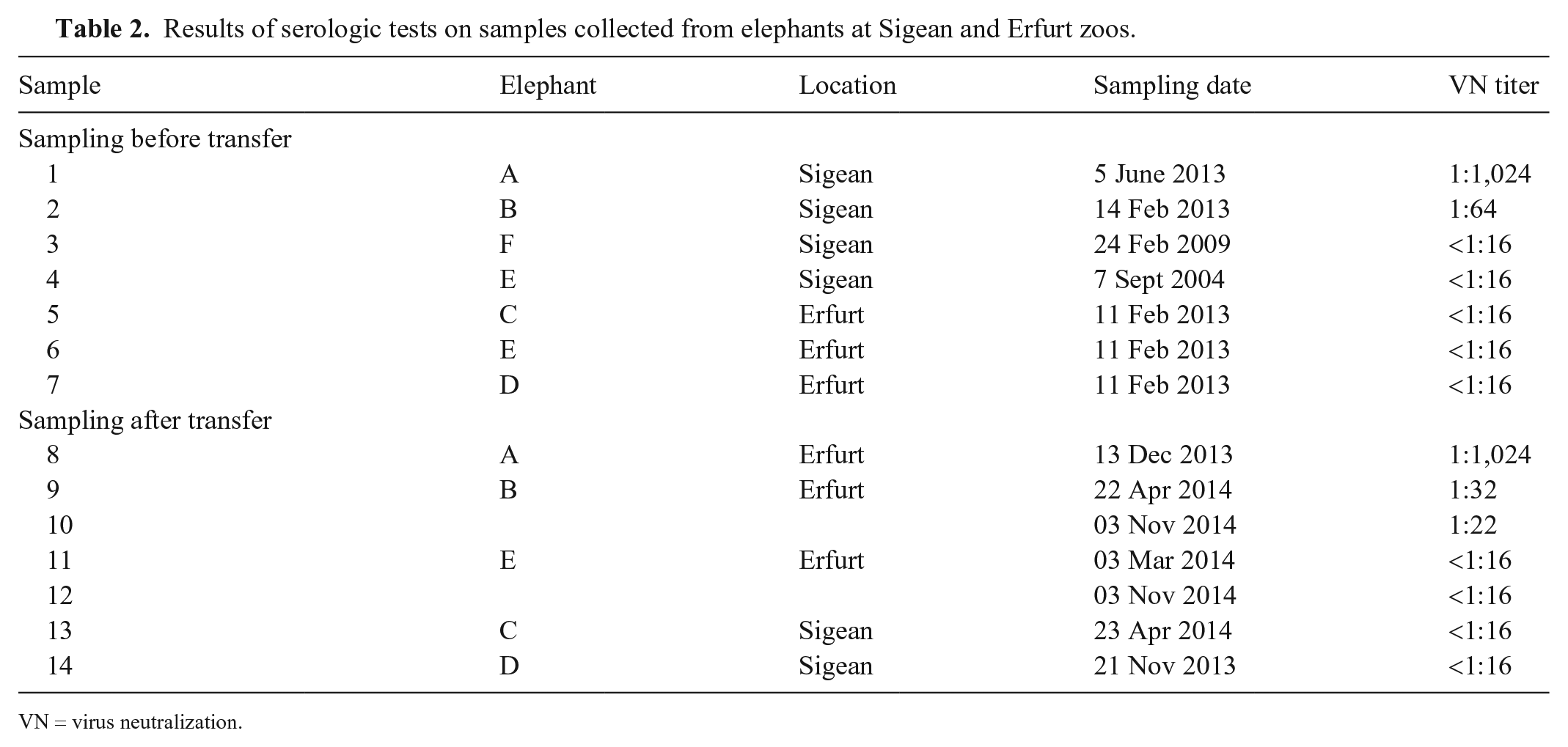
VN = virus neutralization.
Investigations conducted on sera collected between March 2014 and February 2016 from 99 animals representing 28 species revealed the presence of EMCV antibodies in 14 animals (data not shown). The positive animals included nonhuman primates, pigs, porcupines, sitatungas, and giraffes.
Given that rodents are considered a potential reservoir and disseminator of EMCV, pest control was intensified at Sigean zoo, and rodents were captured and tested for EMCV infection. The EMCV genome was detected by RT-rtPCR in samples collected from 9 of 21 tested rats (Table 3); more precisely, in the ileum of 2 rats (10 and 12), in the mesenteric lymph node of 4 rats (2, 3, 10, 14), and in the feces of 6 rats (9, 12, 14, 15, 16, 17). The RT-rtPCR Ct values ranged from 26.5 from the ileum to 36.8 from the mesenteric lymph node. Five of these rats (2, 3, 10, 11, 14) were caught in the elephant barn. EMCV infection of rats was confirmed by the isolation of the virus on BHK-21 cells from 3 RT-rtPCR–positive samples collected from 3 different rats (10, 16, 17). In addition, a high titer of EMCV antibodies (1:16,384) was observed in 1 (rat 9) of the 3 rats tested (9, 20, 21; data not shown). The EMCV genome was also detected by RT-rtPCR from the fecal sample of this rat.
Results of virologic tests on samples collected from rats captured in Sigean zoo.
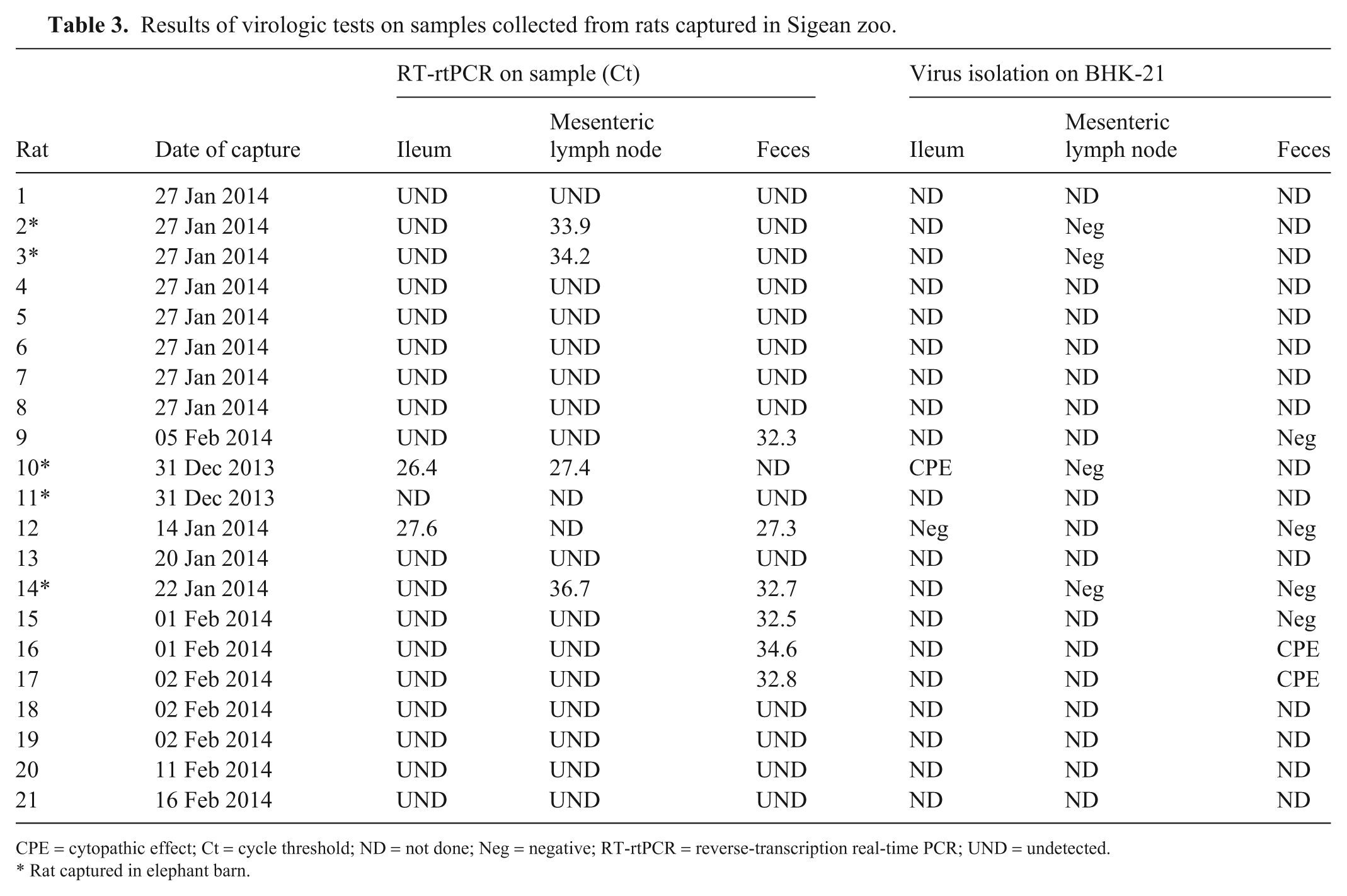
CPE = cytopathic effect; Ct = cycle threshold; ND = not done; Neg = negative; RT-rtPCR = reverse-transcription real-time PCR; UND = undetected.
Rat captured in elephant barn.
Genomic characterization
To investigate the genetic relationship of the Sigean EMCV strains with existing EMCV isolates, we sequenced the full genome of the circulating virus directly from the heart sample of the dead elephant (FRA/SIG/1/2013) as well as from the virus isolated from the ileum of rat 10 (FRA/SIG/1/2014), from the feces of rat 16 (FRA/SIG/2/2014), and from the feces of rat 17 (FRA/SIG/3/2014). By using next-generation sequencing and read assembly, we generated genomic sequences of 7,875 nucleotides. The viral genome contains a single ORF of 6,876 nt encoding a polyprotein of 2,292 aa. The ORF is flanked by a 5’-UTR (untranslated region) of 846 nt including the poly (C) (~145 nt), a 3’-UTR of 123 nt, and ends by a poly (A) tail of 27 nt. Multiple sequence alignments showed that the elephant isolate shared with the rat isolates a high nucleotide sequence identity of 98.1–99.9% in the full genome (Table 4). The highest sequence identity was observed with isolate SIG/1/2014 obtained from rat 10 that was caught in the elephant barn (99.9%). The nucleotide sequence identities between rat isolates were 98.1–98.9% (data not shown). In the entire ORF, the sequence identities between the elephant isolate and rat isolates were 98.0–99.9% at the nucleotide sequence level and 99.5–99.9% at the amino acid sequence level. Sequence alignment based on each coding region indicated that the 2B nucleotide sequence showed the highest variation among the encoded regions. The L, VP2, VP3, 3A, and 3B regions are the most conserved based on amino acid sequence comparison (Table 4).
Uncorrected pairwise distance values (p-distance; %) among the encephalomyocarditis virus (EMCV) strain isolated from the dead elephant (FRA/SIG/1/2013) and EMCV strains isolated from rats (FRA/SIG/1/2014 to FRA/SIG/3/2014).
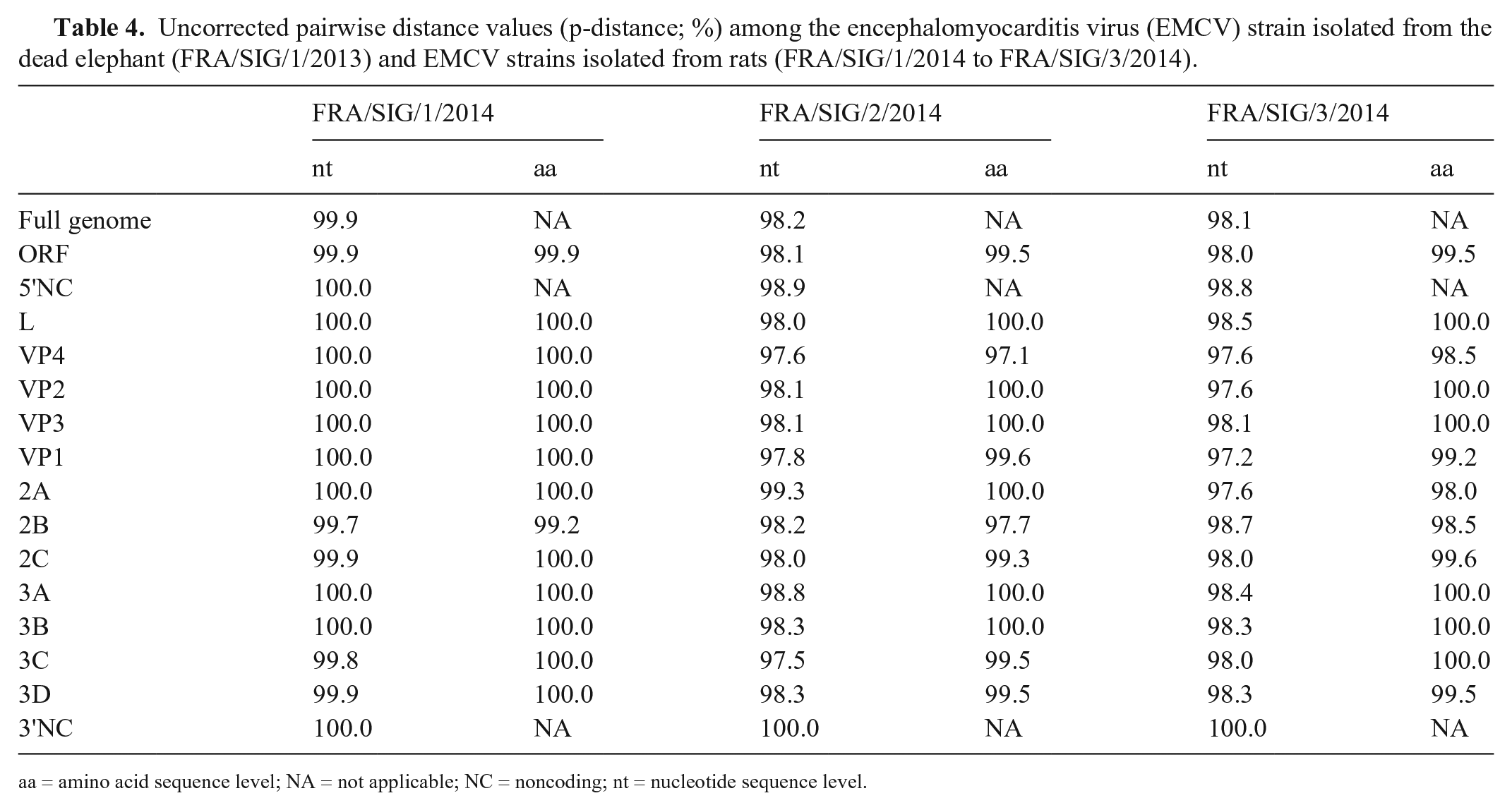
aa = amino acid sequence level; NA = not applicable; NC = noncoding; nt = nucleotide sequence level.
Phylogenetic analysis
To investigate the genetic relationship of the Sigean EMCV with available EMCV strains in GenBank, we performed multiple nucleotide and amino acid sequence comparisons for the ORF, P1, VP1, and 3D regions. The extent of nucleotide sequence identities between the Sigean strains and other EMCV strains was 65.3–84.8% in the ORF region, 58.7–84.0% in the P1 region, 48.9–84.1% in the VP1 region, and 74.3–94.4% in the 3D region. At the amino acid level, sequence identities were 79.7–96.5% in the ORF region, 72.9–99.2% in the P1 region, 59.6–99.6% in the VP1 region, and 89.6–100% in the 3D region. The highest nucleotide sequence identity was observed with the ATCC VR-129B strain in the ORF region, with the HB1 strain in the P1 and VP1 regions, and with the GRE-432/86 strain in the 3D region. The highest amino acid sequence identity was observed with the ATCC VR-129B strain in the ORF region, with the YM13 strain in the P1 region, with B440/95, 1086C, YM13, and EMCV-30 strains in the VP1 region, and with SAR/1/1979, EMCV-30, and strains isolated in Greece in 1986–1997 in the 3D region (Suppl. Tables 1, 2).
Based on genetic analysis, EMCV isolates were divided into 4 virus groups (EMCV 1–4). The strains belonging to EMCV group 1 were separated into 7 lineages (A–G). 39 The phylogenetic analysis and topology of the trees based on the ORF, P1, VP1, and 3D nucleotide sequences showed that the Sigean EMCV strains belonged to lineage A of EMCV group 1 and formed a cluster (Fig. 1, Suppl. Figs. 1–3). High levels of support were obtained for this grouping with the 4 regions analyzed. The nucleotide sequence similarity of the Sigean EMCV with strains of lineage A was 77.4–84.1% in the VP1 region, 79.1–84.0% in the P1 region, 80.0–84.8% in the complete ORF, and 89.7–94.4% in the 3D region. At the amino acid sequence level, the similarities were 96.9–99.6%, 97.9–99.2%, 95.8–96.5%, and 96.2–100%, respectively (Suppl. Tables 1, 2).
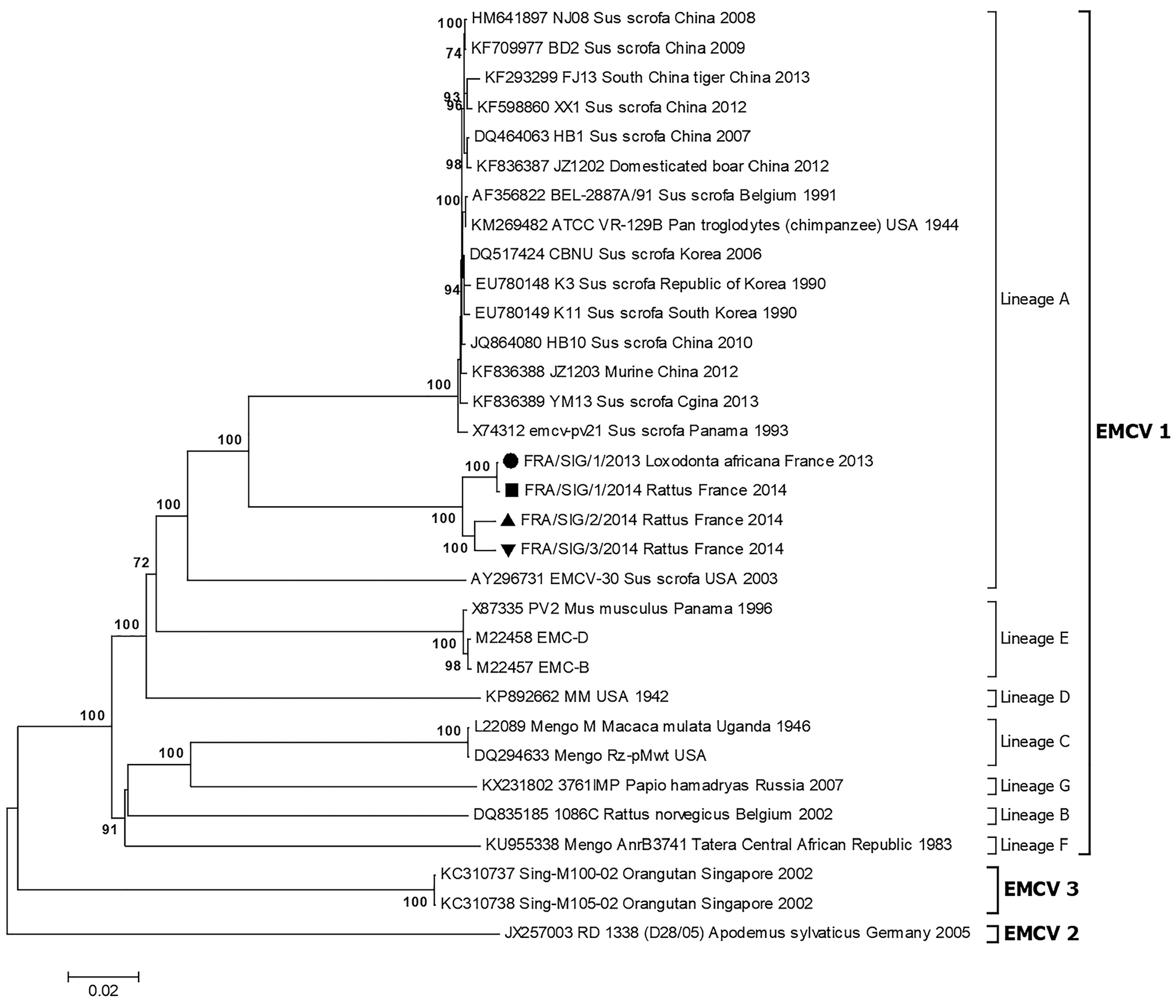

Neighbor-joining tree based on the complete ORF sequence showing relationships between the encephalomyocarditis virus (EMCV) strains isolated at Sigean zoo and EMCV sequences available in GenBank. The evolutionary distances were computed using the Tamura–Nei method. The percentages of bootstrap values >70% of 1,000 replicates that support each branch node are shown next to the branches. The scale bar indicates nucleotide substitutions per site.
Discussion
We observed a higher titer of EMCV in the heart than in other organs, a finding in agreement with the histologic lesions observed in the heart. 22 EMCV has a myocardial tropism in infected elephants that can cause their death from myocarditis.14,18,35,37
Serologic investigations revealed that elephant D and 2 other elephants at Erfurt zoo were negative for EMCV antibodies before the exchange between zoos (Table 2). However, 2 of 4 elephants at the Sigean zoo tested positive for EMCV antibodies before the exchange. The overall serologic results clearly indicated that EMCV was circulating at Sigean zoo before the exchange of elephants and strongly suggested that elephant D was infected at Sigean zoo after the transfer from Erfurt. The fact that elephant D was found negative for EMCV antibodies on the day of death suggests a recent and acute EMCV infection. Serologic investigations conducted on sera collected in Sigean zoo revealed the presence of EMCV antibodies in 14 animals including nonhuman primates, pigs, porcupines, sitatungas and giraffes. Unfortunately, to avoid unnecessary stress to animals, no blood samples were collected from more species to verify their seropositivity. Therefore, the extent of EMCV infection in the zoo is not known. Given that no clinical cases of EMCV infection were reported in the seropositive species, the serologic data suggest subclinical circulation of the virus in the zoo.
Outbreaks of EMCV worldwide have usually been associated with an increase in the rodent population.4,11,14,23,34 Experimental infections have shown that EMCV can be isolated from the feces of infected rats 29 d post-challenge, and that it persists in the rat population by rat-to-rat transmission. 38 Furthermore, the carcasses of infected rodents can be infectious through ingestion by susceptible animals. 28 These observations have led to rodents being considered the natural host and reservoir of EMCV, and therefore an important risk factor for the occurrence of the disease. 6 Our results strongly support these observations. Experimental EMCV infections have shown that elephants can be infected by the oral route and die from acute myocarditis. 18 In addition, it was shown that the virus could not be isolated from the feces of infected animals and no viral transmission occurred between infected elephants and those in contact. Given these observations, our results, and the fact that elephant D died during a cold month when rodents are looking for food and warmth in animal barns, it makes sense to infer that the elephant could have been infected through ingestion of food contaminated with feces from EMCV-infected rats. These findings support the need for an effective rodent control program at zoo level.
The geographic origin of the virus and the timing of its introduction into the zoo remain unclear. The virus may have been introduced into the zoo by infected rats from areas surrounding the zoo or by another infected species residing in the zoo. Serologic investigations conducted in France in 1998–2001 showed an overall EMCV seroprevalence of 2.48–3.27% in domestic pigs and 3.55% in wild boars, suggesting that the virus was circulating in the country.1,26 Our phylogenetic analysis indicated that the Sigean EMCV belongs to lineage A of EMCV group 1. This lineage has a broad geographic distribution, and most of the strains have been associated with myocarditis among pigs. It comprises isolates from China, South Korea, the United States, Panama, South Africa, Belgium, and Greece. Compared to EMCV strains reported previously in Europe, the highest nucleotide sequence homology of the Sigean strain was observed with strain BEL-2887A/91 in the ORF, P1, and VP1 regions, and with strain GRE-432/86 in the 3D region. These 2 strains were isolated in Belgium in 1991 and in Greece in 1986, respectively. The highest homology was observed with the strain GRE-432/86 in the 3D region (94.4%). Interestingly, the Sigean strain showed a lower sequence homology in the 3D region (84.2%) with the EMCV strain FRA-919/95 reported in France in 1995. Full genome sequencing and sequence comparisons suggest that the Sigean EMCV strain diverges significantly from known strains, even from strains previously isolated in Europe and France. Investigations did not identify the origin of this virus. The availability of more sequence data—in particular from EMCV strains possibly circulating in France and neighboring countries—could provide insights into the probable origin of the Sigean EMCV strains. We recommend systematic laboratory investigation of fatal cases of myocarditis among zoo animals and pigs in France to implement control measures in the event of an EMCV outbreak.
Supplemental Material
sj-pdf-1-vdi-10.1177_1040638720978389 – Supplemental material for Molecular characterization of encephalomyocarditis virus strains isolated from an African elephant and rats in a French zoo
Supplemental material, sj-pdf-1-vdi-10.1177_1040638720978389 for Molecular characterization of encephalomyocarditis virus strains isolated from an African elephant and rats in a French zoo by Aurore Romey, Benjamin Lamglait, Yannick Blanchard, Fabrice Touzain, Helene Quenault, Anthony Relmy, Stephan Zientara, Sandra Blaise-Boisseau and Labib Bakkali-Kassimi in Journal of Veterinary Diagnostic Investigation
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Supplementary material
Supplementary material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
