Abstract
Two hundred and twenty-six attaching and effacing Escherichia coli (AEEC) strains (20 enterohemorrhagic E. coli and 206 atypical enteropathogenic E. coli) isolated from calves, lambs, and goat kids with diarrhea and from healthy cattle, sheep, and goats were tested for their resistance to 10 antimicrobial agents by the disc diffusion method. Resistant and intermediate strains were analyzed by polymerase chain reaction for the presence of the major resistance genes. The overall percentage of resistant strains to tetracycline, streptomycin, erythromycin, and sulfamethoxazole was very high (>65%). Moreover, a high level of resistance (approximately 30%) to ampicillin, chloramphenicol, trimethoprim, and trimethoprim-sulfamethoxazole was also detected. The AEEC strains were very susceptible (>90%) to gentamicin and colistin. Because AEEC from ruminants can cause diseases in human beings, the high frequency of antimicrobial resistance detected in the current study is a source of concern. For each antimicrobial agent, the predominant resistance genes in the resistant strains were ampicillin, bla TEM (97.1%); tetracycline, tetA (76.7%); gentamicin, aac(3)II (80%); streptomycin, strA/strB (76.7%) and aadA (71.7%); chloramphenicol, catI (85.1%); trimethoprim, dhfrI (76.3%); and sulfamethoxazole, sul1 (60%) and sul2 (63.3%). In the majority of cases, resistance to a given antimicrobial, except for streptomycin, was caused by a single gene. A negative association between tetA and tetB, between aac(3)II and aac(3)IV, and between dhfrI and dhfrV was observed. The present study gives baseline data on frequency and molecular basis of antimicrobial resistance in AEEC strains from ruminants.
Attaching and effacing Escherichia coli (AEEC) are a cause of diarrhea in humans and animals. These bacteria cause a characteristic attaching and effacing (AE) lesion in the gut mucosa because of the intimate bacterial adhesion to the enterocytes and effacement of the brush border microvilli. Human enteropathogenic E. coli (EPEC) and human enterohemorrhagic E. coli (EHEC) cause AE lesions in the intestinal mucosa and possess the eae gene. Enterohemorrhagic E. coli strains produce verotoxins (VTs) and cause a broad spectrum of enteric disease from mild gastroenteritis to hemorrhagic colitis, hemolytic uremic syndrome, and even death. 15 Domestic ruminants, mainly cattle, have been implicated as the principal reservoir of these strains for humans. 15 In contrast to EHEC, EPEC strains do not produce VTs. 15 Enteropathogenic E. coli have been classified as typical (possessing the bfpA gene) or atypical (lacking the bfpA gene). Typical EPEC are a major cause of infantile diarrhea in developing countries and are rarely found in animals. Atypical EPEC are an important cause of diarrhea in developing and developed countries and can be isolated from human beings and animals. 1
As a result of its epidemiologic importance, the frequency of antimicrobial resistance in human and animal VT-producing E. coli (VTEC), including EHEC, has been the subject of many studies. 12,14 In contrast, only a limited number of studies of the frequency of antimicrobial resistance have focused on atypical EPEC from human beings 1 and, to the authors' knowledge, none on atypical EPEC from ruminants. Chemotherapy for EHEC infections remains controversial. 12,14 Some studies suggest restriction of the use of antimicrobials for therapy of EHEC infections because antimicrobials may lyse bacterial cell walls, thereby liberating VTs, and/or cause increased expression of VT genes in vivo. 20 On the other hand, other studies suggest that some antimicrobials, if administered early in the course of infection, may prevent disease progression to hemolytic uremic syndrome. 11 In addition, the administration of antimicrobial therapy may be useful for cases of persistent diarrhea associated with atypical EPEC. 1 Assessment of the diversity and distribution of resistance genes in bacterial populations represents a more detailed and potentially useful additional tool for improving diagnosticians' understanding of antimicrobial resistance epidemiology. 2 However, there is little information available on the mechanisms of antimicrobial resistance identified among EHEC strains and, to the authors' knowledge, no data from EPEC strains.
The first objective of the current study was to determine the frequency of resistance to several antimicrobial agents, including those commonly used to treat diarrhea in human beings, in AEEC strains from ruminants. The second objective was to assess the diversity and distribution of the major resistance genes in these strains.
Number (percentage) of susceptible (S), intermediate (I), and resistant (R) strains of attaching and effacing Escherichia coli isolated from ruminants to 10 antimicrobial agents. *
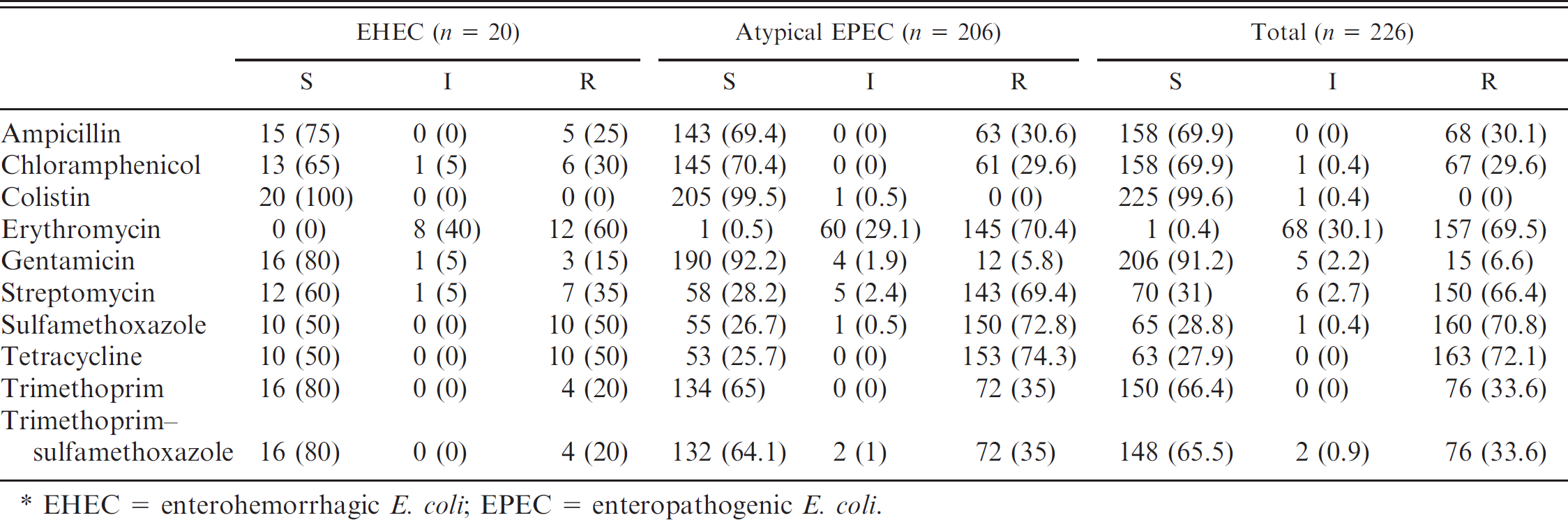
EHEC = enterohemorrhagic E. coli; EPEC = enteropathogenic E. coli.
A total of 226 AEEC strains isolated from calves (24), lambs (17), and goat kids (7) with diarrhea and from healthy cattle (39), sheep (62), and goats (77) in Spain between 1993 and 2005 were used in this study. Twenty of these strains were EHEC (eae +, VT+), and 206 were atypical EPEC (eae +, VT−, bfpA −) and belonged to 74 different serotypes. Only 1 strain with different characteristics (serotype and production of VTs) per animal was included in the study. The 226 selected strains were isolated from 210 animals (48 from diarrheic animals and 162 from healthy animals) in 73 herds and have been described previously. 4,7,16,17
Antimicrobial testing was performed using the disc diffusion method, according to the recommendations of the Clinical and Laboratory Standards Institute. 5 Measurement of growth inhibition areas allowed the classification of each strain as susceptible, intermediate, or resistant, according to data provided by the Clinical and Laboratory Standards Institute. 5 The following antimicrobials belonging to 8 different classes were tested: ampicillin (β-lactams), tetracycline (tetracyclines), gentamicin and streptomycin (aminoglycosides), colistin (polymyxins), erythromycin (macrolides), chloramphenicol (phenicols), trimethoprim (trimethoprim), sulfamethoxazole (sulfonamides), and trimethoprim–sulfamethoxazole. All antimicrobial susceptibility discs were provided from a commercial source. a
The presence of the major resistance genes for ampicillin (bla TEM, bla OXA-1, bla SHV, and bla CMY), tetracycline (tetA, tetB, tetC, and tetM), gentamicin [aac(3)II, aac(3)IV, and ant(2″)I], streptomycin (strA, strB, and aadA), chloramphenicol (catI, floR, and cmlA), trimethoprim (dhfrI and dhfrV), and sulfamethoxazole (sul1, sul2, and sul3) was determined in resistant and intermediate resistant strains by polymerase chain reaction as previously described. 2,3,6,18,19 Significant differences in the frequencies of resistance were determined by the Fisher exact test. A P value of less than 0.05 was considered significant. Analyses were performed using statistical software. b
The antimicrobial susceptibility of the 226 AEEC strains is summarized in Table 1. The overall percentage of resistant strains to tetracycline, streptomycin, erythromycin, and sulfamethoxazole was very high (>65%). Moreover, a high level of resistance (approximately 30%) to ampicillin, chloramphenicol, trimethoprim, and trimethoprim–sulfamethoxazole was also detected. The AEEC strains were very susceptible (>90%) to gentamicin and colistin. Thirty-five strains (15.5%) were resistant to 1 antimicrobial. Most of the strains showed multidrug resistance: 74.8% were resistant to at least 2 antimicrobials, 59.7% were resistant to at least 4 antimicrobials, and 34.9% were resistant to at least 6 antimicrobials. Approximately 14% of the strains were resistant to at least 8 antimicrobials. The lowest susceptibility rate was obtained for erythromycin, as expected, because that antimicrobial is considered to have moderate action on the members of the family Enterobacteriaceae. In contrast, none of the AEEC strains was resistant to colistin (polymyxin E). This antimicrobial has been found previously to be highly effective against E. coli from humans and animals. 9,10 As the use of chloramphenicol in animal husbandry is banned in Europe, the resistance to this antimicrobial found in the current study is probably due to coselection of chloramphenicol resistance genes with other resistance genes.
The percentages of atypical EPEC and EHEC resistant to the antimicrobials used in the present study were, in general, higher than those previously reported for atypical EPEC isolated from children with diarrhea in Brazil 1 and for VTEC (including some EHEC) from human beings, cattle, sheep, and food in Spain. 14 In addition, in a study carried out on E. coli O26 and O111 isolates from cattle, most of which were EHEC, 12 the resistance to gentamicin was lower, and resistance to ampicillin, tetracycline, streptomycin, and chloramphenicol was higher than resistance to those antimicrobials in EHEC in the present study. Because AEEC from ruminants can cause diseases in humans, the high frequency of antimicrobial resistance detected in the current study is a source of concern.
When the frequency of resistance among the AEEC strains was compared, several significant differences in resistance to some of the antimicrobials tested were found: 1) atypical EPEC were more resistant to tetracycline (P < 0.05), streptomycin (P < 0.005), and sulfamethoxazole (P < 0.05) than EHEC; 2) AEEC from animals with diarrhea were more resistant to ampicillin (P < 0.0001), streptomycin (P < 0.05), and chloramphenicol (P < 0.005) than AEEC from healthy animals; 3) AEEC from young animals were more resistant to tetracycline (P < 0.0005), streptomycin (P < 0.0005), chloramphenicol (P < 0.01), and sulfamethoxazole (P < 0.0001) than AEEC from adult animals; 4) caprine AEEC were more resistant to ampicillin (P < 0.005), gentamicin (P < 0.005), streptomycin (P < 0.05), chloramphenicol (P < 0.05), trimethoprim (P < 0.0001), and trimethoprim–sulfamethoxazole (P < 0.0001) than ovine AEEC; 5) caprine AEEC were more resistant to tetracycline (P < 0.0001), streptomycin (P < 0.001), trimethoprim (P < 0.05), sulfamethoxazole (P < 0.0001), and trimethoprim–sulfamethoxazole (P < 0.05) than bovine AEEC; 6) bovine AEEC were more resistant to ampicillin (P < 0.005), gentamicin (P < 0.005), chloramphenicol (P < 0.05), trimethoprim (P < 0.005), and trimethoprim–sulfamethoxazole (P < 0.01) than ovine AEEC; and 7) ovine AEEC were more resistant to trimethoprim (P < 0.0001) and sulfamethoxazole (P < 0.001) than bovine AEEC. As expected, AEEC strains from animals with diarrhea were more resistant than those from healthy animals for some of the antimicrobial agents examined. In addition, AEEC strains from young animals were more resistant to tetracycline, streptomycin, chloramphenicol, and sulfamethoxazole than those from adult animals. Previous studies have shown that the frequency of antimicrobial resistance is usually higher in younger animals. 2 Curiously, caprine AEEC strains showed higher levels of resistance than bovine and ovine AEEC strains against several antimicrobials.
The antimicrobial resistance genes were studied in all resistant strains to ampicillin, gentamicin, chloramphenicol, and trimethoprim, in 60 resistant strains to tetracycline, streptomycin, and sulfamethoxazole (approximately 40% of the all-resistant strains to these antimicrobials), and in all intermediate resistant strains to gentamicin, streptomycin, chloramphenicol, and sulfamethoxazole. For each antimicrobial agent, the predominant resistance genes in the resistant strains were ampicillin, bla TEM (97.1%); tetracycline, tetA (76.7%); gentamicin, aac(3)II (80%); streptomycin, strA/strB (76.7%) and aadA (71.7%); chloramphenicol, catI (85.1%); trimethoprim, dhfrI (76.3%); and sulfamethoxazole, sul1 (60%) and sul2 (63.3%; Table 2). In the majority of cases, resistance to a given antimicrobial, except for streptomycin, was caused by a single gene. The simultaneous presence of more than 1 gene conferring resistance to the same antimicrobial agent was observed in 58 cases (Table 2). Furthermore, in 14 cases, the resistance genes could not be identified (Table 2).
Almost all AEEC strains resistant to ampicillin possessed the bla TEM gene, which is in agreement with previous studies on human and bovine E. coli that are resistant to β-lactams. 6,8,9 Most of the AEEC strains from ruminants resistant to tetracycline possessed the tetA gene. However, in contrast with the data from the present study, the frequency in another study 9 of tetA in bovine E. coli isolates resistant to tetracycline was very similar to that of tetB. Also in contrast with the results of the current study, other studies 3,8 have found that most human and ruminant E. coli strains resistant to tetracycline carry the tetB gene.
As in human Enterobacteriaceae strains, 13 the aac(3)II gene was predominant among the ruminant AEEC strains resistant to gentamicin. However, in another study, 19 the ant(2″)I gene was very frequent in bovine E. coli strains resistant to gentamicin, whereas the aac(3)II gene was present at a low level in these strains. Although the aac(3)II gene was predominant among the AEEC strains resistant to gentamicin, all strains with intermediate resistance to gentamicin had the aac(3)IV gene, which suggests that the aac(3)II gene confers higher levels of gentamicin resistance than the aac(3)IV gene. The present study is believed to be the first to suggest that the aac(3)II gene encodes a higher level of resistance than the aac(3)IV gene. This may be because most previous studies have examined gentamicin resistance genes only in resistant E. coli strains. However, it is also possible that the hypothesis that aac(3)II confers higher levels of gentamicin resistance than aac(3)IV was an artifact of the isolate selection.
In agreement with a German study carried out on E. coli strains from cattle, swine, and poultry, 9 the strA/strB and aadA genes were very frequent in the ruminant AEEC strains resistant to streptomycin. However, the combination of strA/strB and aadA occurred much more frequently in the ruminant AEEC strains of the present study (48.3%) than in the animal E. coli strains of the former study (17.9%). 9 In addition, in a recent study, 8 the percentage of bovine E. coli strains resistant to streptomycin that carried the aadA gene was much lower (36%) than that found in animal E. coli strains in Germany (61%) 9 and in ruminant AEEC strains in the present study (71.7%).
In agreement with previous studies on bovine E. coli, 8,9 the majority of the ruminant AEEC strains resistant to chloramphenicol and trimethoprim carried the catI and dhfrI genes, respectively. The present work is also consistent with earlier reports 8,9 that sulphonamide resistance in bovine E. coli strains is mostly mediated by the sul1 and sul2 genes. However, the results of the current study are not totally in agreement with those obtained in the former studies. For example, in the present study, the frequency of sul1 was very similar to that of sul2, whereas in previous studies, 8,9 the sul2 gene was detected much more often (about twice and 5 times) than the sul1 gene in bovine E. coli strains resistant to sulfamethoxazole. In agreement with a previous work, 9 only a low percentage of the ruminant AEEC strains resistant to sulfamethoxazole carried the sul3 gene.
Altogether, the results of the present study showed a negative association between tetA and tetB, between aac(3)II and aac(3)IV, and between dhfrI and dhfrV. The negative association between the tetA and tetB genes has been observed previously and probably results from plasmid incompatibilities. 2 However, the present study is the first, to the authors' knowledge, that describes the negative associations between the aac(3)II and aac(3)IV genes and between the dhfrI and dhfrV genes. The new information described in the present study on antimicrobial resistance in EHEC and atypical EPEC strains from ruminants will contribute to risk assessment of these pathogens.
Number (percentage) of antimicrobial resistance genes in resistant and intermediate attaching and effacing Escherichia coli strains isolated from ruminants.
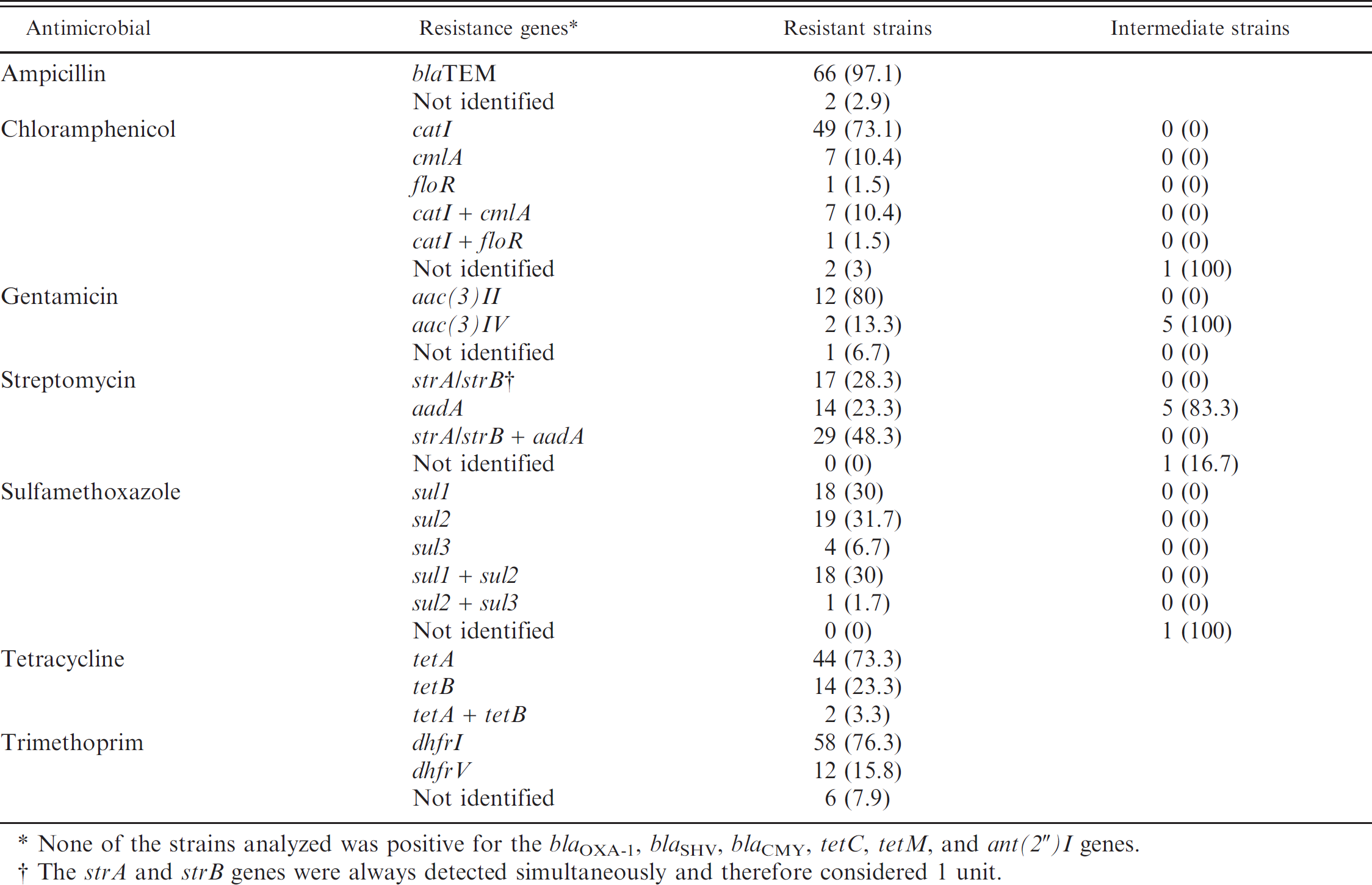
None of the strains analyzed was positive for the bla OXA-1, bla SHV, bla CMY, tetC, tetM, and ant(2″)I genes.
The strA and strB genes were always detected simultaneously and therefore considered 1 unit.
Acknowledgements. Alberto Medina and Pilar Horcajo contributed equally to this work. This study was supported by grants from the Dirección General de Investigación (grant AGL2004-08139) and the Banco Santander Central Hispano-Universidad Complutense (INBAVET 920338). The authors thank L. Fernández for technical assistance and M. A. Moreno, K. Colom, I. Badiola, C. Torres, and F. M. Aarestrup for generously providing the control strains.
Footnotes
a.
Rosco Diagnostica A/S, Taastrup, Denmark.
b.
InStat (version 3), GraphPad Inc., San Diego, CA.
