Abstract
The spread of antimicrobial-resistant bacteria is a significant concern, as it can lead to increased morbidity and mortality in both humans and animals. Whole-genome sequencing (WGS) is a powerful tool that can be used to conduct a comprehensive analysis of the genetic basis of antimicrobial resistance (AMR). We compared the phenotypic and genotypic AMR profiles of 97 Salmonella isolates derived from chicken and turkey diagnostic samples. We focused AMR analysis on 5 antimicrobial classes: aminoglycoside, beta-lactam, phenicol, tetracycline, and trimethoprim. The overall sensitivity and specificity of WGS in predicting phenotypic antimicrobial resistance in the Salmonella isolates were 93.4% and 99.8%, respectively. There were 16 disagreement instances, including 15 that were phenotypically resistant but genotypically susceptible; the other instance involved phenotypic susceptibility but genotypic resistance. Of the isolates examined, 67 of 97 (69%) carried at least 1 resistance gene, with 1 isolate carrying as many as 12 resistance genes. Of the 31 AMR genes analyzed, 16 were identified as aminoglycoside-resistance genes, followed by 4 beta-lactam–resistance, 3 tetracycline-resistance, 2 sulfonamide-resistance, and 1 each of fosfomycin-, quinolone-, phenicol-, trimethoprim-, bleomycin-, and colistin-resistance genes. Most of the resistance genes found were located on plasmids.
In the United States, Salmonella bacteria cause ~1.35 million human infections, 26,500 hospitalizations, and 420 deaths per year. 5 According to Food Safety and Inspection Service data, ~23.0% of foodborne salmonellosis in humans results from the consumption of contaminated chicken and turkey. 38 Despite the 50.0% decrease in Salmonella-positive chicken samples from 2017 to 2021, there has been no significant reduction in human Salmonella infections. 39 In addition, Salmonella bacteria have shown resistance to various antimicrobials belonging to the following classes or subclasses: aminoglycosides, beta-lactams, phenicols, and tetracyclines. The continued emergence and spread of antimicrobial-resistant Salmonella bacteria create a serious public health challenge that calls for better understanding and effective control of antimicrobial resistance (AMR) in these bacteria. 11
It is known that antimicrobial susceptibility testing (AST) of Salmonella bacteria isolated from poultry samples is critical to the judicious use of antimicrobials for treating infections, tracking AMR trends, and formulating public health policies related to foodborne salmonellosis. AST is traditionally carried out using broth dilution and/or disk diffusion methods, which detect phenotypic resistance by measuring bacterial growth in the presence of the antimicrobials being tested. 2 Over the last several decades, commercial semi-automated and automated systems, such as Sensititre (Thermo Fisher), Biomic (Giles Scientific), and Vitek (bioMérieux), have been widely used in clinical or diagnostic laboratories. 7
PCR-based methods have been developed to detect known AMR genes (AMRGs). However, detecting individual AMRGs by PCR does not provide information about the effect of genomic context on resistance mechanisms. 28 It is also laborious to screen a large number of AMRGs in a particular bacterial isolate or specific AMRGs in a large number of isolates by PCR. High-throughput sequencing technologies are increasingly replacing traditional methods across a wide range of applications. For instance, whole-genome sequencing (WGS) could be a one-stop shop approach that accomplishes several objectives in a single test, including taxonomic identification, subtyping, phylogenetic analysis, determination of virulence factors, and detection of AMRGs. 28 WGS provides comprehensive information on potential resistance, ena-bling the identification of all AMRGs with greater accuracy compared to phenotypic AST using predefined antimicrobial panels and concentration ranges or PCR targeting specific AMRGs. 10
WGS is uniquely suitable for uncovering new resistance mechanisms and assessing the impact of genomic content on resistance traits. For example, a previously unknown gentamicin-resistance gene, grdA, was discovered by comparing the genomes of Salmonella isolates that were phenotypically resistant but genotypically susceptible to gentamicin 17 ; grdA in 1 isolate was initially found on a plasmid and flanked by 2 transposase genes, indicating that it could be integrated into the chromosome. Indeed, it was subsequently confirmed that most isolates harbored the grdA gene in their chromosomes. 17 Several studies have compared AMRGs detected by WGS with resistant phenotypes determined in different bacteria, such as Escherichia coli, 34 Salmonella enterica, 21 Campylobacter spp., 42 and Staphylococcus pseudintermedius. 37 These studies have demonstrated a strong association between genotypic resistance and phenotypic resistance, suggesting that WGS could be a viable option for AST and AMR monitoring.
To date, many WGS-based AMR studies have focused on bacteria isolated from human samples and retail meats. 27 With the goal of integrating WGS into diagnostic veterinary bacteriology workflow, we analyzed a collection of Salmonella bacteria isolated from poultry diagnostic samples.
Materials and methods
Bacterial isolates
We obtained 97 archived S. enterica isolates (20 from chickens, 77 from turkeys) from the University of Missouri Veterinary Medical Diagnostic Laboratory (VMDL) Bacteriology Section (Columbia, MO, USA). The bacteria were isolated from diagnostic samples from 2010 to 2017. Species identification was determined using matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS; Bruker), and serotyping of the isolates was conducted in the National Veterinary Services Laboratories (NVSL; Ames, IA, USA).
Antimicrobial susceptibility testing and breakpoints
The antimicrobial susceptibility of Salmonella isolates was initially determined using the Sensititre AVIAN1F plate (ThermoFisher) according to the manufacturer’s instructions. Minimum inhibitory concentration (MIC) and quality control testing were conducted according to Clinical Laboratory Standards Institute (CLSI) guidelines for broth microdilution methods. 6 The AVIAN1F antibiotic concentrations were as follows: amoxicillin (0.25–16 mg/L), ceftiofur (0.25–4 mg/L), clindamycin (0.5–4 mg/L), enrofloxacin (0.12–2 mg/L), erythromycin (0.12–4 mg/L), florfenicol (1–8 mg/L), gentamicin (0.5–8 mg/L), neomycin (2–32 mg/L), novobiocin (0.5–4 mg/L), oxytetracycline (0.25–8 mg/L), penicillin (0.06–8 mg/L), spectinomycin (8–64 mg/L), sulfadimethoxine (32–256 mg/L), sulfathiazole (32–256 mg/L), streptomycin (8–1024 mg/L), tetracycline (0.25–8 mg/L), trimethoprim/sulfamethoxazole (0.5/9.5–2/38 mg/L), and tylosin tartrate (2.5–20 mg/L).
Due to a lack of established breakpoints for antimicrobials against poultry Salmonella isolates, CLSI clinical breakpoints for human Enterobacterales 6 and the National Antimicrobial Resistance Monitoring System (NARMS) clinical breakpoints for human Salmonella 4 were adopted when they were available. The breakpoints to denote resistance were as follows: ceftiofur, 8 mg/L (NARMS); gentamicin, 16 mg/L (CLSI, NARMS); oxytetracycline, 16 mg/L (CLSI); streptomycin, 32 mg/L (NARMS); tetracycline, 16 mg/L (CLSI, NARMS); trimethoprim/sulfamethoxazole, 4/76 mg/L (CLSI, NARMS). The breakpoint for florfenicol (16 mg/L) was based on CLSI guidelines on swine respiratory Enterobacterales. 6 Because there is no CLSI and NARMS clinical breakpoint for amoxicillin alone, we adopted the European Committee on Antimicrobial Susceptibility Testing (EUCAST) clinical breakpoint (8 mg/L) for human Enterobacterales. 32
A rounding principle was applied when a raw MIC read given by the Sensititre system was higher than the highest concentration of an antibiotic in the AVIAN1F plate. For example, when an isolate’s gentamicin AST read was >8 mg/L, the MIC was rounded up to ≥16 mg/L because the actual MIC could be higher than 16 mg/L.
Enrofloxacin, clindamycin, erythromycin, penicillin, novobiocin, and tylosin tartrate are not intended for use in poultry. There are no published clinical breakpoints for spectinomycin, neomycin, sulfadimethoxine, and sulfathiazole. Therefore, all of these antimicrobials were excluded from phenotypic analysis.
Whole-genome sequencing and bioinformatics
Total DNA of individual isolates was purified (DNeasy blood & tissue kit; Qiagen) according to the manufacturer’s instructions. DNA concentrations were measured (Qubit fluorometer, dsDNA HS assay kit; Thermo Fisher) per the manufacturer’s instructions. Sequencing libraries were prepared (DNA Prep kit; Illumina) following the manufacturer’s protocol. Sequencing was conducted (iSeq reagent kit v2 [2 × 150 bp, iSeq 100 platform] or MiSeq reagent kit v2 [2 × 250 bp, MiSeq platform]; Illumina). Bioinformatic analysis was conducted using the GalaxyTrakr server (https://galaxytrakr.org/). 14 Briefly, sequencing reads were uploaded and subjected to the NARMS Salmonella AMR Workflow for quality trimming, de novo assembling, se-rovar identification, and AMR gene identification. Whole genomes of the 97 Salmonella isolates were deposited into GenBank under BioProject accession PRJNA993710 with BioSample accessions SAMN36408382–SAMN36408478. Library preparation and sequencing were repeated when sequencing data from a particular run failed to meet the quality of ≥ 30 average read quality Q score, ≥ 30-fold coverage, or ≤ 300 contigs. 33 To detect AMRGs not included in the NARMS AMRFinder_2020_09_30.1 database, such as grdA, the assembled contigs were imported into CLC Genomics Workbench 22 (Qiagen) and run against the ResFinder database. The assembled contigs were also subject-ed to analysis using GalaxyTrakr MOB-Recon to detect plasmids. 29
Correlation of genotypic and phenotypic resistance
An isolate was defined as genotypically resistant to an antimicrobial agent when one or more AMRGs were identified, and phenotypically resistant to an antimicrobial agent when the MIC was equal to or greater than the breakpoint. The AMR genotype and phenotype of each isolate were compared for the following antimicrobials: gentamicin, streptomycin, amoxicillin, ceftiofur, florfenicol, oxytetracycline, tetracycline, and trimethoprim/sulfamethoxazole. When a discrepancy between genotype and phenotype occurred, the phenotypic AMR was verified using E-Test (bioMérieux) according to the manufacturer’s instructions. The sensitivity of WGS for the detection of AMR was defined as the number of isolates that were both genotypically and phenotypically resistant divided by the number of isolates that were phenotypically resistant and then multiplied by 100%. The specificity of WGS was defined as the number of isolates that were both genotypically and phenotypically susceptible divided by the number of isolates that were phenotypically susceptible and then multiplied by 100%. 21
Results
Phenotypic antimicrobial resistance
We found that 67 of 77 (87%) turkey isolates were resistant to at least one of the above antimicrobials; 15 of 20 (75%) chicken isolates were susceptible (Fig. 1A; Suppl. Table 1tant to aminoglycoside, beta-lactam, and tetracycline classes (Suppl. Table 1). The chicken MDR isolate was resistant to aminoglycoside, beta-lactam, and tetracycline classes (Suppl. Table 1). The prevalence of resistance in the chicken isolates was lower than in the turkey isolates. Most turkey isolates had resistance to streptomycin (58 of 77 isolates; 75%), gentamicin (51 of 77 isolates; 66%), and amoxicillin (41 of 77 isolates; 53%); the most prevalent resistance, streptomycin resistance, and amoxicillin resistance, in chicken isolates, were only 15% (3 of 20 isolates; Fig. 1B).
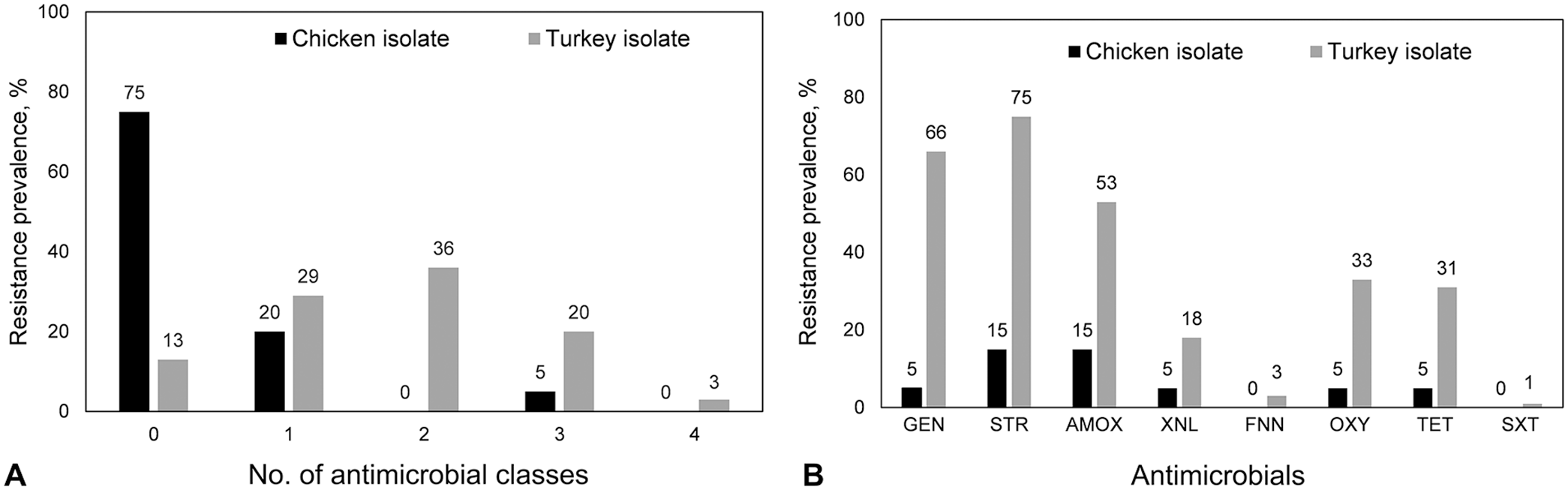
Phenotypic resistance of poultry Salmonella isolates. The 5 classes of antimicrobials in our study were aminoglycosides (GEN = gentamicin; STR = streptomycin), beta-lactams (AMOX = amoxicillin; XNL = ceftiofur), phenicols (FFN = florfenicol), tetracyclines (OXY = oxytetracycline; TET = tetracycline), and trimethoprim (SXT = trimethoprim/sulfamethoxazole).
The 97 poultry Salmonella isolates belonged to 24 serovars, which were initially determined by NVSL and confirmed by WGS. All isolates of the top 6 serovars were susceptible to florfenicol and trimethoprim/sulfamethoxazole (Table 1); the remaining 18 serovars had < 5 isolates per serovar (Suppl. Table 2). All serovar I4,[5],12:i:- isolates were resistant to streptomycin, amoxicillin, oxytetracycline, and tetracycline. Isolates of serovars Senftenberg, Albany, Bredeney, Montevideo, and I4,[5],12:i:- exhibited much higher rates of resistance (50.0–88.9%) to gentamicin than serovar Typhimurium isolates (8.3%).
Phenotypic resistance in Salmonella enterica subsp. enterica serovars with >5 isolates.
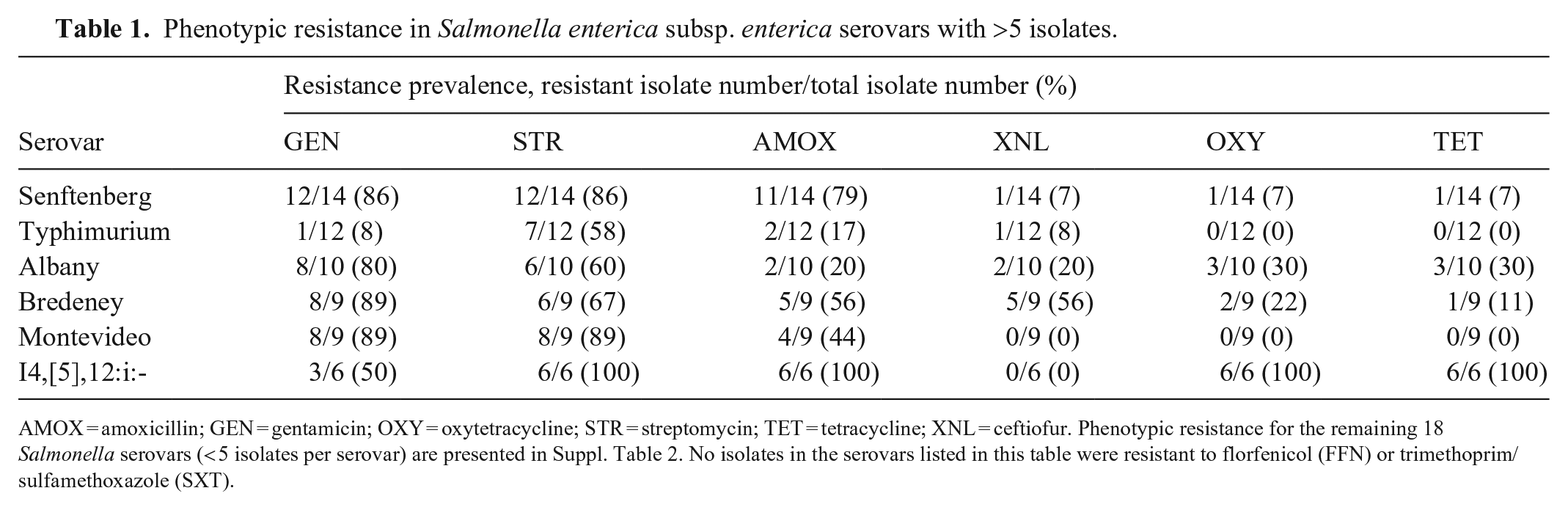
AMOX = amoxicillin; GEN = gentamicin; OXY = oxytetracycline; STR = streptomycin; TET = tetracycline; XNL = ceftiofur. Phenotypic resistance for the remaining 18 Salmonella serovars (< 5 isolates per serovar) are presented in Suppl. Table 2. No isolates in the serovars listed in this table were resistant to florfenicol (FFN) or trimethoprim/sulfamethoxazole (SXT).
No clear AMR trend was observed for any of the antimicrobials in the data collection period from 2010 to 2017 (Suppl. Fig. 1).
Agreement of genotypic and phenotypic AMR
Using WGS, we identified 31 AMRGs, 16 genes responsible for resistance to aminoglycosides, 4 for beta-lactams, 3 for tetracyclines, 2 for sulfonamides, and 1 of each for fosfomycin, quinolone, phenicol, trimethoprim, bleomycin, and colistin (Table 2). Among the 97 isolates studied, 30 isolates (31%) had no known AMRGs; more than half (52 isolates; 54%) carried 3 or more AMRGs, and 1% (1 isolate) carried as many as 12 (Table 3).
Antimicrobial resistance gene (AMRG) prevalence in 97 Salmonella isolates.
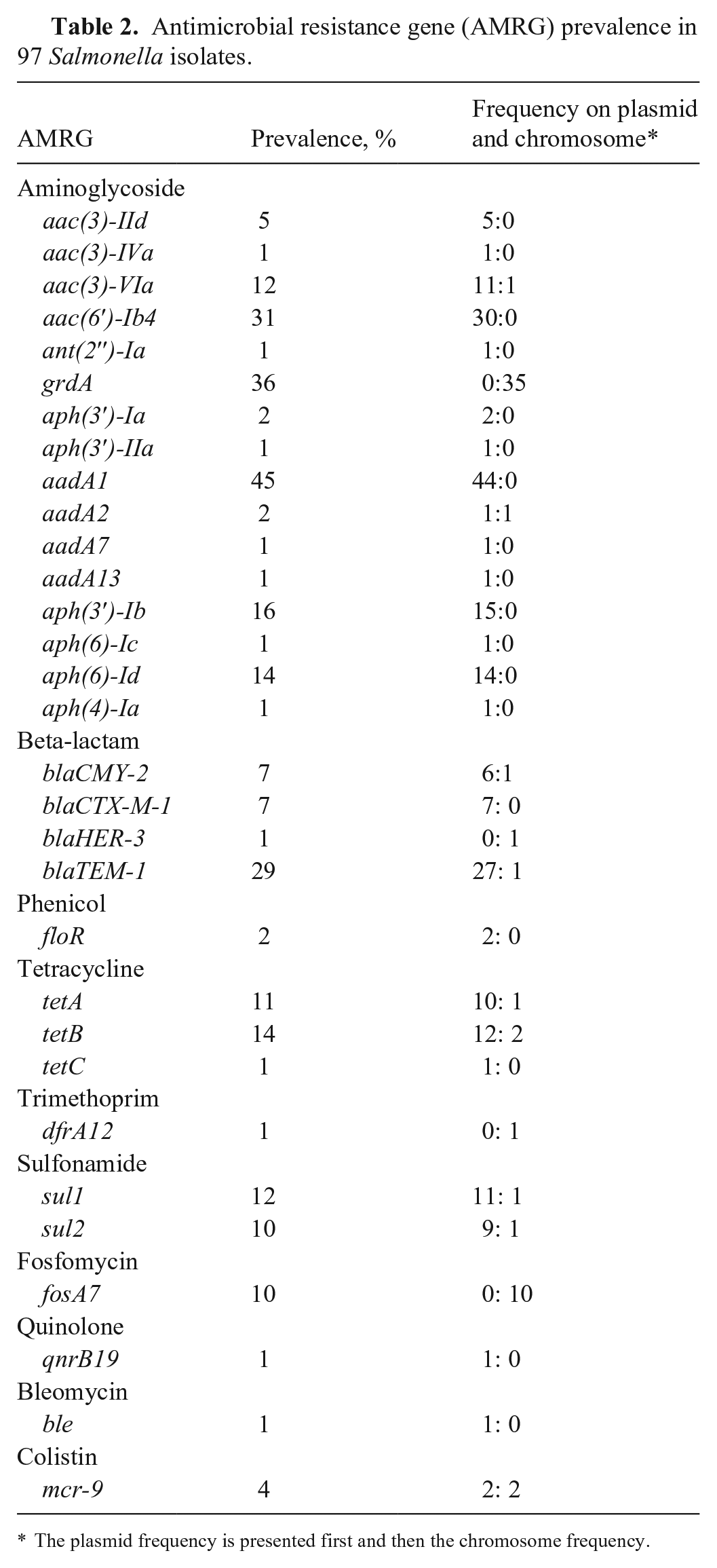
The plasmid frequency is presented first and then the chromosome frequency.
Salmonella isolates carrying multiple antimicrobial resistance genes (AMRGs).
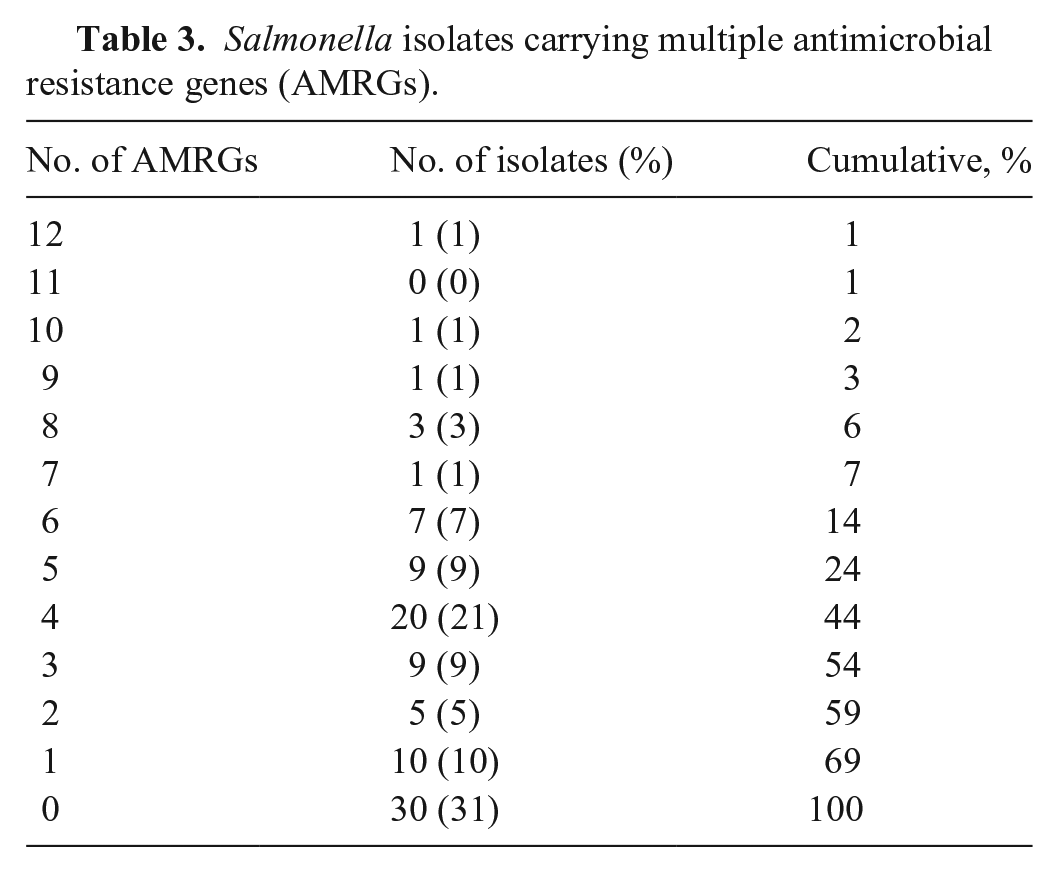
Genotypic resistance determined by WGS was highly consistent with phenotypic resistance (Table 4). The overall sensitivity and specificity of WGS-based prediction of AMR were 93.4% and 99.8%, respectively. There were 16 disagreements involving 15 isolates. Of the 15 isolates, 14 were phenotypically resistant to streptomycin (10), amoxicillin (2), oxytetracycline (1), and amoxicillin/ceftiofur (1); the remaining isolate was phenotypically susceptible but carried AMRGs (Table 4; Suppl. Table 1).
Correlation of genotypic and phenotypic resistance in 97 Salmonella isolates.
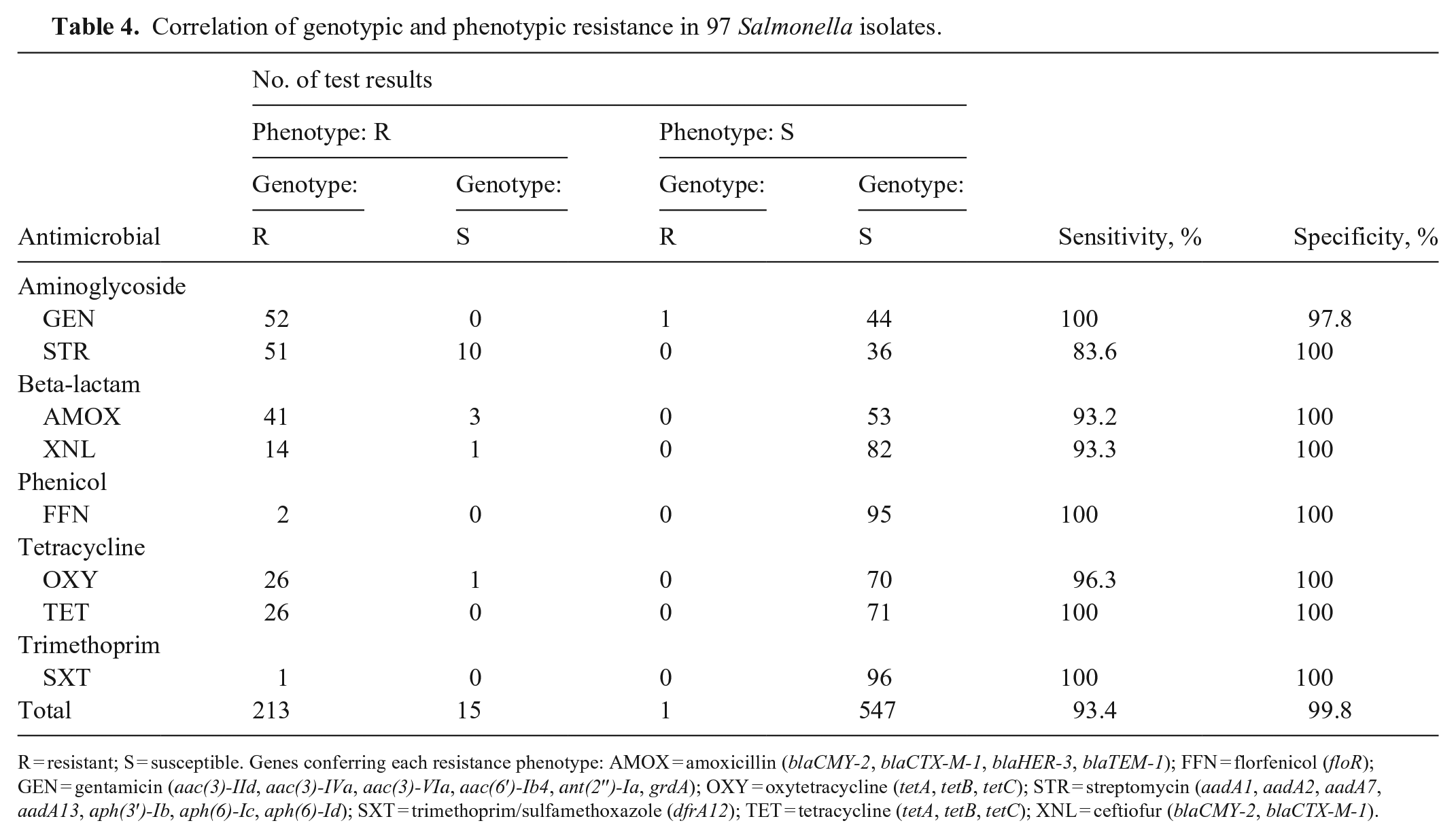
R = resistant; S = susceptible. Genes conferring each resistance phenotype: AMOX = amoxicillin (blaCMY-2, blaCTX-M-1, blaHER-3, blaTEM-1); FFN = florfenicol (floR); GEN = gentamicin (aac(3)-IId, aac(3)-IVa, aac(3)-VIa, aac(6′)-Ib4, ant(2′′)-Ia, grdA); OXY = oxytetracycline (tetA, tetB, tetC); STR = streptomycin (aadA1, aadA2, aadA7, aadA13, aph(3′)-Ib, aph(6)-Ic, aph(6)-Id); SXT = trimethoprim/sulfamethoxazole (dfrA12); TET = tetracycline (tetA, tetB, tetC); XNL = ceftiofur (blaCMY-2, blaCTX-M-1).
Aminoglycoside-resistance genes
The grdA and aac(6′)-Ib4 genes were most frequently associated with gentamicin resistance, and were detected in 35 (36%) and 30 (31%) isolates, respectively (Table 2). The 2 genes co-presented in 27 of the 52 gentamicin-resistant isolates (Suppl. Table 1). An additional 4 gentamicin-resistance genes, aac(3)-IId, aac(3)-IVd, aac(3)-VId, and ant(2′′)-Ia, were detected less frequently (Table 2). WGS accurately predicted gentamicin resistance, except for one turkey isolate that was phenotypically susceptible to gentamicin but carried an aac(6′)-Ib4 gene encoding aminoglycoside-modifying enzymes (Table 4; Suppl. Table 1).
Regarding the aadA genes (aadA1, aadA2, aadA7, aadA13) that confer resistance to both streptomycin and spectinomycin, aadA1 was detected in 44 (45%) isolates; the rest were found in < 5% of isolates (Table 2). Two aph genes, aph(3′)-Ib and aph(6)-Id, often co-presented in the genomes of the streptomycin-resistant isolates, and 14 of 15 aph(3′)-Ib+ isolates also harbored aph(6)-Id (Suppl. Table 1). There were 10 phenotypically streptomycin-resistant isolates (10%), 9 from turkey and 1 from chicken, that did not carry any known resistance genes (Table 4; Suppl. Table 1).
Three neomycin-resistance genes, aph(3′)-Ia, aph(3′)-IIa, and ant(2′′)-Ia, were identified in 2, 2, and 1 turkey isolates (Table 2; Suppl. Table 1). The aph(4)-Ia gene responsible for resistance to hygromycin B was detected in one turkey isolate (Table 2; Suppl. Table 1).
Beta-lactam–resistance genes
Of the beta-lactam–resistance genes identified in our study, blaCMY-2 and blaCTX-M-1 confer both amoxicillin- and ceftiofur-resistance, and blaTEM-1 and blaHER-3 only confer amoxicillin-resistance (Table 2). Two amoxicillin-resistant isolates from turkey carried 2 resistance genes, either blaCTX-M-1 and blaTEM-1 or blaHER-3 and blaTEM-1. The rest of the amoxicillin- or ceftiofur-resistant isolates had only one resistance gene. One turkey isolate that was phenotypically resistant to amoxicillin and ceftiofur, and 2 isolates (1 turkey, 1 chicken) that were phenotypically re-sistant to amoxicillin lacked any known resistance genes (Suppl. Table 1).
Tetracycline-resistance genes
Three resistance genes, tetA, tetB, and tetC, were detected in 11 (11%), 14 (14%), and 1 (1%) isolate, respectively (Table 2). Most isolates, except 1 turkey isolate that carried both tetA and tetB, harbored only 1 of the 3 resistance genes (Suppl. Table 1). One turkey isolate that was phenotypically resistant to oxytetracycline but not tetracycline did not harbor any tet genes. All other isolates resistant to tetracycline and oxytetracycline had identical tet gene profiles (Table 4; Suppl. Table 1).
Phenicol-resistance gene
The floR gene conferring resistance to florfenicol/chloramphenicol was detected in 2 turkey isolates that were also phenotypically resistant to florfenicol (Table 2; Suppl. Table 1).
Trimethoprim-resistance genes
The drfA12 gene conferring resistance to trimethoprim was detected in one turkey isolate, the only isolate that was phenotypically resistant to trimethoprim/sulfamethoxazole. This isolate also carried sul1, a gene conferring resistance to sulfonamide (Table 2; Suppl. Table 1).
Other resistance genes
Other AMRGs were also detected by WGS, including fosA7 for resistance to fosfomycin, qnrB19 for resistance to quinolones, ble for resistance to bleomycin, mcr-9 for resistance to colistin, and sul1 and sul2 for resistance to sulfonamide (Table 2; Suppl. Table 1).
Point mutations that are typically associated with resistance to quinolones (gyrA, gyrB, parC, parE), colistin (pmrA, pmrB), spectinomycin (16S ribosomal DNA), or multidrug resistance (ramR, soxR, soxS) were not detected (Suppl. Table 3).
Resistance genes on plasmids
Plasmid contigs were detected in 92 of the 97 isolates. In total, 27 AMRGs were found 218 times on 119 plasmid contigs in 60 isolates (Suppl. Table 4); 13 AMRGs were detected 58 times on 56 chromosomal contigs in 50 isolates (Suppl. Table 5). Of the 97 isolates, 43 harbored AMRGs on both plasmid and chromosome, 17 carried AMRGs on plasmids only, and 7 had AMRGs on the chromosome only (Table 2). By replicon type, 15 plasmid contigs belonged to IncI-gamma/K1, 13 belonged to IncFIB, 6 belonged to IncQ1, and 2 belonged to IncX1 plasmids, respectively. Other plasmid contigs either lacked the replicon sequences in the contigs or did not have the defined replicons. Among 31 AMRGs that we detected, 18 AMRGs (aac(3)-IId, aac(3)-Iva, aac(6′)-Ib4, aadA1, aadA7, aadA13, ant(2′′)-Ia, aph(3′)-Ia, aph(3′′)-Ib, aph(3′)-Iia, aph(4)-Ia, aph(6)-Ic, aph(6)-Id, blaCTX-M-1, ble, floR, qnrB19, tetC) were located exclusively on plasmid contigs, 4 AMRGs (grdA, blaHER-3, dfrA12, fosA7) were on exclusively chromosomal contigs, and 9 AMRGs (aac(3)-VIa, aadA2, blaCMY-2, blaTEM-1, mcr-9, sul1, sul2, tetA, tetB) were present on both plasmid and chromosomal contigs (Suppl. Tables 4, 5).
Discussion
The NARMS report reveals that 64.3–75.0% of Salmonella isolates from turkey ceca were resistant to at least one class of antimicrobials, with 25.0–47.7% of isolates being MDR in 2013–2017, 41 which is similar to our findings. However, the percentages of resistance to one class antibiotic (52.2–61.5%) and MDR (9.1–24.5%) in chicken cecal isolates in the NARMS report are much higher than the percentages in chicken isolates in our study. 41 The difference might be caused by the small sample size in our study.
We found that WGS accurately predicted the resistance of poultry Salmonella isolates to 8 different antimicrobials, with 16 instances of disagreement between AMR genotypes and phenotypes. The level of accuracy was consistent with previous reports of Salmonella and other bacterial species.21,34,37,42 Of the 16 instances, 15 were phenotypically resistant but genotypically susceptible, indicating a possibility of novel AMRGs carried by these isolates. The remaining instance was an isolate that was phenotypically susceptible but carried silent AMRGs, so-called cryptic genes. 8 Several mechanisms can silence AMRG expression, such as mutation, 18 non-functional promoter, 20 presence of integrons, 44 and presence of negative transcriptional regulators. 15
With regard to streptomycin resistance, NARMS changed the breakpoint from ≥ 64 mg/L to ≥ 32 mg/L in 2013 to better reflect the correlation between phenotype and genotype. 35 We used ≥ 32 mg/L as the breakpoint for all isolates. The broad concentration range of streptomycin (8–1,024 mg/L) in the AVIAN1F plate enabled us to further analyze the relationship between AMRGs and the level of phenotypic resistance. Our data indicated that the most frequently detected streptomycin-resistance gene in poultry Salmonella isolates was aadA1, and this gene alone was associated with moderate MIC levels of 32–256 mg/L. Of the moderately resistant isolates, 3, 7, 19, and 6 isolates had MICs of 256, 128, 64, and 32 mg/L, respectively. The co-presence of aph(3′)-Ib and aph(6)-Id appeared to be associated with high MICs, as evidenced by 9 of 14 isolates having MICs of > 256 mg/L. A synergistic effect of aadA1 and the 2 aph genes was observed in 5 of the 6 isolates having MICs of ≥ 1,024 mg/L. Discrepancies between phenotypic and genotypic resistance to streptomycin were also encountered. Ten isolates classified as resistant based on phenotypic testing lacked any known AMRGs, which warrants an investigation of additional genes involved in streptomycin resistance.
A study reported that different gentamicin-resistance genes contributed to various levels of resistance. 36 With a single gene of aac(3)-IId, aac(3)-VIa, ant(2′′)-Ia, aac(6′)-Ib4, or aac(3)-IVa, 100.0%, 96.3%, 75.0%, 42.9%, and 35.8% of Salmonella isolates had MICs of ≥ 16 mg/L (the clinical breakpoint). 36 We found, however, that the presence of a single resistance gene, aac(3)-IId, aac(3)-VIa, aac(3)-IVa, or grdA contributed to phenotypic resistance to all isolates tested. In addition, 29 of the 30 isolates carrying aac(6′)-Ib4 showed MICs of ≥ 16 mg/L. Of the 30 aac(6′)-Ib4+ isolates, 25 also carried grdA, a gene discovered in 2020. 17 Only 2 isolates carried aac(6′)-Ib4 alone; 1 had an MIC of 1 mg/L and the other had an MIC of ≥ 16 mg/L. Thus, our data cannot support that aac(6′)-Ib4 is associated with a low level of gentamicin resistance as reported previously. 36
We detected 4 genes encoding beta-lactamases, including blaCMY-2, blaCTX-M-1, blaHER-3, and blaTEM-1. We observed that 26 isolates carrying blaTEM-1 alone and 1 isolate carrying both blaHER-3 and blaTEM-1 were phenotypically susceptible to ceftiofur (MICs of 1–2 mg/L), which was consistent with the results of a previous study. 36 It is known that clavulanic acid is a mechanism-based beta-lactamase inhibitor that enhances the antibacterial effectiveness of beta-lactam antimicrobials, resulting in a lower amoxicillin MIC. 12 When clavulanic acid was included in a previous study, 75.2% of Salmonella isolates carrying a single resistance gene, blaTEM-1, had an amoxicillin MIC of ≤ 8 mg/L. 36 In contrast, as shown in our study, when only amoxicillin was used, a single blaTEM-1 gene conferred a higher resistance MIC of ≥ 32 mg/L. Nonetheless, the mechanisms underlying the phenomenon of phenotypically resistant but genotypically susceptible to amoxicillin and ceftiofur remain to be unraveled.
A study reported that most tet+ Salmonella isolates had MICs of ≥ 32 mg/L. 36 In our study, almost all tet+ isolates had MICs of > 8 mg/L to tetracycline and oxytetracycline, which was rounded to ≥ 16 mg/L. It is unclear how many of these isolates have MICs of ≥ 32 mg/L. We did not detect any other tet genes, such as tetD, tetE, tetG, tetM, tetO, or tetX, that have been found in poultry Salmonella isolates.23,25,37
We also identified genes conferring resistance to antimicrobials that are less frequently used in poultry, including fosfomycin, colistin, bleomycin, and quinolone. Fosfomycin, a member of the phosphonic antimicrobial class, inhibits bacterial cell wall biogenesis by inactivating the enzyme UDP-N-acetylglucosamine-3-enolpyruvyltransferase (MurA). 31 This drug is used to treat human urinary tract infections. 24 Colistin, also known as polymyxin E, is a polycationic peptide used as a last-resort drug for the treatment of multidrug-resistant, gram-negative infections in humans. 26 Bleomycin is a glycopeptide antibiotic with a unique mechanism of antitumor activity. 9 Fluoroquinolones were banned by the U.S. Food and Drug Administration (FDA) for use in poultry as of 2005 Sep 12. 40
Mobile genetic elements, including plasmids, phages, and transposons, are known for their role in the horizontal spread of AMRGs between bacterial strains, species, genera, or even higher taxonomic levels. 13 In our study, most Salmonella isolates carried plasmids, including IncI-gamma/K1, IncFIB, IncQ1, and IncX1 plasmids. IncI-gamma/K1 is a subgroup of IncI plasmids, which are frequently detected in E. coli and Salmonella from poultry in Europe. 30 In our study, blaCMY-2, aph(3′′)-Ib, aadA1, aac(3)-VIa, sul1, and tet(A) were all located on the IncI-gamma/K1 plasmid. The IncF plasmids are a group of large plasmids commonly found in Enterobacteriaceae, such as E. coli, S. enterica, and Klebsiella pneumoniae. These plasmids are known to harbor a variety of virulence genes and AMRGs, including those conferring resistance to aminoglycosides, beta-lactams, sulfonamides, tetracyclines, quinolones, macrolides, fosfomycin, trimethoprim, chloramphenicol, and bleomycin. 1 Our data indicated that aadA1, aac(6′)-Ib4, and blaTEM-1 were located on IncFIB, a subgroup of the plasmids. The IncQ plasmids, on the other hand, are small mobilizable plasmids that carry a range of AMRGs, including sul2, strA, strB, tetA, tetB, tetR, cat, aph(3′)-Ia, aph(3′′)-Ib, aph(6)-Id, and blaTEM-1.16,22,43 In our study, all I4,[5],12:i:- isolates carried the same IncQ1 plasmid harboring aph(6)-Id, aph(3′′)-Ib, and sul2. Previously, blaTEM-52 was found on IncX1A plasmids in Salmonella serovars Blockley, Paratyphi, Typhimurium, and Virchow isolated from poultry sources. 3 We detected an IncX1 plasmid contig carrying the blaTEM-1 gene in 2 S. Saintpaul isolates.
The strong relationship that we observed between WGS genotypes and AST phenotypes affirms the reliability of WGS for predicting antimicrobial resistance in diagnostic bacterial isolates. However, there are small numbers of isolates that are either phenotypically resistant yet genotypically susceptible or genotypically resistant but phenotypically susceptible, which indicates that AST cannot be replaced entirely by WGS.
Supplemental Material
sj-pdf-1-vdi-10.1177_10406387241242118 – Supplemental material for Comparison of genotypic and phenotypic antimicrobial resistance profiles of Salmonella enterica isolates from poultry diagnostic specimens
Supplemental material, sj-pdf-1-vdi-10.1177_10406387241242118 for Comparison of genotypic and phenotypic antimicrobial resistance profiles of Salmonella enterica isolates from poultry diagnostic specimens by Zhenyu Shen, C. Y. Zhang, Tamara Gull and Shuping Zhang in Journal of Veterinary Diagnostic Investigation
Supplemental Material
sj-pdf-2-vdi-10.1177_10406387241242118 – Supplemental material for Comparison of genotypic and phenotypic antimicrobial resistance profiles of Salmonella enterica isolates from poultry diagnostic specimens
Supplemental material, sj-pdf-2-vdi-10.1177_10406387241242118 for Comparison of genotypic and phenotypic antimicrobial resistance profiles of Salmonella enterica isolates from poultry diagnostic specimens by Zhenyu Shen, C. Y. Zhang, Tamara Gull and Shuping Zhang in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
We thank the staff of the University of Missouri Veterinary Medical Diagnostic Laboratory Bacteriology Section for providing bacterial isolates and antimicrobial susceptibility testing data to our study.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
