Abstract
The present study describes antimicrobial resistance patterns of Salmonella enterica subspecies enterica serovar Dublin (S. Dublin) in clinical submissions from calves and temporal and farm-type trends in antimicrobial resistance patterns of the isolates. A total of 300 isolates of S. Dublin were obtained from fecal or internal organs of calves fewer than 120 days of age originating from 84 dairies and 18 calf ranches from July 1998 to December 2002. The isolates were susceptibility tested to a panel of 10 antimicrobials using the disk diffusion assay. Temporal and farm-type trends in individual antimicrobial inhibition zone sizes were assessed and antimicrobial resistance patterns were described using cluster analysis. Isolates obtained from calf ranches compared with dairies exhibited decreased susceptibility to florfenicol, gentamicin, neomycin, sulfisoxazole, sulfamethoxazole/trimethoprim, and tetracycline. During the years 1998–2002, decreasing susceptibility was seen for ceftiofur, enrofloxacin, and sulfamethoxazole/trimethoprim. There were 20 different antimicrobial resistance patterns in the isolate set, indicating that S. Dublin has the ability to transfer and pick up resistance genes with relative ease. The trends seen in antimicrobial resistance in S. Dublin may likely be linked to antimicrobial drug use in young calves.
Salmonella enterica subspecies enterica serovar Dublin (S. Dublin) is a host-adapted pathogenic bacteria associated with high morbidity and mortality in young calves. 20 Calf health challenges such as diarrhea, pneumonia, and high mortality caused by S. Dublin have been seen for several decades. 7,18,19 Salmonella Dublin is known for its ability to create a carrier state in cattle. 15 Its environmental persistence 16 makes the infection difficult to manage on farms. A study in the United Kingdom revealed the existence of a variety of clones in both animals and humans. 9 A few prevalent clones seemed widely disseminated among different animal species and showed a diverse geographical and temporal distribution. The same clones were found in animals and humans, which may suggest that both farm and pet animals may act as potential vehicles of infection for humans. Salmonella Dublin is infrequently associated with human infection. However, when a bacteremia is present death is a common sequelae. 10
Salmonella Dublin has been characterized by its multiple antimicrobial resistance patterns. This variability has been attributed to plasmid-borne resistance genes. 6 A clinical trial did not find an association between antimicrobial resistance and virulence. 16 A study using diagnostic laboratory source isolates from bovine animals in Washington detected high levels of multiple antimicrobial resistance in S. Dublin and a decreasing yearly trend in the levels of sensitivity to ceftazidime. 4 It was hypothesized that S. Dublin is highly cattle-adapted, and that the antimicrobial resistance in S. Dublin is expected to be the result of selection within the cattle reservoir. The objective of this study was to describe farm-type and temporal trends in antimicrobial resistance in S. Dublin from clinical submissions from dairy source calves.
Salmonella Dublin isolates from 300 submissions from calves fewer than 4 months old from the central San Joaquin Valley in California submitted to the California Animal Health and Food Safety Laboratory (CAHFS; Tulare, CA) from the period of July 1998 to December 2002 were studied. Salmonella Dublin isolates were obtained from 50 fecal and 250 carcass samples. Only 1 submission per premise within a 1-day period was included in the analysis. The isolate set included isolates from 18 calf ranches and 84 dairies. Salmonella culture and isolation was performed according to standard CAHFS procedures, and serotyping was performed at CAHFS, San Bernardino, California. 3,17
Antimicrobial susceptibility profiles were performed using the disk diffusion assay in accordance with the Clinical and Laboratory Standards Institute (CLSI) guidelines. 14 Ten antimicrobials were evaluated: ampicillin (AMP, 10 μg), ceftiofur (XNL, 30 μg), enrofloxacin (ENR, 5 μg), florfenicol (FFC, 30 μg), gentamicin (GEN, 10 μg), neomycin (NEO, 30 μg), spectinomycin (SPT, 100 μg), sulfamethoxazole/trimethoprim (SXT, 23.75/1.25 μg), sulfonamides (SUL, 250 μg), and tetracycline (TET, 30 μg). The Clinical and Laboratory Standards Institute has determined zone size cut points for classification of bacteria as sensitive (S), intermediate (I), or resistant (R) to an antimicrobial, based on human source Escherichia coli. These ranges are based on therapeutic levels necessary to treat E. coli–associated disease in humans, and are not directly applicable to Salmonella isolates obtained from animals. When looking at temporal or environmental trends, microbiological cut points based on the distribution of the zone sizes for the isolate set may be more appropriate. 13 Distributions of the inhibition zone sizes to the 10 antimicrobials were assessed through histograms and break points for classification as susceptible or resistant (S/R) were determined based on the trough in the bimodal distribution. In the case of ENR, where no bimodal distribution was detected, the S/R cut point was set at the lower limit of the normal distribution (19 mm).
Mean antimicrobial inhibition zone sizes (in mm) of Salmonella enterica subspecies enterica serovar Dublin from clinical submissions from dairy source calves in central San Joaquin Valley in California during the years 1998–2002.
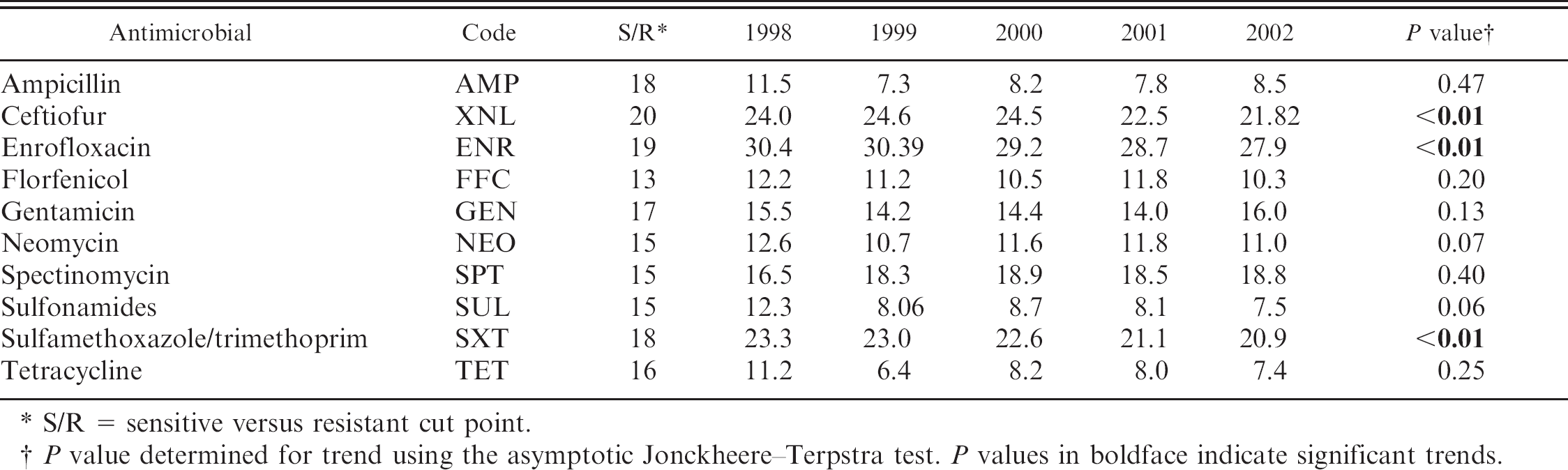
S/R = sensitive versus resistant cut point.
P value determined for trend using the asymptotic Jonckheere-Terpstra test. P values in boldface indicate significant trends.
The resistance patterns of the isolates were described using cluster analysis. Using the squared Euclidean distance as a dissimilarity measure and Ward's minimum variance method, 5 the isolates with similar inhibition zone patterns were grouped into antimicrobial resistance (AR) clusters as previously described. 2 The clusters were ordered in increasing levels of multiple antimicrobial resistance.
Associations between year of isolation (1998–2002) and decreasing mean zone sizes were tested with the non-parametric asymptotic Jonckheere–Terpstra test. 8 Associations between farm-type and decreasing mean disk diffusion zone sizes were tested with the nonparametric asymptotic Kruskal–Wallis test. Decreasing susceptibility over the years were significant (P value ≤ 0.05) for XNL, ENR, and SXT (Table 1). Decreased susceptibility in isolates from calf ranches compared with dairies was significant for FFC, NEO, SUL, SXT, and TET (Table 2). For the antimicrobials that had significant farm-type and year trends, multivariate analysis was performed. A general linear regression model 11 was used to test for both temporal and farm-type effect for the 2 antimicrobials that had a normal zone size distribution (i.e., XNL and ENR). The model indicated a yearly trend of decreasing zone sizes for both XNL and ENR, and S. Dublin isolates from calf ranches had smaller zone sizes for ENR than dairies (Table 3). Logistic regression analysis was performed for the antimicrobial SXT with non-normal bimodal zone size distribution having both significant year and farm-type trend; outcome (dependent) variable was resistant or sensitive as defined in Table 1. 11 This analysis did not reveal any significant interaction effects between farm type and year, thus confirming the univariate analysis (Tables 1, 2).
The isolate set contained 20 AR clusters exhibiting resistance to 0–7 antimicrobials (Table 4). The antimicrobials classified in order of most to least frequent resistance in the data set were AMP, TET SUL, NEO, FFC, GEN, SPT, XNL, and SXT. No isolates exhibited zone sizes classified as resistant for ENR. The most common cluster included resistance to AMP, TET, SUL, NEO, FFC, and GEN. The distribution of AR clusters revealed a large variability on both dairies and calf ranches (data not shown).
The current study has shown that routine diagnostic animal health data can be of value to monitor trends in antimicrobial resistance in production animal pathogenic and zoonotic bacteria. Because of the nature of the data set, the increased number of isolates over the years could either reflect increased prevalence of S. Dublin, increased submissions to the diagnostic laboratory, or a combination of these.
The S. Dublin isolate set demonstrated significant decreasing susceptibility to 3 of 10 antibiotics (ENR, SXT, and XNL) over the 5 years, with SUL exhibiting a decreasing trend (P value = 0.06). Decreasing susceptibility over time to antimicrobials has been noted in S. Dublin in the National Antimicrobial Resistance Monitoring System (http://www.ars.usda.gov/main/site_main.htm?modecode=66–12–05–08). This decrease was also noted for ceftazidime in S. Dublin in Washington, 4 which may be a result of the adaptation of the bacteria to antimicrobial pressure, including adaptive mutations and acquired resistance mechanisms. 12 As S. Dublin is associated with clinical disease in bovine animals, it is likely that antimicrobials may be used on farms when this bacteria is present, creating a selection pressure for increasingly resistant strains.
Mean antimicrobial inhibition zone size (in mm) of Salmonella enterica subspecies enterica serovar Dublin from clinical submissions from calves on calf ranches and dairies.
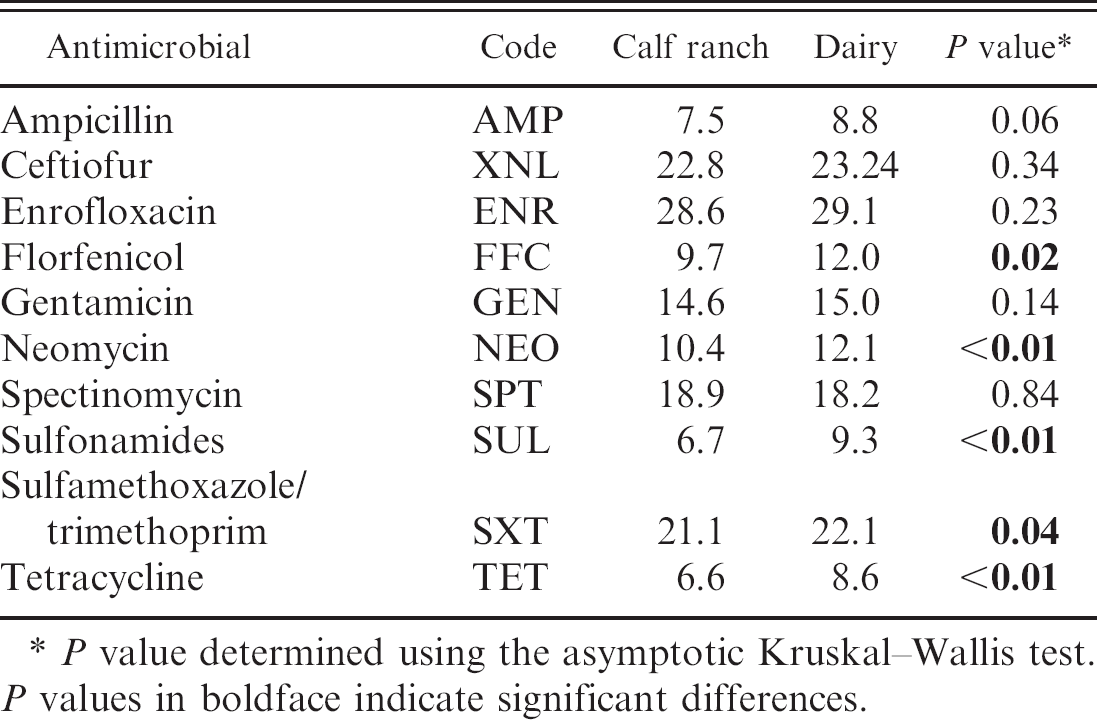
P value determined using the asymptotic Kruskal–Wallis test. P values in boldface indicate significant differences.
Mean zone size trends (in mm) for enrofloxacin in Salmonella enterica subspecies enterica serovar Dublin from clinical submissions from calves on calf ranches and dairies.*
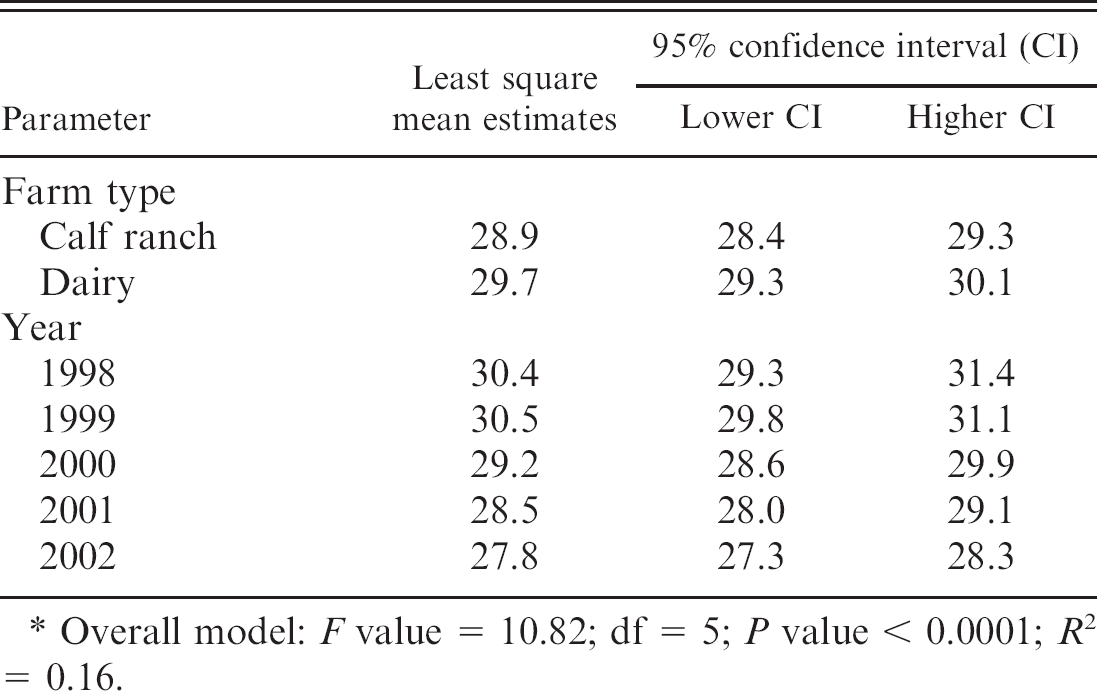
Overall model: F value = 10.82; df = 5; P value < 0.0001; R 2 = 0.16.
The differences in levels of antimicrobial resistance and decreasing susceptibility in calf ranches compared with dairies for 5 of 10 antimicrobials tested may be a result of the common use of prophylactic antimicrobials added to the feed for calf ranch calves. An increased therapeutic antimicrobial use on calf ranches may also create a selection pressure for higher levels of resistance in S. Dublin. There may be increased disease treatments on calf ranches because of the transport of neonatal calves from the dairies to the calf ranch, commingling of calves from several dairies, a large number of colostrum-deprived calves, suboptimal feeding, and other stressors. Although these hypotheses were not corroborated by a study of general fecal shedding of Salmonella in dairy source preweaned calves in California, 1 S. Dublin was only 3% of the isolates in that study, and it would be of interest to evaluate the influence of these farm management–specific factors on a calf-adapted pathogenic serovar, such as S. Dublin. A previous study found more resistance to 6 antibiotics tested from dairy source isolates than from beef breed source isolates indicating that the antimicrobial use, which is likely to be more intense on dairies than in beef cow calf operations, may influence the levels of resistance in S. Dublin. 4
The current study suggests that antimicrobial susceptibility should be evaluated when S. Dublin–associated disease is present in calves to optimize treatment success. Alternative treatment strategies should be investigated for minimizing S. Dublin–associated disease to minimize treatment failure associated with antimicrobial resistance.
Antimicrobial resistance patterns (AR clusters) of Salmonella enterica subspecies enterica serovar Dublin isolates from diagnostic samples from calves grouped into AR clusters 1–20.
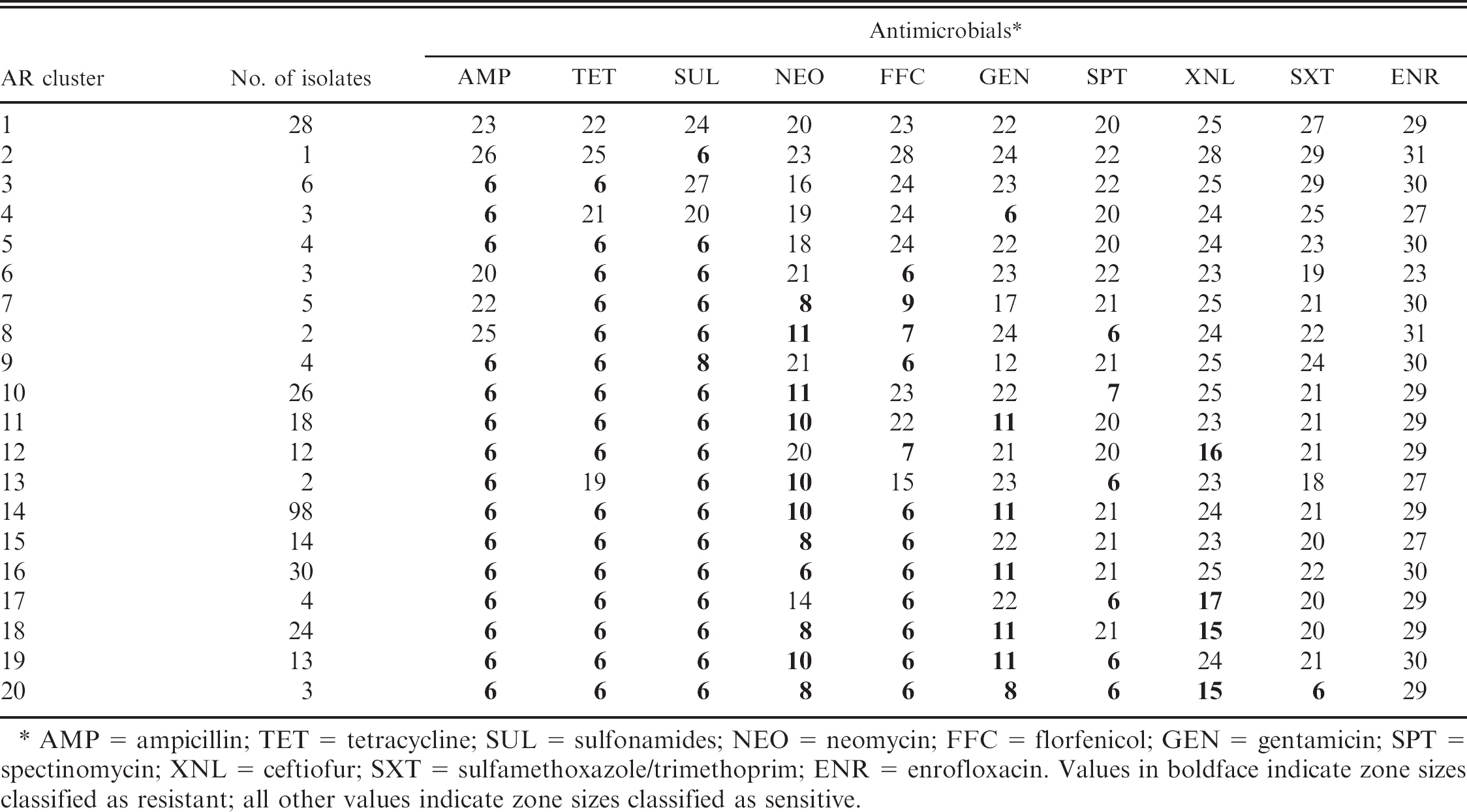
AMP = ampicillin; TET = tetracycline; SUL = sulfonamides; NEO = neomycin; FFC = florfenicol; GEN = gentamicin; SPT = spectinomycin; XNL = ceftiofur; SXT = sulfamethoxazole/trimethoprim; ENR = enrofloxacin. Values in boldface indicate zone sizes classified as resistant; all other values indicate zone sizes classified as sensitive.
Acknowledgements. The authors would like to express their gratitude to CAHFS for saving all diagnostic isolates and associated data, and making them available for this study.
