Abstract
The objective of the current study was to determine the capability of 3 recently described one-step TaqMan real-time reverse transcription polymerase chain reaction (real-time RT-PCR) assays targeting the nucleoprotein (NP), matrix (M), and hemagglutinin (HA) genes of H3N8 Equine influenza virus (EIV NP, EIV M, and EIV HA3 assays, respectively) to detect Canine influenza virus (CIV). The assays were initially evaluated with nucleic acid extracted from tissue culture fluid (TCF) containing the A/canine/FL/43/04 strain of Influenza A virus associated with the 2004 canine influenza outbreak in Florida. The EIV NP, EIV M, and EIV HA3 assays could detect CIV nucleic acid at threshold cycle (Ct) values of 16.31, 23.71, and 15.28, respectively. Three assays using TCF or allantoic fluid (AF) samples containing CIV (n = 13) and archived canine nasal swab samples (n = 20) originally submitted for laboratory diagnosis of CIV were further evaluated. All TCF and AF samples, together with 10 nasal swab samples that previously tested positive for virus by attempted isolation in embryonated hens' eggs or Madin–Darby canine kidney cells, were positive in all 3 real-time RT-PCR assays. None of the 3 assays detected the H1N1 Swine influenza virus strain in current circulation. These findings demonstrate that previously described real-time RT-PCR assays targeting NP, M, and H3 HA gene segments of H3N8 EIV are also valuable for the diagnosis of CIV infection in dogs. The assays could expedite the detection and identification of CIV.
Keywords
Influenza A viruses infect a variety of mammalian and avian species, including pigs, horses, birds, certain wildlife species, and human beings. 17 Interspecies transmission of Influenza A virus (FLUAV; family Orthomyxoviridae, genus Influenzavirus A) plays an important role in its ecology and epidemiology. 16 A significant outbreak of canine influenza (CI) was reported in Greyhound dogs in Florida in early 2004, and it was shown to be due to cross-species transmission of H3N8 Equine influenza virus (EIV). 2 Furthermore, there are reports of experimental transmission of H5N1 and natural transmission of H3N2 avian influenza viruses to dogs. 1,7,8,10,12–14 More recently, it has been reported that a novel H1N1 FLUAV, a reassortant of human, swine, and avian influenza viruses, is capable of infecting dogs (ProMED-mail: 2009, Influenza pandemic [H1N1] 2009, animal [30]: China, canine. Available at http://www.promedmail.org/pls/apex/f?p=2400:1202:5360013597818245::NO::F2400_P1202_CHECK_DISPLAY, F2400_P1202_PUB_MAIL_ID:X, 80277. Accessed June 25, 2010). 5,6 Samples from 2 sick dogs tested positive for H1N1 FLUAV, and both virus isolates were 99% identical with the strains isolated from human beings (ProMED-mail: 2009, Influenza pandemic [H1N1] 2009). However, the significance of transmission of influenza A viruses to dogs from other mammal and avian species including human beings has yet to be determined.
Since the initial CI outbreak in Florida, evidence of virus activity has been reported in 30 states in the United States, reportedly affecting tens of thousands of dogs. 2,4 Sequence analysis of Canine influenza virus (CIV) from the original outbreak (A/canine/FL/43/04) revealed that 8 gene segments of this virus shared ≥96% nucleotide sequence homology with that of the H3N8 EIV subtype, suggesting that CIV resulted from direct interspecies transmission of EIV to dogs without genetic reassortment. 2,11 The sequence changes in the hemagglutinin gene (H3 HA) are indicative of adaptive evolution of the virus in its new host. Phylogenetic analysis of 4 Greyhound isolates and 2 pet dog isolates revealed that these viruses were closely related to EIV strains recovered from horses since 2000, belonging to the Florida sublineage of the American lineage of H3N8 EIV. 2,11 The isolation of 4 closely related H3N8 EIV subtype strains from dogs that died in different locations over a 2-year period, together with serologic evidence of widespread infection among Greyhounds in different states, strongly confirmed sustained circulation of CIV involving dog-to-dog transmission of the virus. 2,11,18
Primer and probe sequences used in the Equine influenza virus (EIV) nucleoprotein (NP) gene, EIV matrix (M) gene, EIV hemagglutinin (HA3) gene, and H1N1 real-time reverse transcription polymerase chain reaction (real-time RT-PCR) assays.

Reporter dye (6-carboxyfluorescein; FAM)-labeled nucleotide.
Nonfluorescent quencher dye (minor groove binding; MGB)-labeled nucleotide.
BHQ1 (black hole quencher 1) dye-labeled nucleotide.
Most outbreaks of CI occur in rescued, kenneled, or boarded dogs. 4 Clinical signs of the disease include low-grade fever, lethargy, anorexia, nasal discharge, and prolonged cough. 4,11 These signs can mimic those caused by other canine respiratory pathogens. Accordingly, it is especially important to be able to distinguish CIV from other agents capable of causing canine respiratory disease. 4 Detection of CIV at the early stage of infection is especially important for facilitating management and control of the disease. To date, few publications have described specific diagnostic assays to detect CIV in clinical specimens. Detection of CIV is frequently attempted by traditional virus isolation (VI) in embryonated hens' eggs or in Madin–Darby canine kidney (MDCK) cells. 4 There have been some conflicting reports over the sensitivity of antigen-capture enzyme-linked immunosorbent assays for the detection of CIV in dogs. 4,15 Presence of CIV nucleic acid in clinical specimens has been confirmed by traditional reverse transcription polymerase chain reaction (RT-PCR) and real-time RT-PCR assays targeting the matrix (M) gene of Avian influenza virus or by an real-time RT-PCR assay specific for the H3 HA or the M gene of CIV. 4,11,15 Recently, 3 real-time RT-PCR assays targeting the nucleoprotein (NP), M, and H3 HA genes of H3N8 subtype of EIV (EIV NP, EIV M, and EIV HA3 assays, respectively; Table 1) were developed and evaluated. 9 The EIV NP, EIV M, and EIV HA3 primers and probe sets were 100% identical to the published NP, M, and H3 HA sequences of CIV, respectively. The purpose of the current study was to evaluate the capability of the EIV NP, EIV M, and EIV HA3 assays to detect not only H3N8 CIV but also the novel H1N1 FLUAV that has been in widespread circulation around the world and that has the proven capacity to infect dogs (ProMED-mail: 2009, Influenza pandemic [H1N1] 2009). 2
Tissue culture fluid (TCF) containing the A/canine/FL/43/04 isolate and 13 other archived CIV isolates from the Animal Health Diagnostic Center at Cornell University (Ithaca, New York) and the University of Kentucky Veterinary Diagnostic Laboratory (Lexington, Kentucky) were tested in the study. Also included were 20 archived nasal swabs from dogs with signs of respiratory illness, 10 of which had previously tested positive for CIV by VI in embryonated hens' eggs or MDCK cells according to standard laboratory protocols. Nasal specimens were collected using cotton or Dacron swabs moistened with a few drops of normal saline or viral transport medium. The presence of CIV in TCF and allantoic fluid (AF) had previously been confirmed by sequencing the entire viral genome (n = 11) or the H3 HA gene (n = 3). Of the 10 CIV-positive nasal swab samples, 3 were previously diagnosed as positive by real-time RT-PCR targeting the M gene. These 3 samples were subsequently tested by an N1 neuraminidase-specific RT-PCR, thus ruling out H1N1 FLUAV. The 7 remaining CIV-positive nasal swab samples were part of a study in which H3N8 virus was continuously circulating in a particular animal shelter. The H3 HA gene from 3 isolates (representative of the total of 7 samples) had been sequenced to confirm that they were H3N8 CIV. A total of 38 nasal swabs from pigs that were experimentally infected with the A/CA/04/2009 strain of H1N1 FLUAV and swabs from 12 control pigs were also included in the study. The presence of H1N1 FLUAV in pig samples was previously confirmed through VI in embryonated hens' eggs. All archived nasal swab specimens and TCF and AF samples were stored at −80°C prior to nucleic acid extraction. Using a commercial magnetic particle processor, a viral nucleic acid was isolated from 50 μl of TCF, AF, or nasal swab samples with a commercial viral RNA isolation kit. b Viral RNA was eluted in 50 μl of nuclease-free water and stored at −80°C until further use. The primers and probes used in the current study were identical to those previously described for the EIV NP, EIV M, and EIV HA3 real-time RT-PCR assays (Table 1). 9 The primers and probe set specific for the H1 HA gene of the novel H1N1 strain were previously described (Table 1). 3 A one-tube real-time RT-PCR assay was performed using a commercial kit and machine c , d under the exact same conditions as previously reported. 9 For each assay, each sample was tested in duplicate.
Detection of Canine influenza virus (CIV) or Swine influenza virus nucleic acid by real-time reverse transcription polymerase chain reaction (real-time RT-PCR) assays targeting nucleoprotein (NP), matrix (M), and hemagglutinin (HA) genes of H3N8 Equine influenza virus (EIV NP, EIV M, and EIV HA3). *
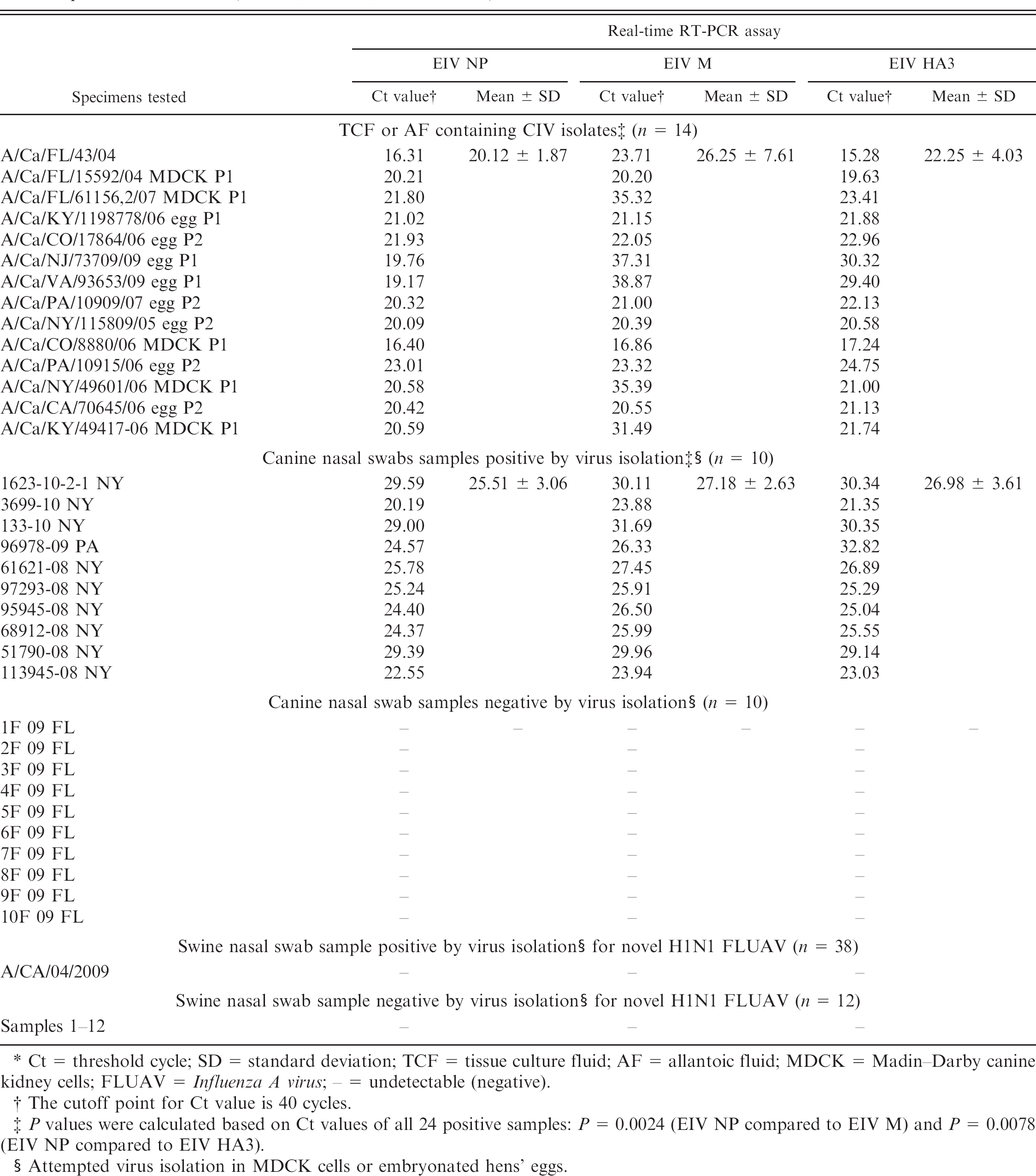
Ct = threshold cycle; SD = standard deviation; TCF = tissue culture fluid; AF = allantoic fluid; MDCK = Madin–Darby canine kidney cells; FLUAV = Influenza A virus; – = undetectable (negative).
The cutoff point for Ct value is 40 cycles.
P values were calculated based on Ct values of all 24 positive samples: P = 0.0024 (EIV NP compared to EIV M) and P = 0.0078 (EIV NP compared to EIV HA3).
Attempted virus isolation in MDCK cells or embryonated hens' eggs.
Initially, the ability of all 3 primer sets to detect CIV using RNA extracted from TCF containing the A/canine/FL/43/04 strain was determined. The real-time RT-PCR assays for EIV NP, EIV M, and EIV HA3 detected CIV RNA with average threshold cycle (Ct) values of 16.31, 23.71, and 15.28, respectively. In contrast, all 3 assays failed to detect influenza virus nucleic acid extracted from the A/CA/04/2009 strain of H1N1 FLUAV. These assays with RNA extracted from the 13 TCF and AF samples containing other CIV isolates were subsequently evaluated. All samples tested positive, with mean Ct values ranging from 20.12 ± 1.87 (mean ± standard deviation) with EIV NP, to 26.25 ± 7.61 with EIV M, and to 22.25 ± 4.03 with EIV HA3 assays. Comparison of Ct values indicated that the EIV NP assay had the lowest Ct value compared to the EIV M and EIV HA3 assays (P = 0.0095 and P = 0.0447, respectively).
Moreover, of the 20 canine nasal swab samples tested in the present study, 10 of the canine nasal swabs, which were positive through VI, were also found to be positive by all 3 real-time RT-PCR assays, with mean Ct values ranging from 25.51 = 3.06 with EIV NP, to 27.18 = 2.63 with EIV M, and to 26.98 = 3.61 with EIV HA3 assays. The remaining 10 nasal swab samples that were negative by VI were also negative in all 3 real-time RT-PCR assays. The EIV NP assay had the lowest Ct values with all 24 samples (TCF, AF, and nasal swab samples) compared to the EIV M and EIV HA3 assays (P = 0.0024 and P = 0.0078, respectively).
Results of the 3 real-time RT-PCR assays using nucleic acid extracted from nasal swabs from dogs were highly correlated with the VI results (100% sensitivity and specificity; Table 2). These assays provide a rapid and convenient means of detecting CIV nucleic acid in nasal swab samples and confirming a diagnosis of CI. The sensitivity of the assays has been previously established to be ≥10 RNA molecules. 9 The primers and probe specific for the H1 HA gene of the novel H1N1 strain failed to recognize the A/canine/FL/43/04 strain or any other H3N8-positive samples included in the present study. In contrast, the assay readily detected the H1N1 viral nucleic acid in the 38 nasal swabs collected from the H1N1 experimentally infected pigs (mean Ct value: 24.78; 100% sensitivity). 3 None of the 12 nasal swab samples collected from control pigs reacted with the H1 HA gene–specific primers and probe (100% specificity). The data clearly demonstrated that the 3 EIV real-time RT-PCR assays and the real-time RT-PCR assay specially designed to detect the novel H1N1 FLUAV are highly specific for each influenza subtype and could be used successfully to diagnose cases of respiratory disease due to either of these viruses in dogs.
In summary, the 3 developed real-time RT-PCR assays recognized both equine and canine H3N8 viruses in clinical specimens and could be used as a routine laboratory test for the diagnosis of influenza in both animal species. The assays are highly specific and do not detect the recently emerged H1N1 strain of influenza virus. Because the EIV NP assay has the highest sensitivity, it is recommended for the routine diagnosis of CIV in dogs. Furthermore, the EIV HA3 assay has the capability to distinguish CI cases resulting from the H3N8 subtype from H1N1 subtype infections, which will not be detected by the developed assay.
Acknowledgements. The present work was supported with funds from the American Quarter Horse Association, the Frederick Van Lennep Chair endowment fund at the Maxwell H. Gluck Equine Research Center, and the Kentucky Agricultural Experiment Station, College of Agriculture, University of Kentucky (project nos. 017027 and 014028). Dr. Zhengchun Lu is the recipient of a Geoffrey C. Hughes Foundation graduate fellowship.
Footnotes
a.
KingFisher® magnetic particle processor, Thermo Fisher Scientific, Waltham, MA.
b.
MagMAX™-96 viral RNA isolation kit, Applied Biosystems, Foster City, CA.
c.
TaqMan® one-step RT-PCR master mix, Applied Biosystems, Foster City, CA.
d.
7500 Fast real-time PCR system, Applied Biosystems, Foster City, CA.
