Abstract
A duplex quantitative real-time polymerase chain reaction (qPCR) assay was developed to differentiate between Bolbophorus damnificus and Bolbophorus type II species cercariae. Both trematode species are prevalent throughout the commercial catfish industry, as both infect the ram's horn snail, Planorbella trivolvis, which is commonly found in catfish ponds. Identification of cercaria to species is important in catfish disease challenge experiments, as only B. damnificus has been shown to have negative impacts on channel catfish. Oligonucleotide primers and fluorescence resonance energy transfer hydrolysis probes were designed to amplify the 18S small subunit ribosomal DNA gene of each species. The quantification cycle indicative of the number of cercariae in the sample prep was determined, and standard curves correlating to cercaria numbers were established. For both species, the assay was found to be highly repeatable and reproducible, with a linear dynamic range covering 7 orders of magnitude. The sensitivity limit of the assay was ∼1/256th of a cercaria, regardless of species, and there was no remarkable interference between the 2 assays when run simultaneously within the same reaction. In a field study, identification of cercaria by the duplex real-time qPCR assay was in complete agreement with previously established end-point PCR protocols, demonstrating the assay to be a more rapid, quantifiable means of parasite identification.
Digenetic trematodes of the genus Bolbophorus have been implicated in significant economic losses throughout the commercial catfish industry. 9,10 As such, this parasite has garnered significant attention from aquatic disease researchers. Two distinct species of Bolbophorus are commonly associated with commercial catfish operations, both of which use the ram's horn snail (Planorbella trivolvis) as an intermediate host. 2,4,7 To gain a better understanding of the pathology associated with infestation by these parasites, disease challenge experiments have been performed by exposing naïve channel catfish (Ictalurus punctatus) to Bolbophorus-type cercaria. This research has shown only one of these species, B. damnificus, to have negative impacts on channel catfish. Naïve fish exposed to high numbers of B. damnificus cercaria died within 6-7 days, whereas high numbers of Bolbophorus type II sp. cercaria did not appear to have any adverse affects. 4 Therefore, to ensure consistency in disease challenge studies, researchers working with channel catfish must ensure they are exposing fish to only B. damnificus cercaria and not Bolbophorus type II sp. In addition, since snails can be infected simultaneously with both trematode species, it is important to identify snails releasing only B. damnificus cercaria and not combinations of the two. Unfortunately, the morphological similarities between the cercaria of these 2 closely related parasites make identification based solely on physical characteristics challenging and time-consuming.
Currently, researchers use endpoint polymerase chain reaction (PCR) to differentiate between B. damnificus and Bolbophorus type II sp. 2,4 Although these methods are reliable, they require postreaction processing and are not readily quantifiable. Quantitative real-time PCR (qPCR) assays are more favorable compared with conventional endpoint PCR as they collect data as the PCR occurs, thereby combining amplification and detection into a single step. Consequently, there is no need for postreaction processing, and since the reaction can be monitored as it progresses, results can be obtained more rapidly than endpoint PCR. 12 The development of real-time qPCR primer and probe combinations specific to each species that can be run simultaneously within the same reaction will provide a more rapid method to identify of Bolbophorus-type cercaria.
Primer sequences used in the current study.
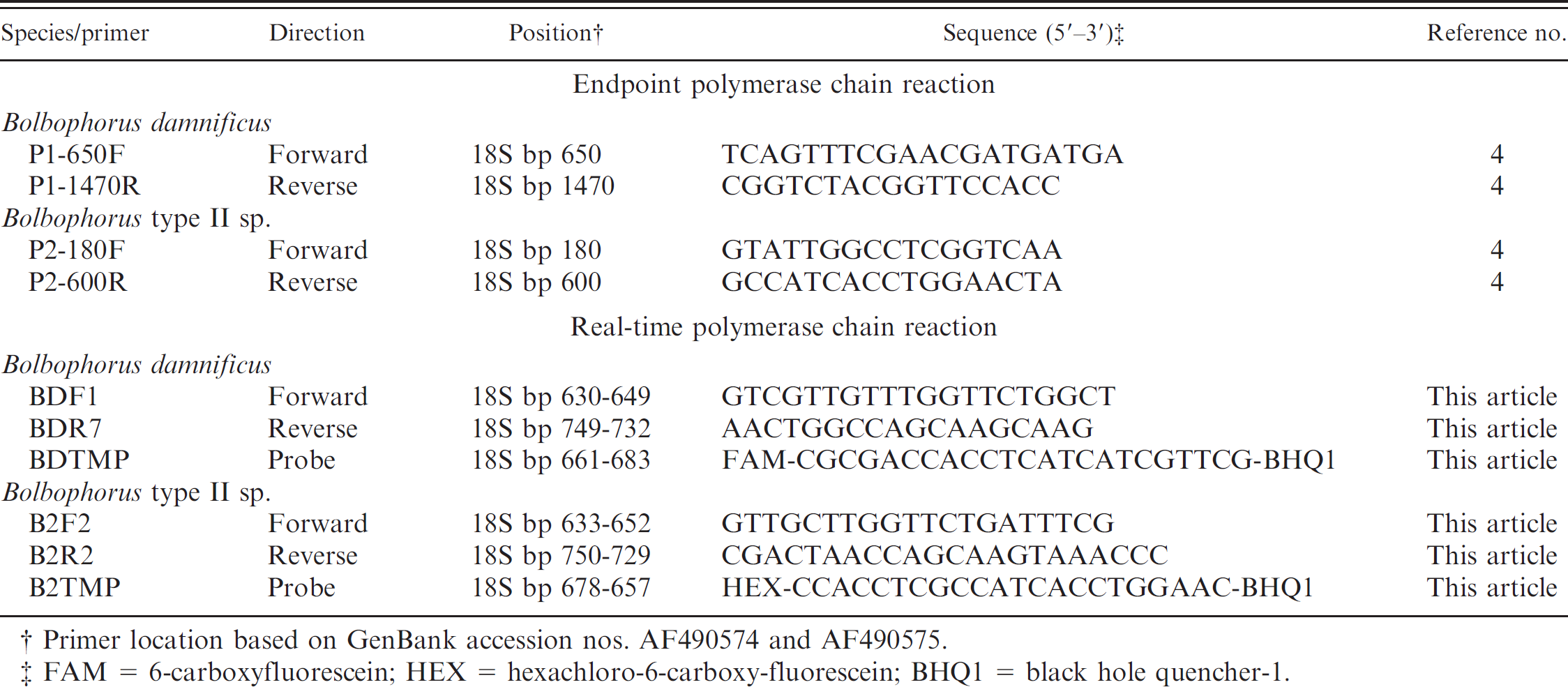

Primer location based on GenBank accession nos. AF490574 and AF490575.
FAM = 6-carboxyfluorescein; HEX = hexachloro-6-carboxy-fluorescein; BHQ1 = black hole quencher-1.
Ram's horn snails (n = 500), collected from a commercial operation with a history of trematode infestations, were rinsed and placed in individual glass scintillation vials with 10 ml of autoclaved pond water that had been passed through a 20-μm nylon mesh screen. The snails were maintained at an ambient temperature of 27°C for 24 hr and then examined microscopically for the release of the characteristic longifurcate cercariae of Bolbophorus spp. 2 Forty-five snails were observed to be shedding cercariae, from which batches of cercariae (∼25 cercariae) were collected from individual snails and placed directly into a 1.5-ml microcentrifuge tube with 70% molecular-grade ethanol. Three different cercaria morphotypes 8 were identified microscopically and collected for standardization of the real-time qPCR primers. The morphotypes included the Bolbophorus-type cercaria, 2 an unidentified amphistome-type cercaria, and an unidentified furcocercous-type cercaria, presumably of the genus Clinostomum (C. Doffitt, unpublished data).
Genomic DNA was isolated from cercariae pools using a commercial DNA isolation kit
a
following the manufacturer's suggested protocol. All cercariae pools were identified as either B. damnificus, Bolbophorus type
Oligonucleotide primers and fluorescence resonance energy transfer hydrolysis probes specific to each species (B. damnificus or Bolbophorus type II sp.) were developed. Primer and probe sequences were designed by exploiting nucleotide differences in the 18S small-subunit ribosomal DNA sequences of the 2 species obtained from the National Center for Biotechnology Information (NCBI) GenBank sequence database (B. damnificus: AF490574; Bolbophorus type II sp.: AF490575). A subsequent BLASTn search for somewhat similar sequences of the NCBI nonredundant nucleotide (nr/nt) database confirmed the primer and probe combinations to be specific to their target sequences. The B. damnificus probe consisted of a 5′ reporter fluorophore (6-carboxyfluorescein [FAM]) while the Bolbophorus type II sp. probe used the 5′ reporter hexachloro-6-carboxyfluorescein (HEX). Black hole quencher-1 served as the 3′ quencher for both probes. All primers and probes are described in Table 1.
Standard curves were developed for both species to serve as a measure of reaction efficiencies. For both curves, standard DNA was obtained by amplifying the target amplicons of the real-time PCR primers by the endpoint PCR protocol described above, and the generation of target amplicons was confirmed through electrophoresis. The amplicons were purified using a centrifugal filter device, d quantified spectrophotometrically, e and serially diluted 10-fold.
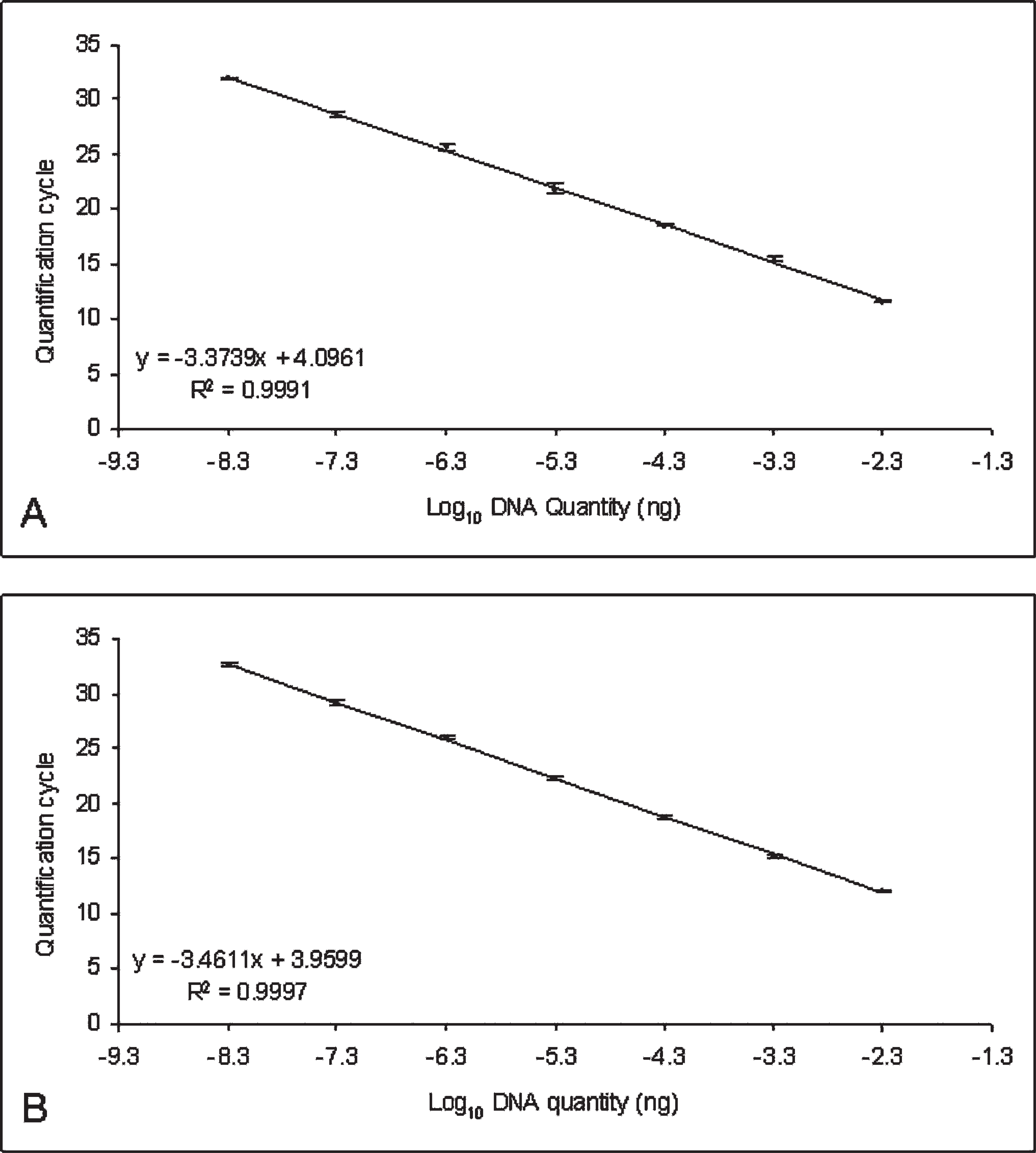
The linear dynamic ranges for detection of Bolbophorus damnificus (
Real-time PCR for simultaneous detection and quantification of B. damnificus and Bolbophorus type II sp. DNA was performed in a quantitative thermal cycler using the accompanying software to conduct data analysis. f Preliminary testing was carried out to optimize the performance of the duplex real-time PCR assay by determining the appropriate primer concentrations and amplification conditions. The optimal 15-μl reaction mixture included 7.5 μl of PCR supermix, 5 pmol of each primer, 1.25 pmol of each probe, and 5 μl of template DNA. The thermal cycling consisted of activation of the Taq DNA polymerase at 95°C for 10 min, followed by 35 cycles of denaturation at 95°C for 15 sec, and annealing/extension at 61°C for 1 min. Data collection was carried out during the annealing/extension step. Reactions were characterized by their quantification cycle (Cq), the point in time (or PCR cycle) at which the target amplification achieved a predetermined threshold where fluorescence intensity was significantly greater than background fluorescence. This threshold was manually set the same for all runs within the region of exponential amplification of the standard positive controls.
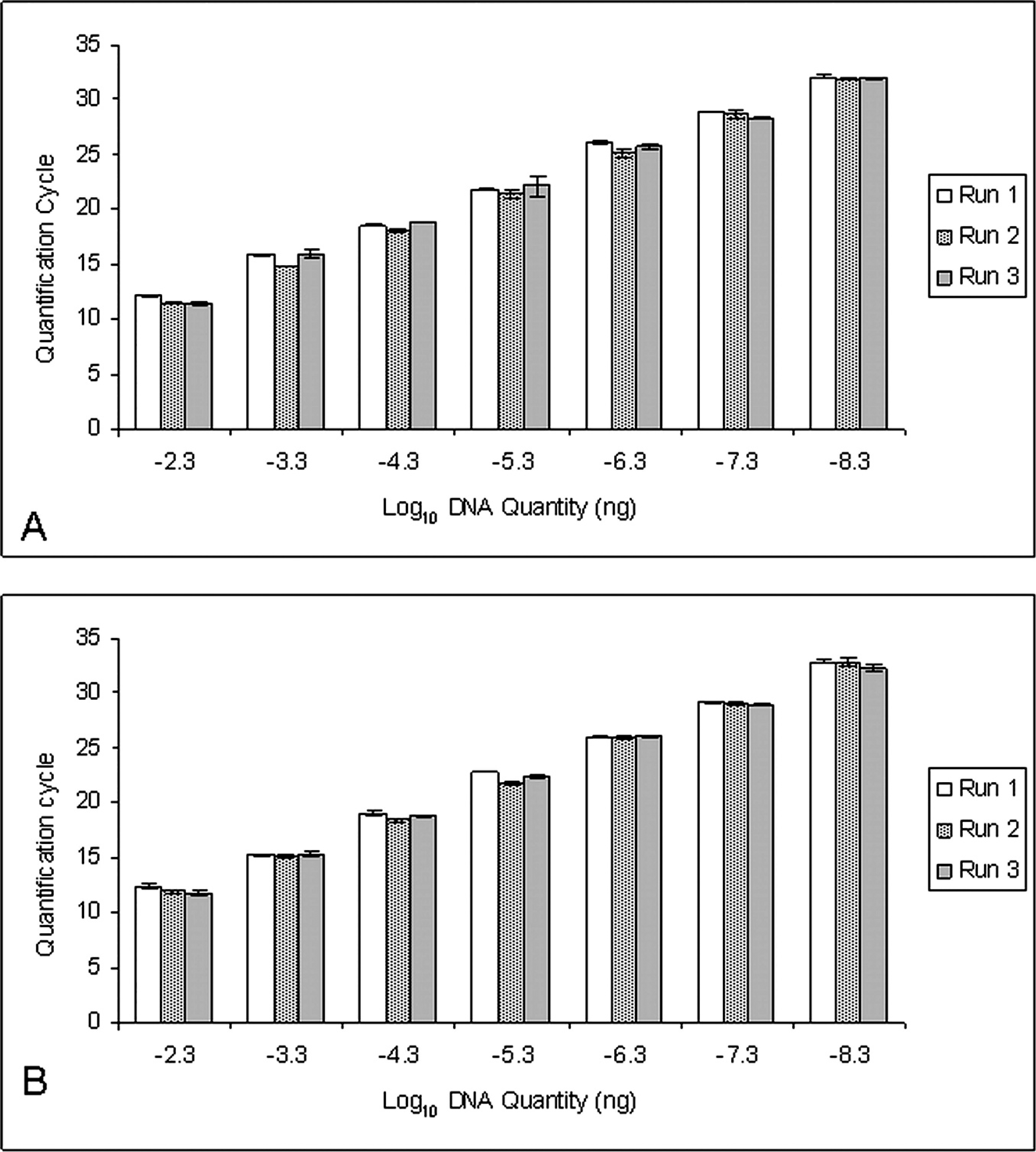
The interassay variability of the Bolbophorus damnificus (
Intra-assay variability of the Bolbophorus damnificus and the Bolbophorus type II sp. duplex real-time quantitative polymerase chain reaction assay.†
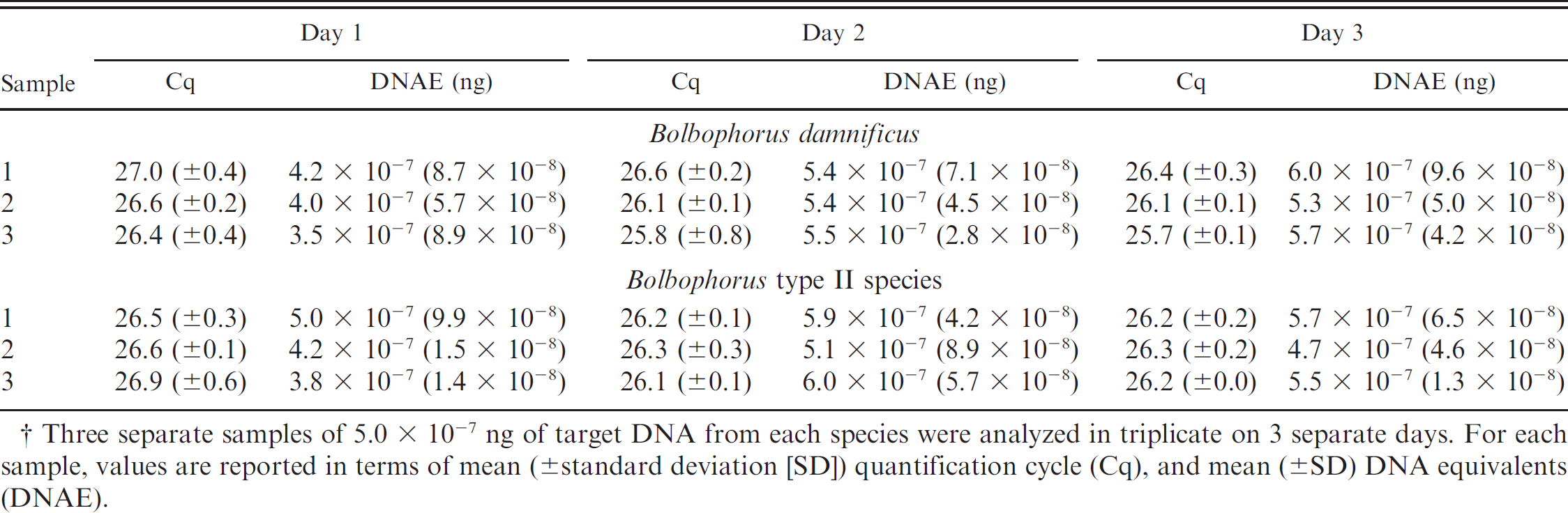
Three separate samples of 5.0 × 10−7 ng of target DNA from each species were analyzed in triplicate on 3 separate days. For each sample, values are reported in terms of mean (±standard deviation [SD]) quantification cycle (Cq), and mean (±SD) DNA equivalents (DNAE).
To determine the ability of the duplex assay to discriminate between and accurately quantify B. damnificus and Bolbophorus type II sp. in the same sample, DNA mixtures composed of combinations of the 2 DNA standards in various quantities (10−3, 10−5, 10−7 ng, with all possible combinations between the 2 species) were artificially generated. Positive controls consisted of target DNA quantities from each species combined with nuclease-free water. All combinations were tested simultaneously by duplex real-time qPCR. The obtained Cq values for each DNA combination were compared with positive controls of target DNA in the absence of nontarget DNA (nuclease-free water) to identify any interference or cross-reactivity between the 2 assays.
The analytical sensitivity of the assay was evaluated using known numbers of cercaria collected from snails determined to be releasing a single Bolbophorus-type by PCR. 4 Cercaria for each species were pooled and enumerated by placing 100 μl of water from the cercaria pool onto a glass counting cell, and the approximate number of cercaria ml−1 was estimated for each species. For each species, 8 aliquots of volumes corresponding to 25 and 125 cercaria were collected and transferred to a 1.5-ml microcentrifuge tube. Eight separate aliquots of 1, 5, and 10 cercaria were collected and placed in a 1.5-ml micro-centrifuge tube using a dissecting microscope and a fine glass pipette. DNA isolation of cercaria aliquots and qPCR analysis were then carried out as described previously.
The linear dynamic range of the assay was established by determining the Cq values for a 10-fold dilution series of the aforementioned DNA standards. Each DNA quantity was analyzed in triplicate on 3 separate days. In addition, the interassay and intra-assay variability were determined by analyzing known amounts of target DNA. Separate preparations of 5 × 10−7 ng (corresponding to approximately 1 cercaria) were analyzed in triplicate on 3 different occasions. The samples were run concurrently with a 10-fold serial dilution of target DNA to evaluate the assay's ability to determine DNA equivalents within each prepared sample.
Based on the fluorescence signals (FAM or HEX) generated during amplification, each primer and probe combination recognized its specific target. The FAM fluorescence was exclusively generated by samples identified by endpoint PCR as B. damnificus, whereas the HEX signals were detected only in samples identified as Bolbophorus type II sp. There was no detectable fluorescence signal generated from the no-template negative controls or the non-bolbophorid cercariae, confirming the assay to be specific for the detection of and discrimination between B. damnificus and Bolbophorus type II sp.
Samples spiked with combinations of low (10−3 ng), intermediate (10−5 ng), and high (10−7 ng) quantities of synthesized DNA of B. damnificus, and Bolbophorus type II sp. showed no remarkable interference during detection and quantification of the 2 Bolbophorus spp. standards in the same sample. Even samples spiked with high quantities of B. damnificus DNA and low quantities of Bolbophorus type II sp. DNA, and vice versa, showed no significant differences in Cq from the controls (analysis of variance [ANOVA]; P > 0.05; data not shown). g Evaluation of serial dilutions of generated DNA standards demonstrated both assays to be linear over 7 orders of magnitude (5 × 10−3 to 5 × 10−9 ng) with a mean slope of −3.37 for the B. damnificus assay and −3.46 for the Bolbophorus type II assay. These slopes represented reaction efficiencies of approximately 98% and 95%, respectively (Fig. 1). 1,12 In addition, both assays were also found to be highly repeatable and reproducible (Fig. 2; Table 2). Analysis of known numbers of cercaria also showed the assay to be sensitive enough to detect a single cercaria of each species (Table 3; Fig. 3). The cutoff Cq value of the assay is 35 cycles, corresponding to approximately 1/256th of a cercaria.
Mean representative quantification cycle (Cq) values (±standard error of mean [SEM]) for aliquots (n = 8) of known numbers of Bolbophorus damnificus and Bolbophorus type II sp. cercariae determined by duplex real-time quantitative polymerase chain reaction.
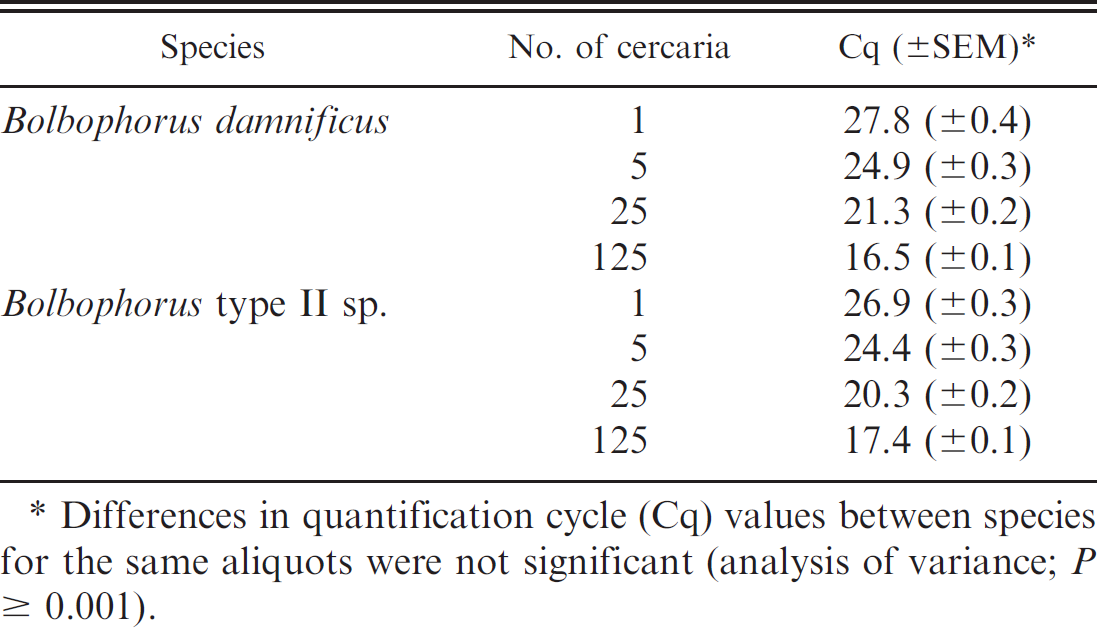
Differences in quantification cycle (Cq) values between species for the same aliquots were not significant (analysis of variance; P ≤ 0.001).
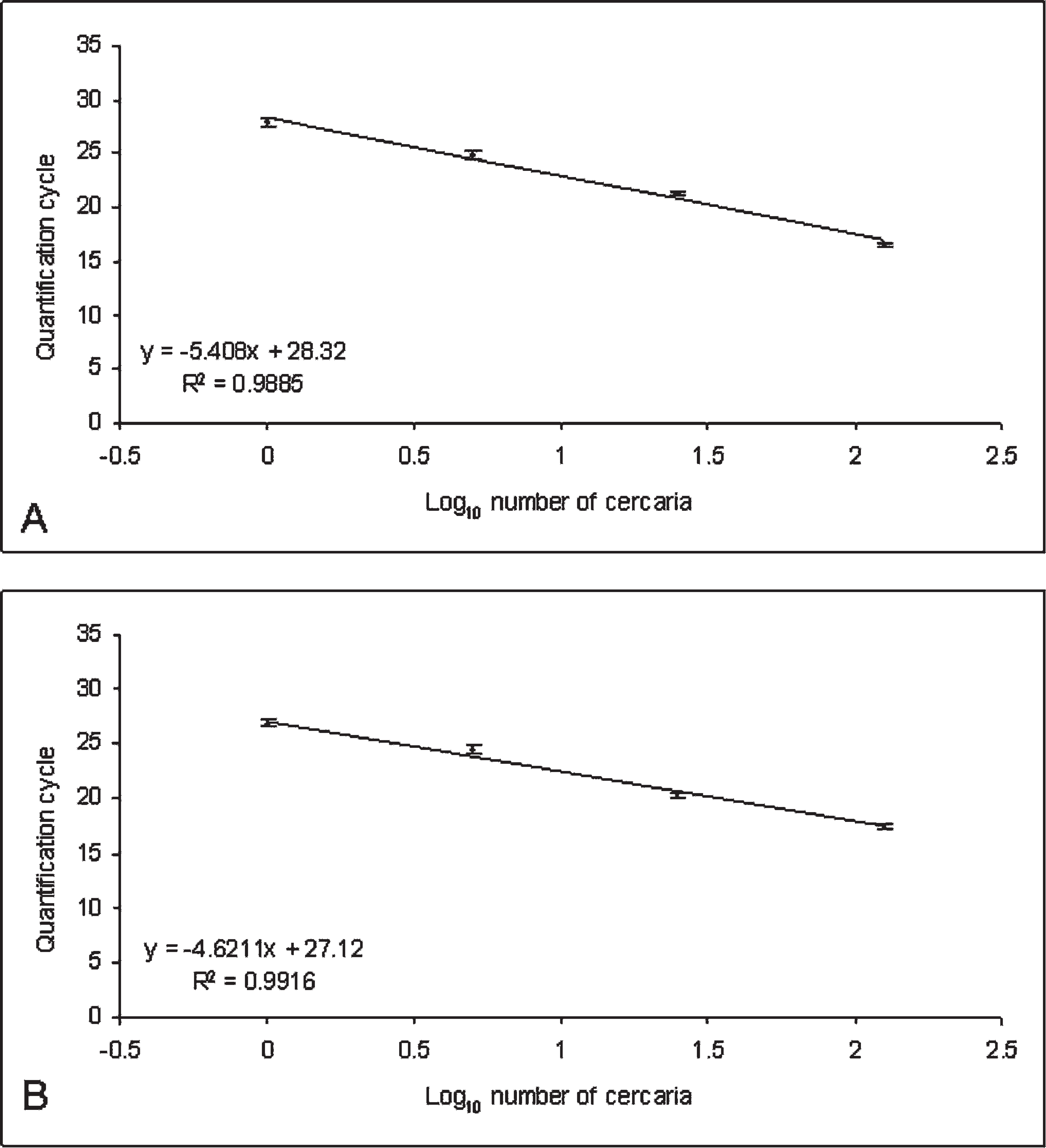
Standard curves generated from known quantities of Bolbophorus damnificus (
As a consequence of PCR chemistries, the more target DNA in the reaction, the faster the fluorescence reaches the predetermined threshold, resulting in an inverse relationship between Cq and template quantity. In other words, the lower the Cq, the greater the amount of initial target DNA in the reaction. Although Cq values for Bolbophorus type II sp. were slightly lower than those for B. damnificus cercaria, these differences were not significant (ANOVA; P ≤ 0.001) g and are likely an artifact of the DNA isolation procedure, pipette error, or variability in the actual number of cercaria added to each aliquot, and they do not necessarily represent differences in the target gene between species. The results for parasite identification by duplex real-time qPCR were in complete agreement with the duplex endpoint PCR assay, 4 confirming the assay described herein to be a reliable means of parasite identification.
To obtain adequate numbers of viable B. damnificus cercariae for disease challenge, large numbers of actively shedding snails are required. A typical challenge requires the collection of upwards of 1,000 snails, of which 5–20% may be shedding 1 or both Bolbophorus-type cercariae. In addition, the cercaria released from these snails must be identified in a relatively short period of time, while the snails are still actively releasing cercariae. Conventional PCR requires postreaction processing, which is laborious and time-consuming, especially when dealing with the large numbers of snails described above. Conversely, real-time PCR performed in 96-well format, excluding 2 wells for controls, can qualitatively identify cercaria types from 94 snails in a single run, with results available immediately following the completion of the reaction, thus expediting the process of identifying snails shedding only B. damnificus cercariae, increasing the numbers of snails that can be evaluated in short periods of time.
In addition, the quantitative nature of the assay suggests this protocol could be employed in a similar fashion to other established protocols for the detection and quantification of pathogens from channel catfish ponds. 3 The insidious nature of trematode infections on pond-raised channel catfish argues strongly for a planned formal program to continually assess disease prevalence, as even light infestations can result in significant economic losses. 10 Currently practiced control measures involve the chemical eradication of the snail host from infested ponds. 5,6,10,11 Unfortunately, nonspecific prophylactic use of chemicals to control the parasite at the farm level is cost prohibitive. As such, formal disease surveillance and monitoring programs are necessary to identify ponds requiring treatment because low-grade infections are difficult to detect by casual observation. To be cost-effective, chemotherapeutants can be applied only to ponds with snail populations actively shedding the infective cercarial stage. Current recommendations for identifying ponds requiring treatment call for the physical examination of 30–40 catfish from each pond, where infestations are verified by the presence of metacercarial cysts presenting as raised nodules just under the skin. 10 This surveillance methodology proves cumbersome, labor intensive, and time-consuming and ultimately does not determine the numbers of cercaria in the pond, which are the indirect target of control. The development of a molecular-based assay to directly detect and quantify the free-swimming cercaria in pond water could potentially overcome the labor-related limitations associated with current disease-monitoring programs.
The duplex assay described in the current study is a reliable, more rapid means of parasite identification than conventional endpoint PCR. In addition, the ability to quantify cercaria from pond water samples, similar to protocols used for other catfish pathogens, 3 could prove invaluable in determining the impact of chemical treatments on cercaria concentrations as well as serve an essential role in the development of treatment application schedules designed to maximize treatment efficiency. Future research will focus on the integration of this assay into protocols currently in place for pathogen detection and the validation of its use in on-farm pond disease monitoring and surveillance applications.
Acknowledgements
The authors thank Josh Walker, Monroe Walker, and Todd Byars for help in snail collection. This research was funded through the Mississippi Agricultural and Forestry Experiment Station (MAFES) Strategic Research Initiative program and was supported by the College of Veterinary Medicine, Mississippi State University. This is MAFES publication J-11736.
Footnotes
a.
Powersoil DNA Isolation Kit, Mo Bio Laboratories Inc., Carlsbad, CA.
b.
TaqMan Environmental Mastermix 2.0, Applied Biosystems, Foster City, CA.
c.
MJ Research PTC-200 Gradient Cycler, GMI, Ramsey, MN.
d.
QIAquick PCR purification kit, QIAGEN Inc., Valencia, CA.
e.
NanoDrop spectrophotometer and software version 3.2.1, NanoDrop Technologies, Wilmington, DE.
f.
Bio-Rad iCycler version 3.1, Bio-Rad Laboratories, Hercules, CA.
g.
SAS software version 9.2, SAS Institute, Cary, NC.
