Abstract
The current study was carried out to set up a fast and specific technique for porcine tuberculosis diagnosis in formalin-fixed, paraffin-embedded tissues. A retrospective study was carried out using 54 samples fixed in 10% neutral buffered formalin from 29 slaughtered Iberian pigs. Most of the pigs showed tissue samples positive to immunohistochemical staining (70.4%), and mycobacteria were detected within or near the necrotic cores of the lesions. However, diagnosis by this technique was time-consuming and tedious because of the paucibacillar nature of porcine tuberculous lesions. Classic polymerase chain reaction (PCR) was unsuccessful in mycobacteria genome amplification in all of the examined samples; however, real-time PCR amplified the mycobacteria genome in 23 of 29 examined pigs, identifying the Mycobacterium tuberculosis complex in all but one, which amplified Mycobacterium avium complex. Moreover, when reamplification of the DNA was performed, classic PCR amplified the mycobacteria genome in all the examined pigs (29/29), identifying the M. tuberculosis complex in 28 of 29 studied pigs and M. avium complex in only 1 pig. Results of the current study point out that both real-time and classic PCR assays, with genome reamplification, represent sensitive, fast, and specific diagnostic tools for porcine tuberculosis in formalin-fixed, paraffin-embedded tissues.
Purulent and/or chronic caseous lymphadenitis are caused by several microorganisms triggering gross lesions in the lymph nodes, several organs, and carcasses. The multiple microorganisms, which may be involved in these lesions, make it hard to identify the etiologic agent by visual inspection at the slaughterhouse. 18 Tuberculosis is a chronic granulomatous infectious disease caused by acid-fast bacilli from the Mycobacterium genus, which has 2 large differentiated complexes: M. tuberculosis complex and M. avium complex. Mycobacterium tuberculosis complex includes, among others, M. tuberculosis, M. bovis, M. africanum, and M. microti, whereas M. avium complex includes M. avium subsp. avium, M. avium subsp. hominissuis, M. avium subsp. paratuberculosis, and M. intracellulare. 9 Mycobacterium bovis has been reported as the main etiologic agent of porcine tuberculosis. 4,12,14 However, tuberculosis in commercial crossbred pigs is rare because of the tight control in the intensive breeding systems, which has led to an increase in the incidence of M. avium complex and atypical mycobacteria 2 in several European countries. 12,16 Iberian pigs are bred in extensive or semi-extensive breeding systems and share habitat with other domestic and wild animals, which may increase the risk of mycobacterial infection by M. bovis in pigs. 12
Wild animals represent a reservoir of mycobacteria, which facilitates the persistence and spread of mycobacteria in a specific geographic area. 1,14 The transmission of different M. bovis isolates between wild animals species and between wild and domestic animals has been previously reported. 4,14 Indeed, cyclic infections may appear between domestic and wildlife animals. 4
The initial lesions observed in porcine tuberculosis are located mainly in the mandibular lymph nodes, and it is possible to find lesions in lung, liver, or spleen when there is lymphohematic dissemination. 8,11,12 However, several microorganisms, such as Rhodococcus equi or Arcanobacterium pyogenes, may induce similar lesions, making it necessary to identify the causative agent in order to make an accurate diagnosis. 18 Several diagnostic tests have been developed for porcine tuberculosis diagnosis. Specific culture media may be considered the gold standard technique making it feasible to differentiate between mycobacteria species; nonetheless, this technique requires fresh tissues and an extended period for mycobacterium isolation. Other techniques allow for the detection of mycobacteria in fixed-tissue sections, such as Ziehl–Neelsen staining (ZN) or immunohistochemistry; however, it is not always easy to identify them with these techniques. 13,17 Molecular techniques represent a fast method for diagnosis, showing higher sensitivity and specificity, 1,2,4,10,12,13,17 although their efficiency is rather limited when formalin-fixed samples are used. 6,15,20 Thus, the aim of the current study was to develop an objective, sensitive, and specific technique for porcine tuberculosis diagnosis in formalin-fixed, paraffin-embedded (FFPE) tissues, which could also differentiate between M. tuberculosis and M. avium complexes.
A retrospective study was carried out using 54 tissue samples fixed in 10% neutral buffered formalin for 2–5 days. The samples corresponded to lymph node, tonsil, lung, and/or liver taken from 29 Iberian pigs that were slaughtered because of tuberculosis-like lesions. Formalin-fixed samples were embedded in paraffin wax, and 4-μm sections were stained with hematoxylin and eosin for the histopathologic study and with ZN for the detection of acid-fast bacilli. Immunolabeling for mycobacterium antigen was carried out in tissue sections using the polyclonal antibody M. bovis BCG. a The cross-reaction of this antibody against mycobacteria that belonged to M. avium complex was confirmed using sections from positive avian tuberculosis cases. Avidin–biotin–peroxidase complex was used for the immunohistochemical technique. Briefly, antigen retrieval was performed using a 0.07% protease b solution in phosphate buffered saline for 10 min in a stirrer. The primary antibody, diluted 1 in 750, was incubated overnight at 4°C in a humid chamber. A biotinylated goat anti-rabbit immunoglobulin G secondary antibody, c diluted 1 in 200, was incubated at room temperature for 30 min. Negative controls for the technique consisted of the primary antibody replacement by normal goat serum or isotype negative controls.
DNA for polymerase chain reaction (PCR) was extracted from three 20-μm sections from FFPE tissue waxes. The tissue sections were collected into a 1.5-ml sterile DNAsefree microcentrifuge tube and pelleted by centrifugation at 4,000 rcf for 5 min. Blades were changed between each sample to avoid cross contamination. Next, 200 μl of Tween 20 d diluted to 0.5% was added to each tube with a tissue sample, which were subjected to 3 consecutive cycles of heating for 10 min at 100°C followed by liquid nitrogen immersion for 5 min. Tubes were centrifuged at 2,200 rcf for 20 min. The paraffin ring formed after centrifugation was removed, and 60 μl of the supernatant was taken and frozen at −80°C until PCR reaction was complete. DNA extraction was carried out using a commercial PCR template preparation kit. e Polymerase chain reaction detection of DNA sequences of M. tuberculosis and M. avium complexes was performed as previously described. 3 Oligonucleotide primers myc1 and myc3 simultaneously amplified the M. tuberculosis complex (M. bovis and M. tuberculosis) and the M. avium complex (M. avium and M. avium subsp. paratuberculosis) producing different amplicon sizes of 178 bp and 257 bp, respectively. Primer sequences were: myc1: 5'-GAGTAGGTCATGGCTCC TCC-3′ and myc3: 5′-CATGCACCGAATT AGAACGT-3′. Classic PCR reactions were performed in a 50-μl volume containing 5 μl of template, 10 pmol of each primer, 0.25 mM of each deoxynucleoside triphosphates, and 0.625 U of Taq DNA polymerase. Reactions were run in a thermocycler f under the following conditions: initial denaturation at 95°C for 10 min, 45 cycles of amplification consisting of denaturation at 95°C for 30 sec, primer annealing at 58°C for 45 sec, and extension at 72°C for 30 sec, with an increase of 1 sec per cycle for the denaturation and extension steps. All reactions were run in duplicate. To check the specificity of the amplified products, M. tuberculosis complex–positive and M. avium complex–positive samples were included in each assay. Cross-contamination was ruled out by including negative controls (PCR analysis without DNA template). Mycobacteria isolates typified as M. bovis (2003, 2005, 2012 and 10/0 88 isolates) or M. avium (AA isolate), and positive controls for M. bovis and M. avium from FFPE samples diagnosed histopathologically, were used as positive controls for the validation of the technique. Because classic PCR resulted in no mycobacteria genome amplification from any of the tested samples, both real-time and classic PCR assays with reamplified genome were carried out to avoid false-negative results caused by low DNA template. Reamplification of the DNA was carried out to improve the threshold of the classic PCR, as previously described. 5 The secondary PCR was identical to the classic PCR described previously, but in the current case, 1 μl from the product obtained in the classic PCR was used as template.
Real-time PCR was performed to assess an alternative, faster molecular technique than classic PCR. Real-time PCR reactions were performed using an SYBR Green master mix g in 50-μl reaction volumes containing 5 μl of template and 10 pmol of each primer. Plus–minus reactions were run in a thermocycler under the same conditions described previously for classic PCR. To check the specificity of the amplified products, dissociation curve analysis was performed (Fig. 1). Mycobacterium tuberculosis complex–positive and M. avium complex–positive samples were differentiated based on the melting temperatures for each amplicon, 87.8 ± 0.4°C and 89.6 ± 0.2°C, respectively. The technique was validated using the same positive controls described earlier. The coefficient agreement between the different techniques was assessed by means the Kappa coefficient.
Granulomatous or pyogranulomatous lesions were identified in the 54 tissue samples examined, and ZN showed acid-fast bacilli in most of the tissue samples examined. In 70.4% of tissue samples (28/29 pigs), the immunolabeled mycobacteria were observed within the cytoplasm of epithelioid cells and giant cells (Figs. 2, 3), as well as in the necrotic core of the lesion and their surroundings. Classic PCR was unable to amplify the DNA of mycobacteria in any of the pigs studied. Therefore, both real-time and classic PCR assays were performed subsequent to reamplification of the DNA. Using real-time PCR, the genome of M. tuberculosis complex was amplified in 22 of 29 pigs, and the DNA belonging to M. avium complex was amplified in 1 pig. No fragment was amplified from the samples obtained from the 6 remaining pigs (Table 1). Reamplification of the DNA allowed amplifying a 178-bp amplicon belonging to the M. tuberculosis complex in 28 of 29 pigs, whereas a 257-bp amplicon corresponding to M. avium complex–specific DNA was amplified in 1 pig (Table 1).
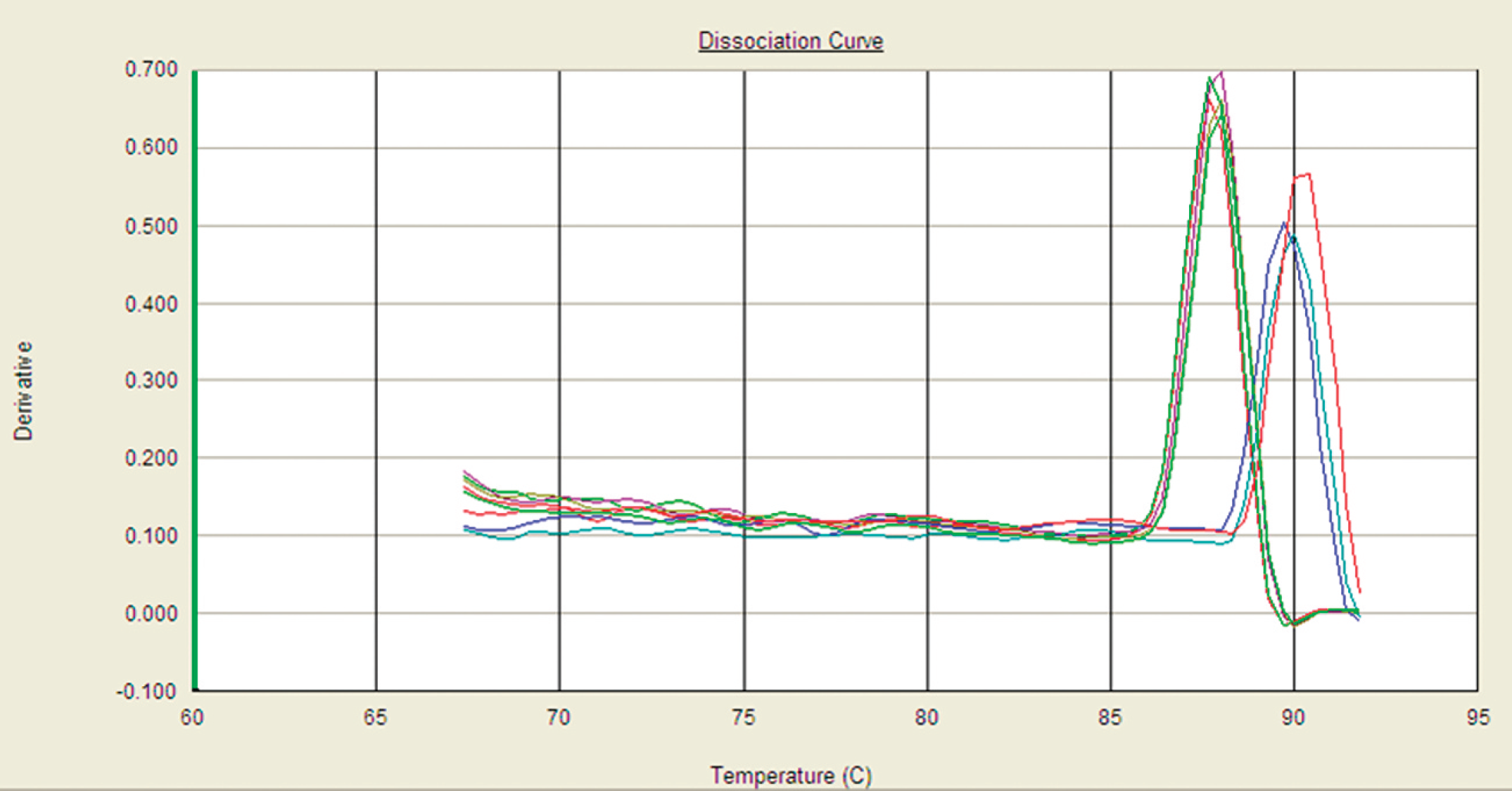
Dissociation curve analysis for differentiation of Mycobacterium tuberculosis complex– and Mycobacterium avium complex–positive samples based on the melting temperatures for each amplicon.
Because the current study is a retrospective study using FFPE tissue samples, it was not possible to isolate mycobacteria by means of specific culture media, which is considered the gold standard. Although most of the tissue samples showed positive immunolabeling for mycobacterial antigen in the present study, the diagnosis of mycobacteria by immunohistochemistry was still far from being easy and fast. Although a high level of efficiency as achieved with immunohistochemistry, the subjectivity of the pathologist and the paucibacillar character of porcine tuberculosis constitute important disadvantages of this diagnostic tool. Nonetheless, in the current study, immunolabeling of mycobacteria antigen displayed an almost perfect agreement with respect to both real-time (κ = 0.83) and classic (κ = 0.97) PCR with reamplification of the DNA.
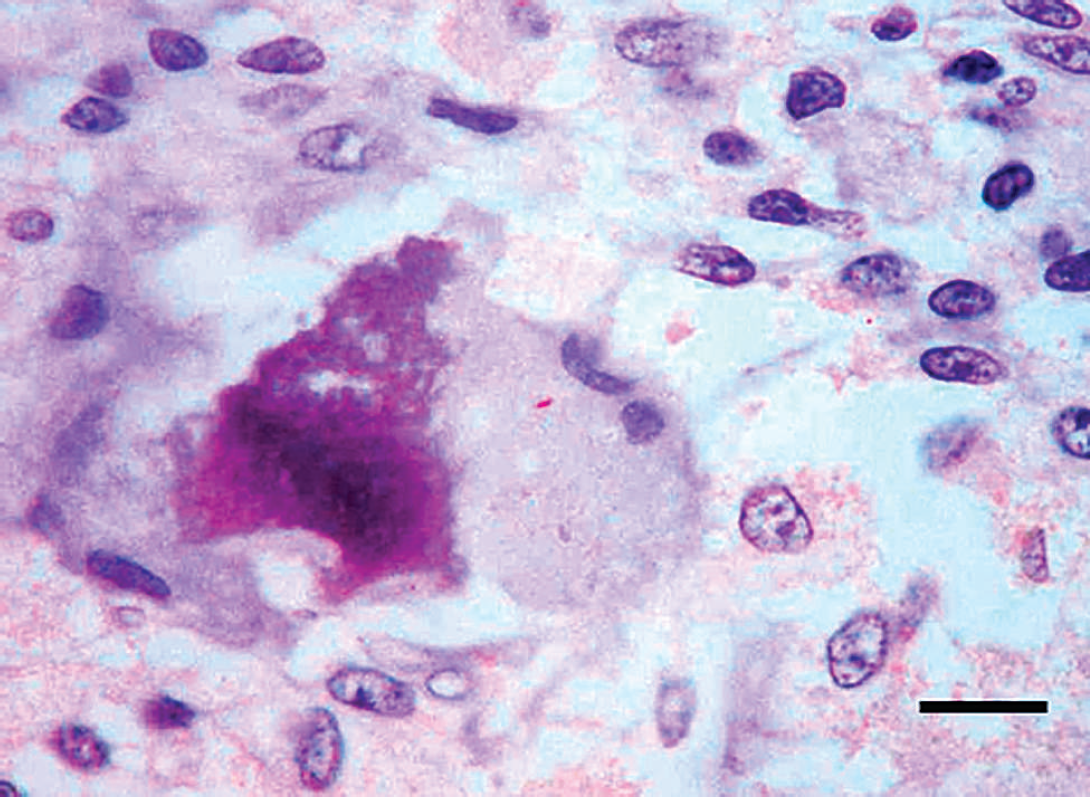
Lymph node. Immunolabeled mycobacteria within the cytoplasm of a Langhans giant cell. Avidin–biotin–peroxidase complex. Mayer's hematoxylin counterstaining. Bar = 5 μm.
Real-time PCR resulted in a more sensitive diagnostic tool than classic PCR, because DNA reamplification was required to amplify the mycobacteria genome from FFPE tissues by means of classic PCR. Several factors may explain such differences, such as insufficient DNA template, which is a cause for false-negative results in PCR reactions. 19 Porcine tuberculosis induces paucibacillar lesions, which means that the DNA template per section is very low. 7 In addition, formalin fixation has some negative consequences for DNA amplification. The DNA present in the sample may be disrupted by formalin fixation, reducing the detection threshold of the PCR. Several authors 15,20 have concluded that formalin fixation limits the detection of DNA fragments that are more than 225 bp, and examination of smaller fragments is more accurate. Fixation time is also significant as genome amplification decreases after 24 hr of fixation. 6,20 Despite the inconvenience of formalin fixation, real-time PCR was the most successful technique in the current study, because classic PCR required DNA reamplification for successful amplification of fragments belonging to the Mycobacterium genus. The results obtained from both real-time PCR and classic PCR with DNA reamplification showed a substantial agreement (κ = 0.79) by means of Kappa statistics.
In the current study, 96.5% of pigs were found to be positive for M. tuberculosis complex compared with 3.5% of pigs found to be positive for M. avium complex. Similar results have been previously reported in isolating M. bovis in 37 wild boars and 25 Iberian pigs suffering from tuberculosis. 12 The detection of only one pig infected by M. avium complex in the present study might be due to the involvement of M. avium subsp. hominissuis, an environmentally ubiquitous mycobacteria, which is nonpathogenic for birds but an opportunistic pathogen for other animals, such as the pig. 10 On the other hand, the habitat of the Iberian pig makes contact easier between it and other domestic or wild species, such as cattle or cervids, which are commonly infected by M. bovis in tuberculosis cases. 11 Moreover, because of the participation of different mycobacteria species in porcine tuberculosis, specific and fast diagnostic tools are required to determine the Mycobacterium complexes involved in each case. Although in the current study, M. tuberculosis complex was the agent responsible for most of the examined pigs, attention should be paid to emerging cases associated with M. avium complex infection. This aspect is especially significant because different serovars from M. avium isolated from immunosuppressed patients are also isolated from tuberculous lesions in pigs. 18
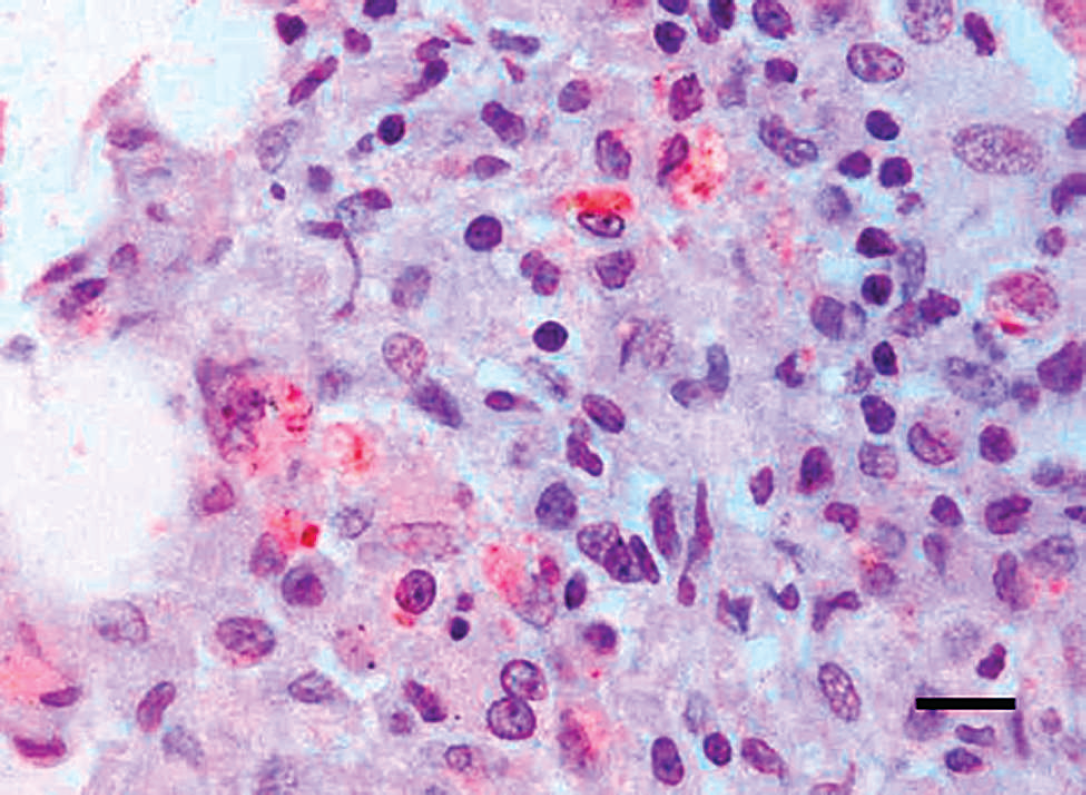
Lung. Immunohistochemical staining of mycobacteria antigen within the cytoplasm of macrophages infiltrating the alveolar septa of the lung of a slaughtered pig. Avidin–biotin–peroxidase complex. Mayer's hematoxylin counterstaining. Bar = 5 μm.
The current study revealed that real-time PCR represents the fastest and most accurate technique for detecting Mycobacterium complexes in FFPE tissues. This approach may help to encourage further retrospective studies to evaluate changes in mycobacterial species in different regions, which might determine the significance of each mycobacteria species in tuberculosis outbreaks.
Immunohistochemistry (IHC) and polymerase chain reaction (PCR) results observed in each pig. *
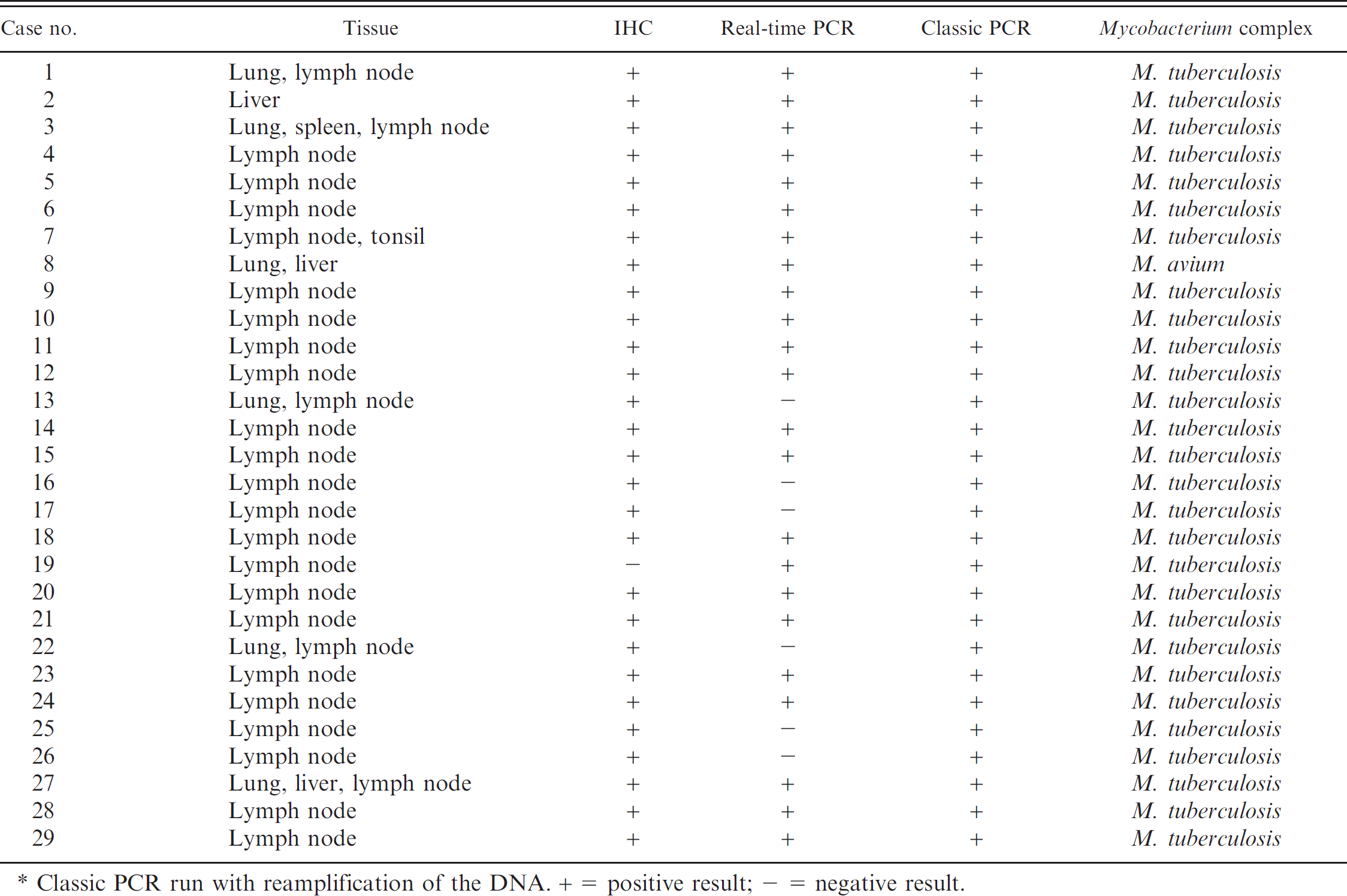
Classic PCR run with reamplification of the DNA. + = Positive result; — = negative result.
Acknowledgements. The authors would like to thank Dr. A. Parra for his kind gift of mycobacterial isolates 2003, 2005, 2012, 10/0 88, and AA; and G. Muñoz for her technical assistance. J. Gómez-Laguna and J. J. Quereda were supported by a doctoral grant from the Spanish Ministry of Education and Science (AP-2004-0395 and AP-2005-3468, respectively).
Footnotes
a.
Polyclonal Mycobacterium bovis BCG antibody, Dako Denmark A/S, Glostrup, Denmark.
b.
Protease from Bacillus licheniformis, Sigma-Aldrich Inc., Burlingame, St. Louis, MO.
c.
Biotinylated goat anti-rabbit IgG secondary antibody, Vector Laboratories Inc., Burlingame, CA.
d.
Tween 20, Sigma-Aldrich, St. Louis, MO.
e.
High Pure PCR Template Preparation Kit, Roche Diagnostics GMBH, Mannheim, Germany.
f.
Thermocycler ABI 7300, Applied Biosystems, Foster City, CA.
g.
Power SYBR Green Master Mix, Applied Biosystems, Foster City, CA.
