Abstract
Real-time reverse transcription loop–mediated isothermal amplification (real-time RT-LAMP) holds substantial potential as a highly sensitive, specific, and easy-to-perform molecular technique for pathogen detection in clinical samples. In the current study, the analytical and diagnostic performance of 2 commercial realtime RT-LAMP kits, Avian Flu H5 and Avian Flu H7, in detecting Avian influenza virus (AIV) infections were evaluated and compared with validated real-time reverse transcription polymerase chain reaction (RT-PCR) assays using RNA from reference virus isolates of subtypes H5 (n = 24) and H7 (n = 25) and of phylogenetically related subtypes (n = 20). When real-time RT-LAMP was carried out according to the recommendations of the manufacturer, 3 out of 24 H5 isolates and 8 out of 25 H7 reference strains were not detected. Prolonging the amplification phase resulted in detection of all H5 isolates but also in false positive detection of 2 non-H5 isolates. Real-time RT-LAMP specific to H7 failed to detect 2 H7 isolates after prolonged amplification. According to the examination of RNA log dilutions, the sensitivity of the real-time RT-LAMP assays, for a number of historic but also recent strains, was considerably lower compared with subtype-specific real-time RT-PCR assays. Application of the real-time RT-LAMP assays for analysis of diagnostic samples from wild birds confirmed their lower sensitivity. Commercial real-time RT-LAMP as tested in this study with a broad range of AIV H5 and H7 strains of phylogenetically diverse yet recent origin, holds some promise for routine veterinary diagnostic purposes, although real-time RT-LAMP was markedly more vulnerable to a reduction of detection limits because of strain-specific sequence variation than subtype-specific real-time RT-PCR.
Keywords
Highly pathogenic Avian influenza virus (HPAIV) can cause severe economic losses in poultry production and could also threaten human health. The currently panzootically circulating HPAIV subtype H5N1 of Asian origin is considered a candidate for another new human pandemic. During recent decades, HPAIV outbreaks in poultry and human AIV infections have increased in expansion and frequency, respectively, and have prompted intensified AIV surveillance projects in both wild birds and poultry. Molecular diagnostics of AIV was revolutionized by the introduction of real-time reverse transcription polymerase chain reaction (real-time RT-PCR) since 2000 and is now firmly implemented in routine diagnostics. 1 Recent developments, like low-density microarray analysis or combined rolling-circle and PCR amplification using padlock primers, are powerful tools to further speed up and expand AIV diagnostic possibilities where fully equipped laboratories are available. 3,4,12 Given, however, that the majority of outbreaks of currently circulating HPAIV H5N1 occurs in regions where such laboratories are only infrequently encountered, the need is urgent to implement highly specific and sensitive rapid molecular tests for detection of AIV in less well equipped laboratories. Such assays might also be used directly in the field as so-called pen site tests.
Because of its simplicity, the principle of loop-mediated isothermal amplification (LAMP) seems to be a promising technology to develop such assays for AIV diagnosis. 10,11 Basically, the minimal technical requirement is a water bath or a thermoblock to incubate the reaction mix. 10 The reportedly high specificity of the LAMP reaction is based on the recognition of target sequences by 2 pairs of primers, whereas the sensitivity is driven by 1 additional pair of loop primers for acceleration of the reaction. 9 Special requirements for LAMP primer design, however, hamper the development of diagnostic assays when conserved regions in the target gene are scarce. The extremely high variability of the AIV hemagglutinin (HA) gene as a target for HA subtype–specific diagnostics poses a particularly difficult case not only for the design of primers for conventional PCR, but even more for appropriate LAMP primer sets. 5,12,13
The need for new rapid AIV assays that can be carried out with a minimum of specific equipment prompted the present evaluation of the performance characteristics of a commercial real-time RT-LAMP amplification kit a and the corresponding primer kits a designed to detect RNA of the notifiable H5- and H7-AIV subtypes. Until then, no data were available concerning performance characteristics of the H7-specific real-time RT-LAMP, whereas H5-specific kits had been evaluated with some selected virus isolates of the HPAIV H5N1 and were found to have a matching, if not superior, performance when compared with conventional RT-PCR. 5,6 However, real-time RT-PCR is considerably more sensitive than conventional RT-PCR and now represents the gold standard for AIV RNA detection, 12 and no data on comparisons between RT-LAMP and subtype-specific real-time RT-PCR are available.
For a basic evaluation of the real-time RT-LAMP assays, RNA extracted manually b from established reference virus isolates of the H5 and H7 subtypes, as well as RNA of phylogenetically closely related subtypes, namely H1, H2, H6 (related to H5), and H10 (related to H7), were used. The analytical sensitivity of the Avian Flu H5 a primer kit was analyzed by using 24 virus isolates of the H5 subtype comprising the neuraminidase (NA) subtypes N1 (n = 12), N2 (n = 8), N3 (n = 2), N6 (n = 1), and N9 (n = 1) belonging to biotypes of high (HPAIV) and low (LPAIV) pathogenicity (Table 1). Furthermore, 12 isolates of the HA subtypes H1, H2, and H6 were analyzed to challenge the specificity of the H5-specific primers (Table 1). The Avian Flu H7 a primer kit was tested by examination of 25 isolates of the H7 subtype and 8 isolates belonging to the closely related H10 subtype (Table 2).
Diagnostic performance was assessed with RNA preparations of samples obtained from wild birds in Germany in 2007 (Table 3). RT-LAMP was performed a with the use of H5- and H7-specific primer kits Avian Flu H5 and Avian Flu H7, a respectively. Amplification was performed with an amplification kit a at 62.5°C for 35 min as recommended by the manufacturer or extended to 60 min in an attempt to increase sensitivity, followed by an inactivation (80°C, 5 min). Sensitivity of the real-time RT-LAMP test was compared with established real-time RT-PCR assays by examination of log dilutions of RNA from a selection of egg-propagated reference strains (Tables 1, 2) with the use of a 1-step real-time RT-PCR c employing DNA polymerase c using PCR primers specific for fragments of the matrix (M), H5, or H7 genes as described previously. 14,15
The H5-specific real-time RT-LAMP when run with recommended settings failed to detect 3 of 24 H5 isolates (Table 1). Suspensions of the isolates A/ck/Scotland/59 (HPAIV H5N1), A/teal/Germany/Wv632/05 (LPAI H5N1), and A/duck/Potsdam/1701/86 (LPAIV H5N2), which tested false negative in the real-time RT-LAMP, were clearly positive by H5-specific real-time RT-PCR with threshold cycle (Ct) values of 20.7, 26.5, and 26.7, respectively, indicating that ample amounts of specific RNA were present. An extension to 60 min of the original amplification time of 35 min resulted in a detection of these 3 isolates after 37 min (A/teal/Germany/Wv632/05), 51 min (A/ck/Scotland/59), or 52 min (A/duck/Potsdam/1701/86). However, the extended amplification time when applied to detection of isolates of related subtypes resulted in false-positive signals for 2 non-H5 isolates (A/turkey/Grub/R41/98 [H6N5] and A/mallard/Germany/Wv672/04 [H2N3]) after 50 min and 59 min, respectively (Table 1). These isolates were not detected when using the recommended amplification time.
Eight out of 25 RNA samples from different H7-specific AIV isolates were not detected by the real-time RT-LAMP assay after 35 min of amplification, although all 8 samples tested positive in the H7-specific real-time RT-PCR with Ct values between 24 and 33. A prolongation of the amplification time to 60 min resulted in detection of 6 more samples after 36, 37, 47, 50, 51, and 57 min, so that finally, only the isolates A/FPV/Rostock/45/34 (H7N1) and A/dk/Alberta/48/76 (H7N3) stayed negative. In real-time RT-PCR, Ct values of 33.2 and 25.1, respectively, were obtained for these 2 isolates. Eight further AIV strains belonging to the H10 subtype were not detected by the H7-specific real-time RT-LAMP, even when applying the prolonged amplification protocol (Table 2).
In addition, all RNA samples that tested specifically positive in the real-time RT-LAMP were also assayed after preparing log dilutions to assess the sensitivity of real-time RT-LAMP compared with real-time RT-PCR. The highest dilution step that was positive in real-time RT-LAMP had a mean Ct value of 28.6 for the Avian Flu H5 primer kit and 26.7 for the Avian Flu H7 primer kit when using the recommended amplification protocol of 35 min. In realtime RT-PCR assays, mean Ct values of the highest positive dilutions were 37.1 for both H5- and H7-specific real-time RT-PCR assays. Taking into account that a difference of 3.3 Ct values in real-time RT-PCR amounts to approximately 1 log step of target copies, the real-time RT-LAMP in comparison to real-time RT-PCR must be considered approximately 1,000-fold less sensitive.
To test the diagnostic performance of the H5- and H7-specific real-time RT-LAMP assays, RNA isolated from diagnostic swab samples of wild birds and poultry from Germany were analyzed by real-time RT-LAMP and realtime RT-PCR. The samples comprised 21 H5 positives, 6 H7 positives, 5 non–H5- and non–H7-AIV positives, and 2 samples that were positive for Newcastle disease virus (NDV; Table 3). The H5-specific real-time RT-LAMP detected 4 out of 21 H5 real-time RT-PCR–positive samples. These 4 samples had Ct values below 30. Performing the H7-specific real-time RT-LAMP, 4 real-time RT-PCR–positive samples were also considered positive by real-time RT-LAMP (after 13–16 min run time), but 2 H7 swab samples with low viral RNA loads were not detected by real-time RT-LAMP (Table 3). One of the H5-negative samples, identified to harbor an H3N8 isolate, was false positive by real-time RT-LAMP. Four further AIV RNA–positive diagnostic samples that tested negative for H5- and H7-RNA in the real-time RT-PCR and 2 samples harboring RNA of NDV were correctly identified as negative in the subtype-specific realtime RT-LAMP by both primer sets.
In summary, the mean detection limit of the real-time RT-LAMP kits investigated in the current report was approximately 1,000-fold lower than that of optimized subtype-specific real-time RT-PCR assays. Concerning sensitivities of AIV subtype–specific LAMP approaches, only a few reports on H5 and 1 report on H9 have been published so far, but these focus more on the suitability for detection of a selection of very recent strains rather than a wide variety of distinct isolates. 2,5,8 Surprisingly, no data is available for the diagnosis of H7 subtype AIV RNA with the use of LAMP-based assays, although H7 AIV infections of poultry are notifiable to the O.I.E (World Organization for Animal Health). All AIV-targeted RT-LAMP assays developed so far have been compared with conventional RT-PCR, and sensitivity was calculated on the basis of infectivity, not taking into account the amount of RNA copies. 2,5,6,8 In this study, sensitivity of real-time RT-LAMP assays was compared with highly sensitive realtime RT-PCR assays, which now represent the gold standard for AIV RNA detection. 12
Detection of H5 subtype–specific Avian influenza virus RNA by H5-specific real-time reverse transcription polymerase chain reaction (real-time RT-PCR) and by Avian Flu H5 real-time reverse transcription loop-mediated isothermal amplification (realtime RT-LAMP).
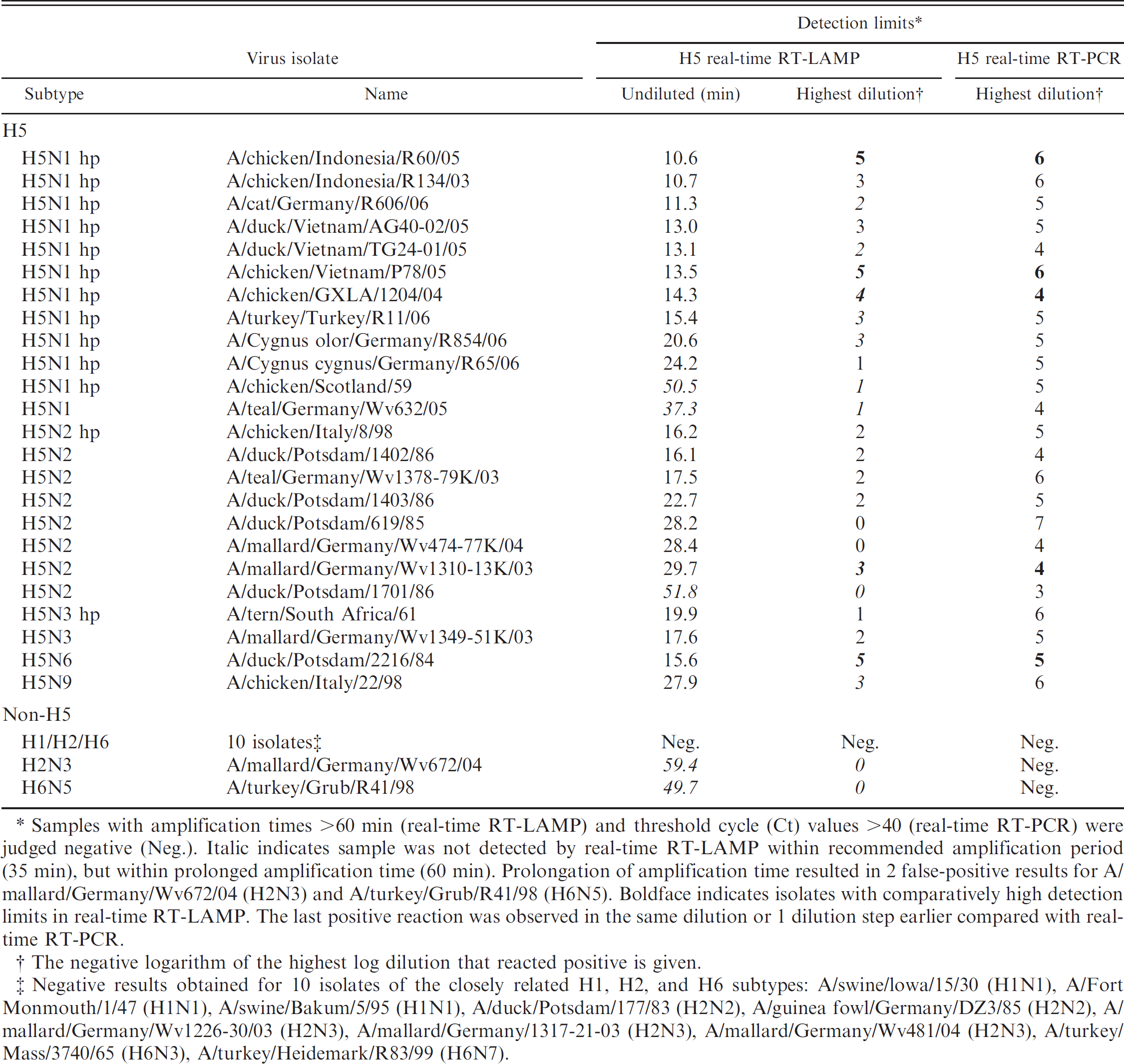
Samples with amplification times >60 min (real-time RT-LAMP) and threshold cycle (Ct) values >40 (real-time RT-PCR) were judged negative (Neg.). Italic indicates sample was not detected by real-time RT-LAMP within recommended amplification period (35 min), but within prolonged amplification time (60 min). Prolongation of amplification time resulted in 2 false-positive results for A/mallard/Germany/Wv672/04 (H2N3) and A/turkey/Grub/R41/98 (H6N5). Boldface indicates isolates with comparatively high detection limits in real-time RT-LAMP. The last positive reaction was observed in the same dilution or 1 dilution step earlier compared with realtime RT-PCR.
The negative logarithm of the highest log dilution that reacted positive is given.
Negative results obtained for 10 isolates of the closely related H1, H2, and H6 subtypes: A/swine/lowa/15/30 (H1N1), A/Fort Monmouth/1/47 (H1N1), A/swine/Bakum/5/95 (H1N1), A/duck/Potsdam/177/83 (H2N2), A/guinea fowl/Germany/DZ3/85 (H2N2), A/mallard/Germany/Wv1226–30/03 (H2N3), A/mallard/Germany/1317–21–03 (H2N3), A/mallard/Germany/Wv481/04 (H2N3), A/turkey/Mass/3740/65 (H6N3), A/turkey/Heidemark/R83/99 (H6N7).
A conventional RT-LAMP assay developed for detection of the nucleoprotein gene of human Influenza B virus and the HA fragments of the Influenza A virus subtypes H1 and H3 was reported to be less sensitive than virus isolation but more sensitive than a commercial immunochromatography test. 7 An AIV H9 subtype–specific RT-LAMP was considered to be approximately 10 times more sensitive than the corresponding conventional RT-PCR and was able to detect clinical samples that were missed by this RT-PCR. 2 All clinical samples originating from a recent AIV outbreak in China and examined with this H9 subtype–specific RT-LAMP had also been virus isolation–positive, which indicates that comparatively high virus loads had been present in these samples. 2
Detection of H7 subtype–specific Avian influenza virus RNA by H7-specific real-time reverse transcription polymerase chain reaction (real-time RT-PCR) and by Avian Flu H7 real-time reverse transcription loop-mediated isothermal amplification (realtime RT-LAMP).
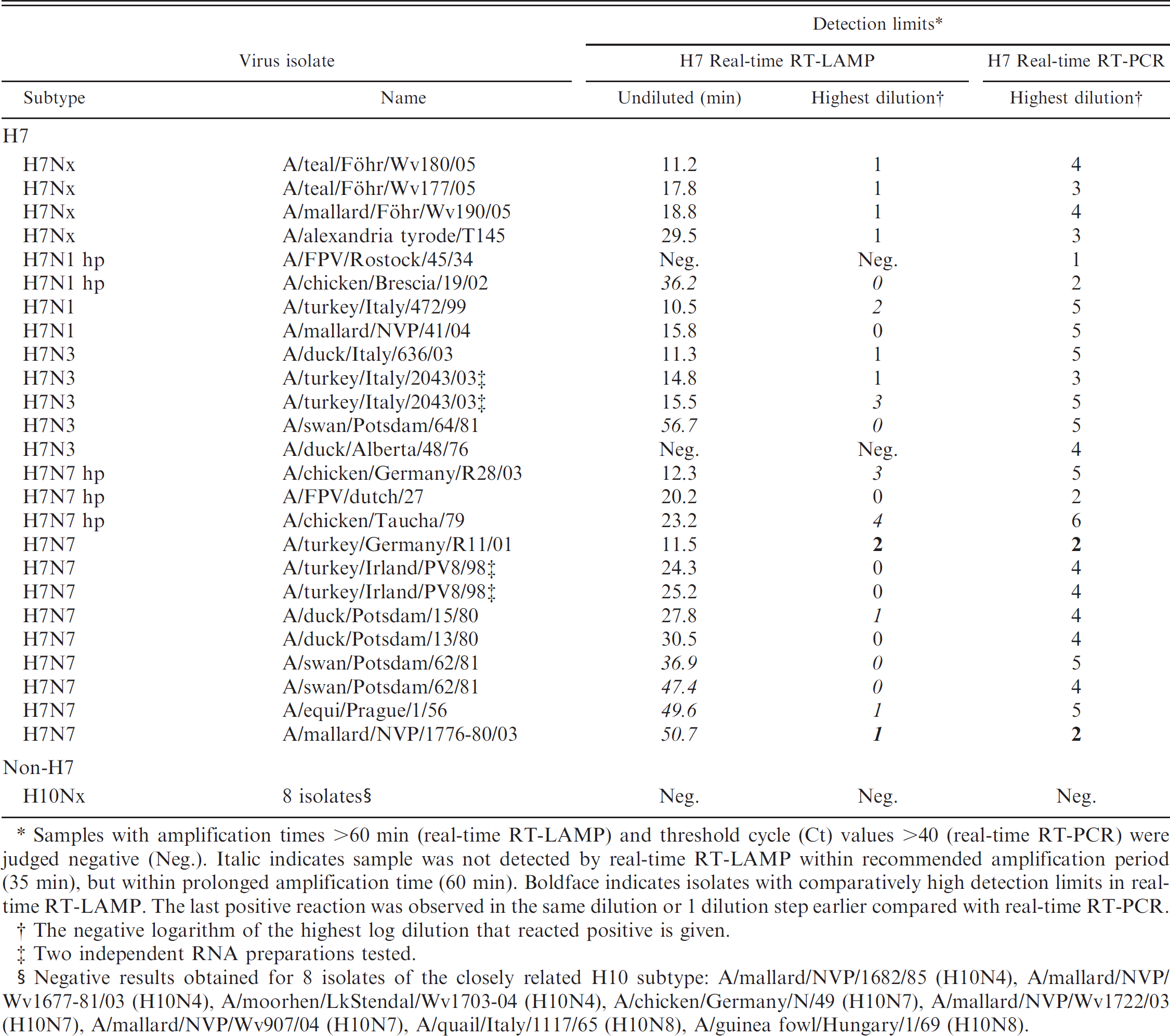
Samples with amplification times >60 min (real-time RT-LAMP) and threshold cycle (Ct) values >40 (real-time RT-PCR) were judged negative (Neg.). Italic indicates sample was not detected by real-time RT-LAMP within recommended amplification period (35 min), but within prolonged amplification time (60 min). Boldface indicates isolates with comparatively high detection limits in realtime RT-LAMP. The last positive reaction was observed in the same dilution or 1 dilution step earlier compared with real-time RT-PCR.
The negative logarithm of the highest log dilution that reacted positive is given.
Two independent RNA preparations tested.
Negative results obtained for 8 isolates of the closely related H10 subtype: A/mallard/NVP/1682/85 (H10N4), A/mallard/NVP/Wv1677–81/03 (H10N4), A/moorhen/LkStendal/Wv1703–04 (H10N4), A/chicken/Germany/N/49 (H10N7), A/mallard/NVP/Wv1722/03 (H10N7), A/mallard/NVP/Wv907/04 (H10N7), A/quail/Italy/1117/65 (H10N8), A/guinea fowl/Hungary/1/69 (H10N8).
Two H5-specific RT-LAMP assays differed very much in their sensitivities. 5,6,8 An H5N1-specific real-time RT-LAMP with a long 2-hr amplification phase revealed detection limits comparable to conventional RT-PCR for recent highly pathogenic H5N1 strains belonging to clades 1 and 2, respectively. 8 However, no comparison with real-time RT-PCR was reported.
In line with the previously described study, an extension of the recommended amplification time from 35 to 60 min also increased the sensitivity for some strains in our study (Tables 1 and 2, indicated in italic) but, at the same time, decreased the specificity of the H5-specific primer set. For the RT-LAMP assays evaluated here, the possibilities of a prolongation of the amplification phase are limited, in that, already, a 1-hr amplification period led to false positive results. It might be possible to reduce the likelihood of false positives during prolonged amplification by adjusting the chemistry of the reaction toward minimized non-specific primer binding. On the other hand, prolonged amplification times would interfere with the aspired point-of-care and the pen site use of real-time RT-LAMP.
The results obtained in the current study are in line with other studies about H5-specific real-time RT-LAMP assays showing a highly variable sensitivity for different H5 isolates. 5,6 Specific H5N1 virus isolates of Asian origin belonging to different phylogenetic clades, as well as some Eurasian LPAI H5 viruses, were particularly well detected (Table 1, indicated by boldface). However, a number of H5 isolates, including several Southeast Asian HPAIV strains, as well as LPAIV isolates of recent Eurasian origin, were detected with diminished sensitivity. It had been reported previously that, for some of the older H5 isolates like A/tern/ SouthAfrica/61, LAMP-based RNA detectionisabout 1,000 times less sensitive, even compared with conventional RT-PCR. 5,6 This observation is supported by the finding that this isolate also owned the lowest detection limits in the evaluated H5 real-time RT-LAMP. Real-time RT-PCR proved to be even 100,000 times more sensitive in this case.
Comparison of Avian influenza virus–specific RNA detection by real-time reverse transcription polymerase chain reaction (real-time RT-PCR) and real-time reverse transcription loop-mediated isothermal amplification (real-time RT-LAMP) in diagnostic samples originating from wild birds and poultry in Germany in 2007.
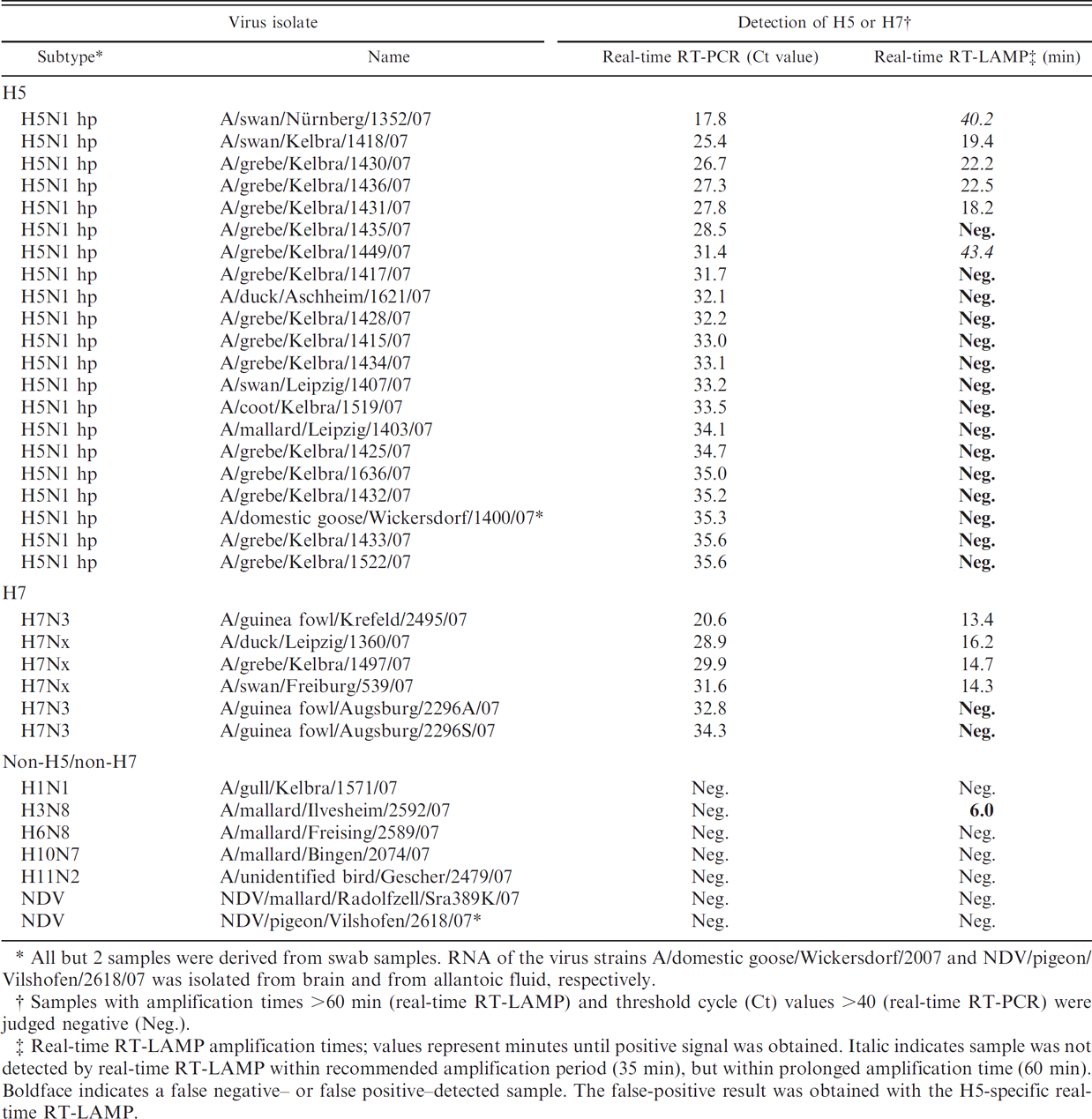
All but 2 samples were derived from swab samples. RNA of the virus strains A/domestic goose/Wickersdorf/2007 and NDV/pigeon/Vilshofen/2618/07 was isolated from brain and from allantoic fluid, respectively.
Samples with amplification times >60 min (real-time RT-LAMP) and threshold cycle (Ct) values >40 (real-time RT-PCR) were judged negative (Neg.).
Real-time RT-LAMP amplification times; values represent minutes until positive signal was obtained. Italic indicates sample was not detected by real-time RT-LAMP within recommended amplification period (35 min), but within prolonged amplification time (60 min). Boldface indicates a false negative– or false positive–detected sample. The false-positive result was obtained with the H5-specific realtime RT-LAMP.
The need for optimally matching primers governs the sensitivity of LAMP assays, and long stretches of highly conserved regions are required in the target sequence. Problems of designing an optimally matching universal set of broadly reactive H5- or H7-specific LAMP primers are inherent in the high variability of the Influenza A virus HA sequences and the special constraints of LAMP primer design. 5,6 The observed lower average sensitivity of the H5-and H7-specific real-time RT-LAMP assays in our study might be due to primers that do not equally well match with the phylogenetically very divergent isolates used in this broad evaluation. LAMP primers tailored to detect with high sensitivity a limited set of, e.g., H5 isolates currently circulating in a circumscribed region are expected to yield better sensitivities. 2,5,6 In this line, it is interesting to note that for the Influenza A virus M gene, which harbors a series of highly conserved sequence stretches, a sensitive generic RT-LAMP assay was successfully developed by using degenerated primers and an amplification phase of 2 hr. 13
On the basis of this study, the real-time RT-LAMP technology using the commercial primer kits Avian Flu H5 and Avian Flu H7 is currently less suitable for generalized diagnostic approaches in the veterinary field when compared with real-time RT-PCR. Nevertheless, the LAMP principle remains a promising technology to develop new field-applicable tests. The power is demonstrated when detecting certain isolates for which the primer sets seem to be specifically adapted (Tables 1 and 2, indicated by boldface). Although this study reveals the limitations of the 2 evaluated real-time RT-LAMP assays, they can still be considered useful diagnostic tools, provided the user is aware of these limitations. In the case of suboptimally equipped laboratories in countries endemically infected by HPAIV, these assays might be an alternative to rapid antigen testing for the monitoring of AIV on the flock level. In this regard, further constraints need to be observed: Adequate storage (cold chain) of the enzymes is a critical factor for test performance and sensitivity, and a low-titer positive control should be supplied as “sensitivity control” with the kit to monitor for such influences. For the benefit of a rapid, sensitive, and specific point-of-care and pen-site diagnosis of AIV infections, a generic approach-based (e.g., on fragments of the M or NP gene segments) assay might currently be more promising. In regions that experience endemic HPAIV infections but have low prevalence of other AIV infections (e.g., HPAIV H5N1 in Egypt or Indonesia), targeted restriction actions could be initiated on the basis of such a generic AIV RNA detection. If subtype-specific RT-LAMP assays are preferred, a specific, timely, and regular adaptation of the primer sets to presently and regionally circulating virus strains seems to be indispensable to achieve optimal sensitivities.
Acknowledgements. The authors are grateful to Bianka Hillmann, Martina Maron, Diana Wessler, Stephanie Franz, and Cornelia Illing for excellent technical assistance. This study has been funded by the Federal Ministry of Food, Agriculture, and Consumer Protection (BMELV), Germany (FSI project 2–1.2.1), and by the European Network of Excellence ”Epizone.”
Footnotes
a.
Eiken Chemical Co. Ltd., Tokyo, Japan.
b.
Viral RNA, Qiagen GmBH, Hilden, Germany.
c.
Superscript® III, Platinum® Taq; Invitrogen GmbH, Karlsruhe, Germany.
