Abstract
Reoviruses are nonenveloped, segmented, double-stranded RNA viruses capable of infecting a wide range of invertebrate, vertebrate, fungus, and plant hosts. Though sporadic infection has been reported in a variety of reptilian species, infection of rough green snakes (Opheodrys aestivus) has not been previously described. Five wild-caught, adult rough green snakes were obtained by a zoological institution. Clinical deterioration was first noted in all snakes after 3 weeks in quarantine. Despite treatment, clinical decline progressed, and all 5 snakes died or were euthanized by 48 days post-arrival. Moderate, multifocal, acute, necrotizing hepatitis with hepatocellular syncytia was diagnosed in 1 snake. Two additional snakes had severe, diffuse, subacute to chronic pancreatitis. All 5 snakes had gastroenteric cryptosporidiosis. Electron microscopic examination of liver from the snake with hepatic lesions revealed scattered hepatocytes containing 1 or more intranuclear clusters of approximately 90 nm in diameter viral particles arranged in loose arrays. Polymerase chain reaction (PCR) amplification of a segment of the reovirus RNA-dependent RNA polymerase gene was performed on RNA extracted from tissues of all 5 snakes. PCR amplification of samples extracted from the snake with hepatic lesions resulted in a 109–base pair (bp) product. Phylogenetic analyses indicated the virus was a novel strain distinct from other reoviruses at a level consistent with species difference. The source of infection was unknown. PCR amplification of samples extracted from the other 4 snakes was negative.
Introduction
Reoviruses are nonenveloped, segmented, double-stranded RNA viruses capable of infecting a wide range of invertebrate, vertebrate, fungus, and plant hosts. Vertebrate reoviral pathogens are typically associated with disease of the respiratory and/or alimentary systems.
Sporadic reports of reovirus infection have been documented in a variety of reptilian species (Blahak S, Göbel T: 1991, A case reported of a reovirus infection in an emerald tree boa (Corallus caninus). In: Proceedings of the 4th International Colloquium on the Pathology and Medicine of Reptiles and Amphibians, pp. 13–16, Bad Nauheim, Germany, September 27–29. Deutsche Veterinaermedizinische Gesellschaft, Giessen, Germany). 1,3,6,12,14,18,20,21,24,29 To date, reptilian infections have been attributed to viruses in the genus Orthoreovirus. All previously characterized reptile reoviruses have been cultured prior to molecular characterization. 30 Clinically, disease presents as pneumonia and/or neurologic dysfunction. Though clinical presentation may be suggestive, definitive diagnosis is challenging due to similarities in clinical and pathologic findings between reovirus and other reptilian viral infections. Experimental reovirus infection in snakes has resulted in fatal proliferative interstitial pneumonia. 18 Other snake pathogens associated with similar clinical findings and/or lesions, particularly inclusion body disease (IBD) and ophidian paramyxoviruses (OPMVs), must be distinguished from reovirus infection. 15,25 Characteristic viral inclusions may be present in cases of IBD and OPMV to aid in diagnosis; inclusions are not a feature of reovirus infection. 25 Reptilian reoviruses are fusogenic resulting in the formation of syncytia within infected tissues. 7 Syncytia are not unique to reovirus infections, however, and may be noted in other viral infections including OPMV. Definitive confirmation requires additional testing such as virus isolation and/or molecular diagnostic techniques (e.g., polymerase chain reaction [PCR] and sequencing).
Rough green snakes (Opheodrys aestivus) are colubrid snakes native to North America. Their diet consists mostly of arthropods. 23 The only disease process previously described in this species is intestinal cryptosporidiosis. 4 The following report describes reovirus infection in a group of captive rough green snakes confirmed via PCR and sequencing.
Materials and methods
Animals and clinical findings
Five wild-caught, adult rough green snakes were obtained from Florida by a zoological institution. Upon receipt, all snakes were maintained in quarantine facilities at the zoo and housed individually in 18.9-liter acrylic enclosures containing various sized branches, few plastic plants, and dampened paper towels and sphagnum moss substrate. Enclosure temperature was maintained at 24–27°C, and enclosures were misted 2–3 times per day. Each snake was offered 5 adult crickets (Acheta domesticus) and ad libitum water daily. Clinical deterioration was first noted in all snakes after 3 weeks in quarantine. Despite treatment, clinical decline progressed, and all 5 snakes died or were euthanized by 48 days post-arrival.
Pathology
Immediately following death and/or euthanasia, carcasses were submitted for necropsy. Representative sections of all tissues were fixed in 10% neutral buffered formalin, processed routinely, embedded in paraffin, sectioned at 5 μm, and stained with hematoxylin and eosin (HE) for routine light microscopic examination.
Transmission electron microscopy
Formalin-fixed sections of liver from snake 1 were post-fixed in 5% glutaraldehyde followed by 1% osmium tetroxide, then embedded in epoxy resin. Ultrathin sections were contrasted with uranyl acetate and lead citrate and then analyzed with a transmission electron microscope.
RNA extraction
For all snakes, formalin-fixed, paraffin-embedded (FFPE) tissues were used for RNA extraction. RNA was extracted from five 9-μm–thick sections cut from each block of interest. Tissue extracted from snake 1 included liver and lung; snake 2 included liver, heart, stomach, intestine, and lung; snake 3 included pancreas, stomach, and intestine; snake 4 included pancreas, stomach, and intestine; and snake 5 included liver, lung, pancreas, stomach, and intestine. To remove paraffin, sections were incubated in 1 ml of xylene at room temperature on a rocker platform for 10 min. Following incubation, samples were centrifuged at 10,000 × g for 10 min, and supernatant was removed. Incubation and centrifugation were repeated 3 times. The resultant tissue pellet was washed twice with 1 ml of cold 100% ethanol and centrifuged for 3 min at 16,000 × g. Supernatant was removed, and tissue pellets were dried in a vacuum centrifuge for 10 min. Tissue pellets were resuspended in 600 μl of digestion buffer containing 0.5 mg/ml proteinase K, 20 mM Tris–HCl (pH 7.6), 20 mM ethylenediamine tetra-acetic acid, and 1% sodium dodecyl sulfate. Samples were then incubated at 50°C for 4 hr. Following incubation, samples were centrifuged for 15 sec at 16,000 × g, and then 250 μl of buffered phenol and 250 μl of chloroform–isoamyl alcohol were added for extraction. Samples were vortexed and centrifuged at 16,000 × g for 2 min. The upper aqueous phase was removed and placed in a new microcentrifuge tube. The phenol extraction step was repeated for each sample. The resultant aqueous phase was mixed with an additional 500 μl of chloroform–isoamyl alcohol by vortexing. Samples were then centrifuged at 16,000 × g for 2 min, and the upper aqueous phase was again removed and placed into a new microcentrifuge tube. Extracted nucleic acids were precipitated by addition of one-tenth the extracted aqueous phase volume of 3 M sodium acetate (pH 7.0), twice the extracted aqueous phase volume of ice-cold 100% ethanol, and 20 μg of glycogen. Samples were inverted several times and vortexed briefly prior to an overnight incubation at −20°C. Following incubation, samples were centrifuged at 16,000 × g for 30 min. Supernatant was removed, and the pellet was washed with 1 ml of cold 70% ethanol. Samples were centrifuged at 16,000 × g for 15 sec and supernatant removed. Samples were dried in a vacuum centrifuge for 15 min. Finally, dried samples were resuspended in 30 μl of sterile dH2O prior to storage at −20°C.
Reovirus polymerase chain reaction
Polymerase chain reaction amplification of a segment of the reovirus RNA-dependent RNA polymerase gene was adapted from a previously published protocol, 30 but to increase chances of success with probable shorter fragments of RNA present in FFPE tissues, a shorter target was selected. The first round utilized forward primer 2090F (5′-GGBTCMACNGCYACYTCBACYGAGCA-3′) and reverse primer 2334R (5′-CDATGTCRTAHWYCCANCCRAA-3′), which was the second round of the original protocol. The second round utilized 2090F and a different reverse primer, 2200R (5′-CCRTCRTCWCCYTGRCAKACRTARTT-3′). Reaction compositions and conditions were otherwise identical to those previously reported. The PCR products were resolved in 1% agarose gels. Bands were excised and purified using a commercial kit. a The purified products were directly sequenced using a commercial cycle sequencing kit b and analyzed using an automated sequencer. c All products were sequenced in both directions, and primer sequences were edited out prior to analyses.
Phylogenic analysis
The sequences were compared with those in GenBank, European Molecular Biology Laboratory (Cambridge, UK), and Data Bank of Japan (Mishima, Shizuoka, Japan) databases using TBLASTX (http://blast.ncbi.nlm.nih.gov/Blast.cgi). 2 Predicted homologous 36 amino acid sequences of reoviral RNA-dependent RNA polymerase were aligned using 3 methods: 1) ClustalW, 27 2) T-Coffee, 22 and 3) MUSCLE. 8
Bayesian analyses of amino acid alignments were performed using MrBayes 3.1 26 with gamma distributed rate variation and a proportion of invariant sites, and mixed amino acid substitution models. Four chains were run, and statistical convergence was assessed by looking at the standard deviation of split frequencies as well as potential scale reduction factors of parameters. The first 10% of 1,000,000 iterations were discarded as a burn in.
Maximum likelihood (ML) analyses of amino acid alignments were performed using PHYLIP (Phylogeny Inference Package, version 3.66), 10 running each alignment in proml with amino acid substitution models JTT, 16 PMB, 28 and PAM 17 further set with global rearrangements, 5 replications of random input order, less rough, gamma plus invariant rate distributions, and unrooted. The values for the gamma distribution were taken from the Bayesian analysis. Striped bass reovirus (GenBank accession no. AAM93410) was designated as the outgroup. The alignment producing the most likely tree was then used to create data subsets for bootstrap analysis to test the strength of the tree topology (200 resamplings), 9 which was analyzed using the amino acid substitution model producing the most likely tree.
Results
Clinical findings
Approximately 3 weeks after arrival, all 5 snakes were noted to have multiple areas of brown to black cutaneous discoloration. In snakes 2 and 4, areas of discoloration focally corresponded to cutaneous abscesses. In snake 2, the cutaneous lesion extended into the muscular layers of the subjacent body wall. Snakes 2–4 also exhibited dysecdysis, and in snakes 1 and 2, poor body condition was evident. All snakes were treated with a single daily dose of enrofloxacin (10 mg/kg by mouth) for 3–25 days.
Despite treatment, snake 1 was found dead in its enclosure 3 days following onset of clinical signs. The condition of snake 2 continued to deteriorate over the next week. Progressive weight loss, dehydration, and lethargy were noted. Four doses of ceftazidime (20 mg/kg intramuscular) and 3 doses of metronidazole (25 mg/kg by mouth) administered at 48-hr intervals were added to the treatment regimen. Snake 2 died 16 days following initiation of treatment. Cryptosporidiosis was diagnosed in snakes 3 and 5 via gastric lavage. Consequently, snakes 3–5 were euthanized 48 days after initial receipt.
Pathology
Grossly, snake 1 was in marginal body condition with reduced coelomic adipose stores and multifocal cutaneous ulceration. Histologically, the liver contained multiple, random, small to moderate-sized foci of hepatic necrosis. Within affected areas, hepatic cords were discontinuous and disorganized. Hepatocytes were smudged and hypereosinophilic with loss of individual cellular detail and karyolysis. Necrotic foci frequently contained small numbers of multinucleated syncytial cells and few heterophils (Fig. 1). Sinusoids in affected areas frequently contained small amounts of eosinophilic granular debris and scattered melanophages. Other histologic findings included mild gastric cryptosporidiosis and severe, multifocal, necro-ulcerative dermatitis with intralesional yeast and bacteria.
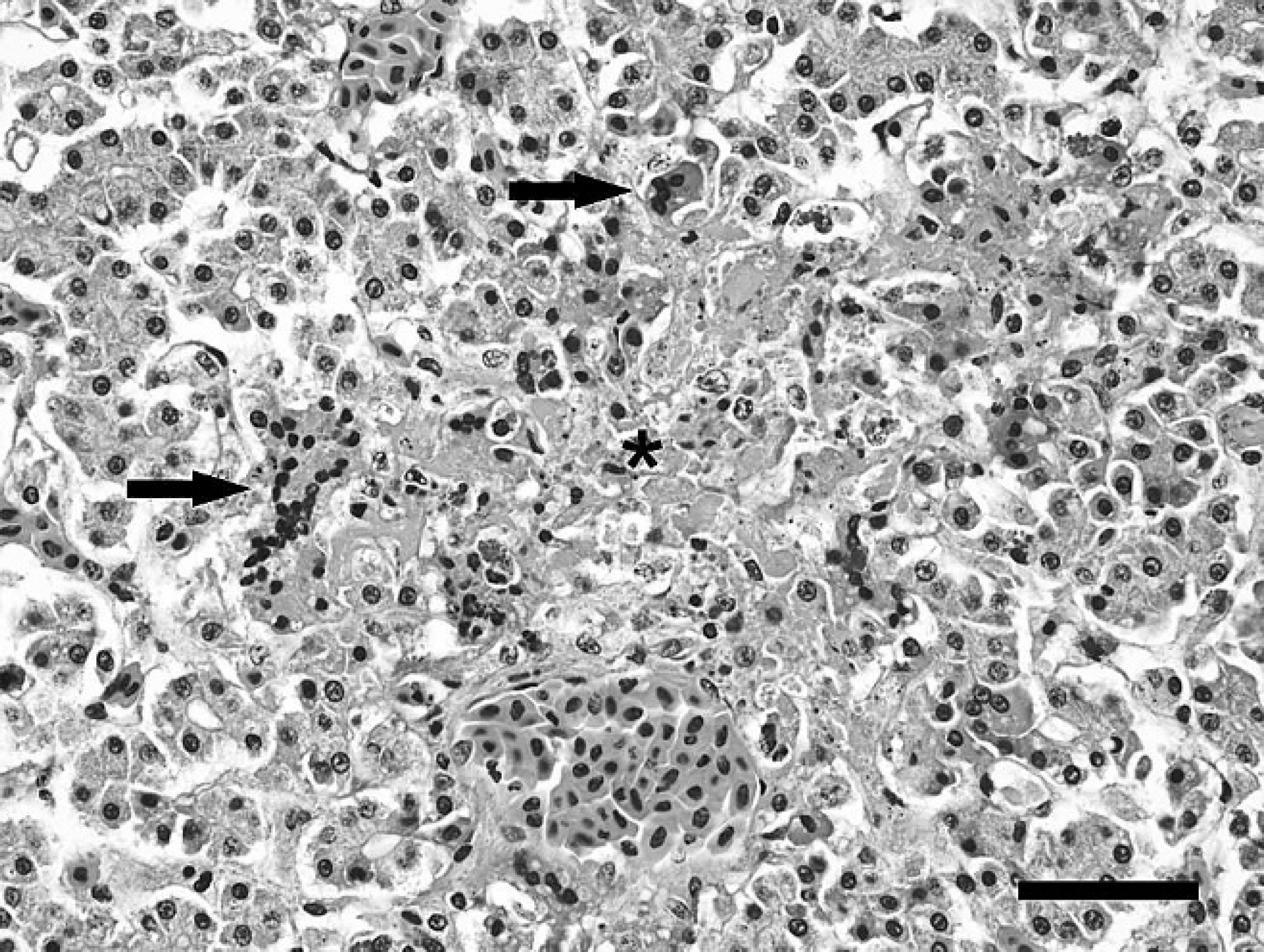
Snake 1, rough green snake (Opheodrys aestivus), liver. Centrilobular hepatocytes have loss of individual cellular detail and karyolysis indicative of necrosis (asterisk). Adjacent to the area of necrosis are hepatocellular syncytia (arrows). Hematoxylin and eosin. Bar = 50 μm.
Snake 2 lacked significant gross abnormalities. Histologically, significant findings included moderate diffuse hepatocellular and coelomic adipose atrophy indicative of inanition, and moderate gastroenteric cryptosporidiosis with gastric glandular atrophy. Other histologic findings included mild pulmonary nematodiasis (Rhabdias spp.) and mild, multifocal ulcerative dermatitis with intralesional yeast.
Grossly, snake 3 was in marginal body condition with moderately reduced coelomic adipose stores and mildly reduced skeletal musculature. Histologically, approximately 98% of the exocrine pancreas was replaced by large confluent accumulations of hyper-cellular, disorganized, collagen-rich fibrous connective tissue with numerous lymphocytes, fewer macrophages, and scattered heterophils (Fig. 2). The few remaining exocrine acini were individualized, widely separated, and disorganized. Pancreatic ducts were variably ectatic and surrounded by concentric accumulations of fibrous connective tissue with mixed inflammatory cells. Ductular epithelial cells were frequently piled and mildly disorganized with large vesicular nuclei (hyperplasia). Some ducts contained intraluminal and/or intraepithelial accumulations of karyorrhectic debris. Other histologic findings included moderate, diffuse hepatocellular, coelomic adipose, skeletal muscular atrophy indicative of inanition; moderate gastroenteric cryptosporidiosis with gastric glandular atrophy; and mild pulmonary nematodiasis.
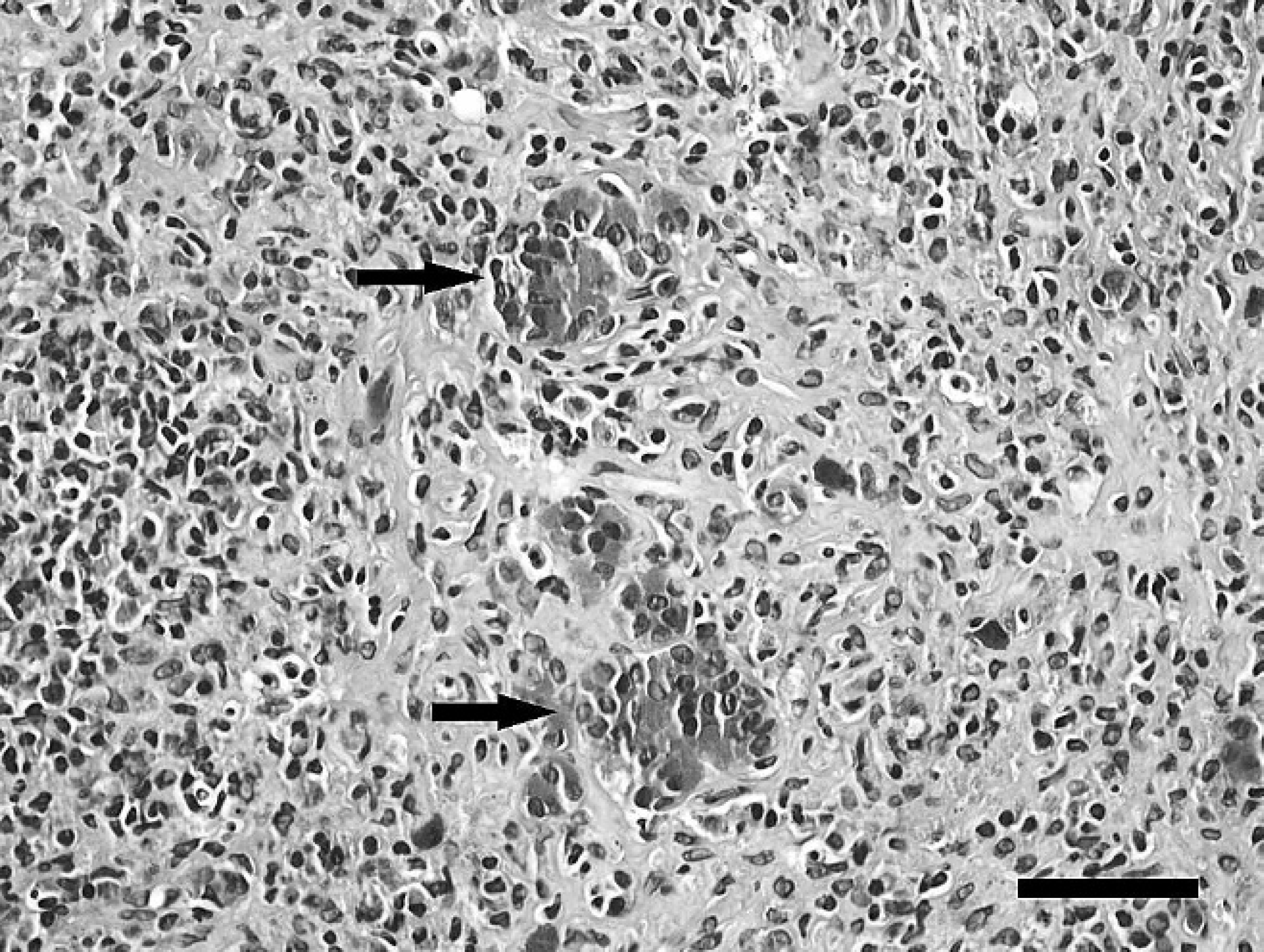
Snake 3, rough green snake (Opheodrys aestivus), pancreas. The pancreatic parenchyma is largely replaced by accumulations of hypercellular, disorganized, collagen-rich fibrous connective tissue with numerous lymphocytes, fewer macrophages, and scattered heterophils that separate and distort the few remaining acini (arrows). Hematoxylin and eosin. Bar = 50 μm.
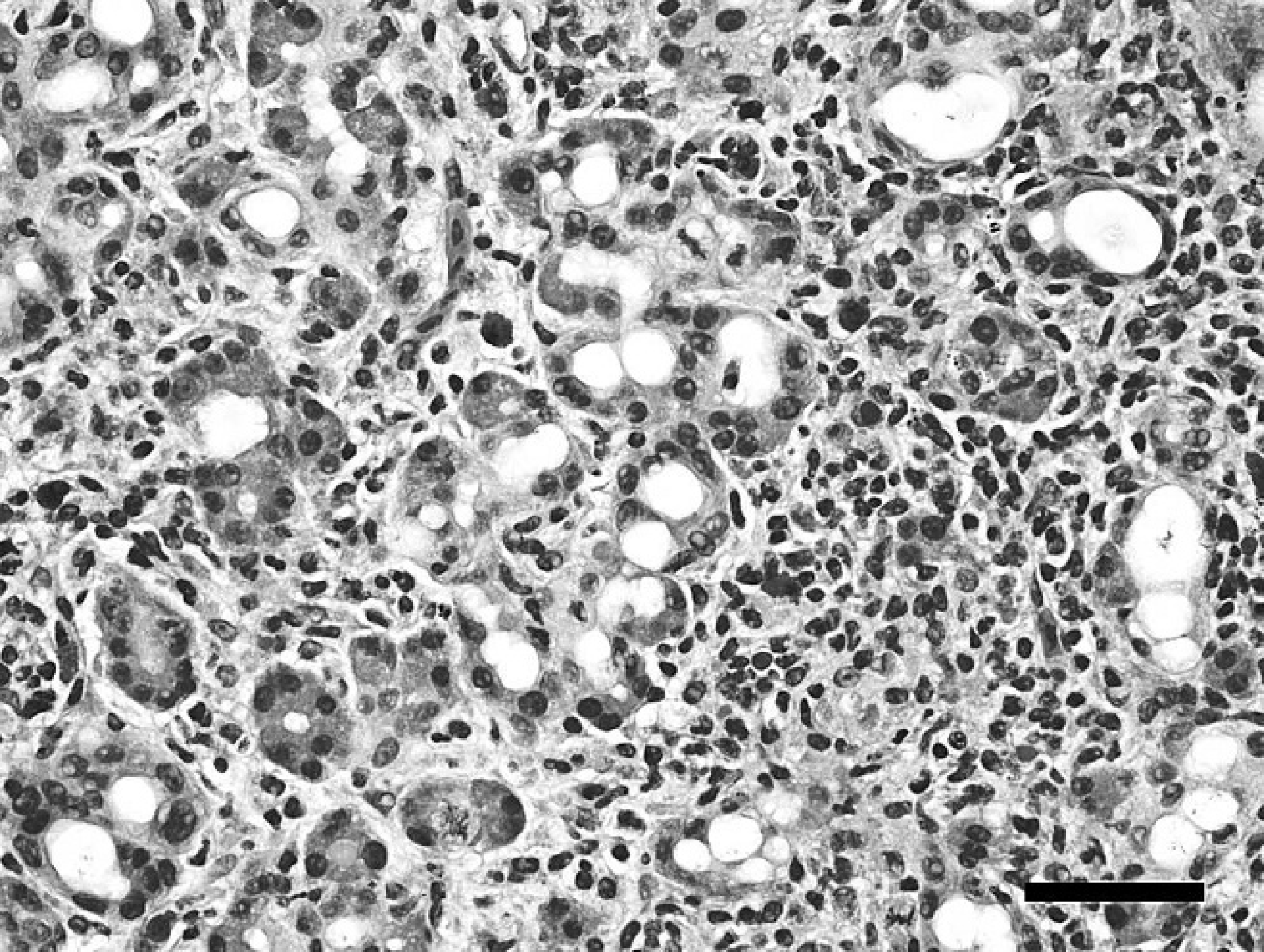
Snake 4, rough green snake (Opheodrys aestivus), pancreas. Pancreatic acini are separated by small to moderate amounts of hypercellular, disorganized, collagen-poor fibrous connective tissue with numerous lymphocytes and scattered heterophils. Acini are frequently mildly dilated and lined by attenuated, disorganized, and multifocally piled epithelial cells lacking zymogen granules (regeneration). Hematoxylin and eosin. Bar = 50 μm.
Snake 4 lacked significant gross abnormalities. Histologically, the pancreatic interstitium contained moderate amounts of hypercellular, disorganized, collagen-poor fibrous connective tissue with numerous lymphocytes and scattered heterophils that separated and distorted acini and ducts. Acini were frequently mildly dilated and lined by attenuated, disorganized, and multifocally piled epithelial cells lacking zymogen granules (regeneration). Ducts were also often ectatic and lined by regenerative epithelium. Rare ducts and acini contained scant cellular debris (Fig. 3). Other significant histologic findings included mild, diffuse hepatocellular and coelomic adipose atrophy indicative of inanition, and mild enteric cryptosporidiosis.
Grossly, snake 5 was in poor body condition with absent coelomic adipose stores and moderately reduced skeletal musculature. Histologically, abnormalities included moderate to marked, diffuse hepatocellular, coelomic adipose, and skeletal muscular atrophy indicative of inanition; moderate gastroenteric cryptosporidiosis with mild gastric glandular atrophy; and mild pulmonary nematodiasis.
Transmission electron microscopy
Within ultrathin sections of liver from snake 1, small numbers of hepatocytes bordering areas of necrosis contained 1 or more intranuclear clusters of approximately 90 nm in diameter viral particles arranged in loose arrays (Fig. 4). Viral particles had an electron-dense, icosahedral nucleocapsid and morphology typical of reoviruses (Fig. 4 inset).
Reovirus polymerase chain reaction
Polymerase chain reaction amplification of samples extracted from snake 1 resulted in a 109-bp product when primer sequences were edited out. The sequence was submitted to GenBank under accession number FJ906820. PCR amplification of samples extracted from snakes 2–5 was negative.
Phylogenic analysis
TBLASTX results showed the highest score with African grey parrot reovirus RNA polymerase (GenBank accession no. EU309695). All 3 alignment algorithms found no indels present, resulting in identical alignments with no gaps. Bayesian phylogenetic analysis showed the Wag model of amino acid substitution was most probable with a posterior probability of 0.624, 31 followed by the Blosum62 model with a posterior probability of 0.360. 13 The Bayesian tree is shown in Figure 5. Maximum likelihood analysis found the most likely tree from the JTT model of amino acid substitution, and this model was used for bootstrap analysis. Maximum likelihood bootstrap values are shown on the Bayesian tree (Fig. 5).
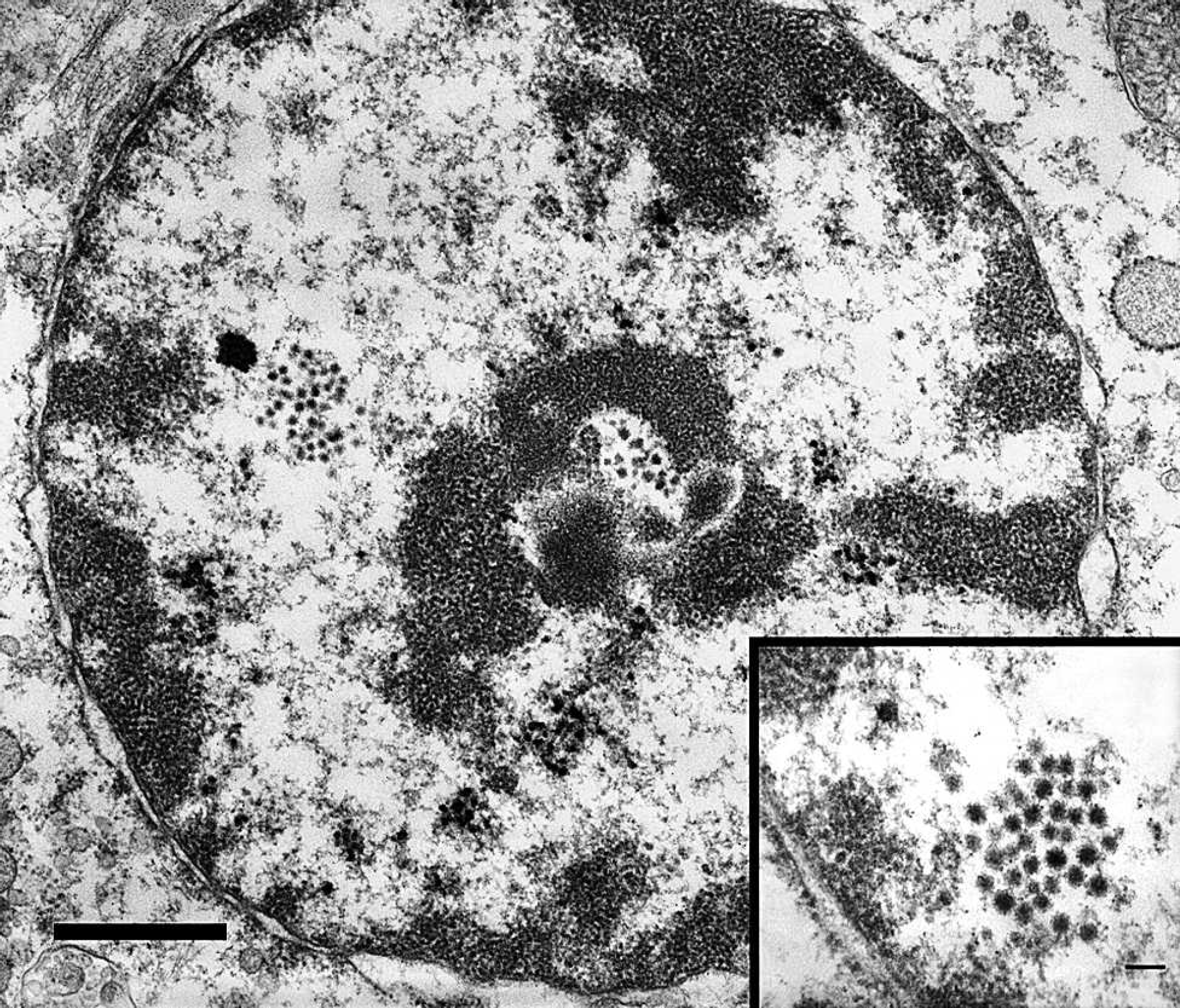
Snake 1, rough green snake (Opheodrys aestivus), liver. The hepatocyte contains few intranuclear clusters of electron dense viral particles arranged in loose arrays. Viral particles are approximately 90 μm in diameter with an electron dense icosahedral nucleocapsid and morphology typical of reoviruses. Uranyl acetate and lead citrate. Bar = 1 μm. Inset: uranyl acetate and lead citrate. Bar = 2 μm.
Discussion
Clinical course and pathologic lesions noted in snake 1 were compatible with acute reovirus infection. Accordingly, viral infection was confirmed in extracted samples from snake 1 via reovirus PCR. Viral particles were also visible within hepatic lesions examined by transmission electron microscopy. Pancreatic fibrosis and regeneration noted in snakes 3 and 4 were compatible with chronic and resolving damage. Due to the chronicity of the lesions, no definitive cause was apparent in examined sections. Chronic pancreatitis similar to that noted in snakes 3 and 4 has been noted in reovirus infections of other reptilian species (Kinsel, Landolfi, and Terio, unpublished data). The morphologic compatibility of the lesions combined with confirmed infection in snake 1 suggested previous reovirus infection was a likely differential for the pancreatic lesions of snakes 3 and 4. Failure to amplify reovirus in samples from these individuals could have been due to either the stage of disease at the time of examination or to false-negative results from use of FFPE tissues. Hepatitis was only documented in snake 1, and thus appeared to be associated with acute disease. Pancreatitis was lacking in snake 1, suggesting pancreatitic involvement in reovirus infection may occur later in the course of disease.
Lesions suggestive of acute or chronic reovirus infection were lacking in tissues examined from snakes 2 and 5. Similarly, reovirus PCR was negative for samples extracted from snakes 2 and 5. Because snake 2 died early in the time course of the outbreak, peracute reovirus infection could have contributed to clinical signs and demise and accounted for the lack of noted lesions. However, reovirus infection has not been previously reported to result in peracute disease in reptiles. Alternatively, snakes 2 and 5 may have been resistant to infection with the reovirus strain responsible for this outbreak. Because all snakes were housed individually at the zoo, it is also possible that snakes 1, 3, and 4 were already incubating the virus at the time of acquisition, and snakes 2 and 5 may never have been adequately exposed to cause productive infection.
The source of this reovirus is unknown. Since Opheodrys aestivus is a dietary arthropod specialist, 5 and orthoreoviruses have not been identified from arthropod hosts, it is considered unlikely that the virus was acquired from a prey item. The most probable source of infection is from another animal at the supplier's facility, where there was significant potential for exposure to other recently acquired herptiles.
The virus strain characterized in the current study is distinct from other reoviruses at a level consistent with species difference. Unlike previously reported reoviruses of reptiles, this virus was not cultured prior to PCR amplification. It is possible that selection for viruses that grow on routinely used cell lines represents a significant filter, resulting in missing some orthoreoviruses. Virus culture was not attempted from these snakes, and frozen tissues were not available.
The addition of this virus to the data set significantly alters the placement of the Nelson Bay/African grey parrot clade; a previous analysis found that this clade diverged prior to the separation of the mammal “Orthoreovirus/Ndelle” clade from the reptile reovirus clade. 30 However, the support in the prior analysis for this branching order was not strong (as defined by a Bayesian posterior probability >90 and ML bootstrap value >70). Despite the shorter sequence length, the analysis produced by the current study finds that the addition of the rough green snake reovirus results in strong support for clustering of the Nelson Bay/African grey parrot clade with the reptile reovirus clade after divergence of the Mammal Orthoreovirus/Ndelle clade. This would place the divergence between fusogenic and nonfusogenic orthoreoviruses as basal within the genus. The availability of a more complete representation of existing species for comparison results in greater phylogenetic resolution. 11 Significant errors can occur in phylogenetic analyses due to incomplete taxa sampling, despite very large sequence length. 19 Work toward discovery of additional orthoreoviruses is indicated.
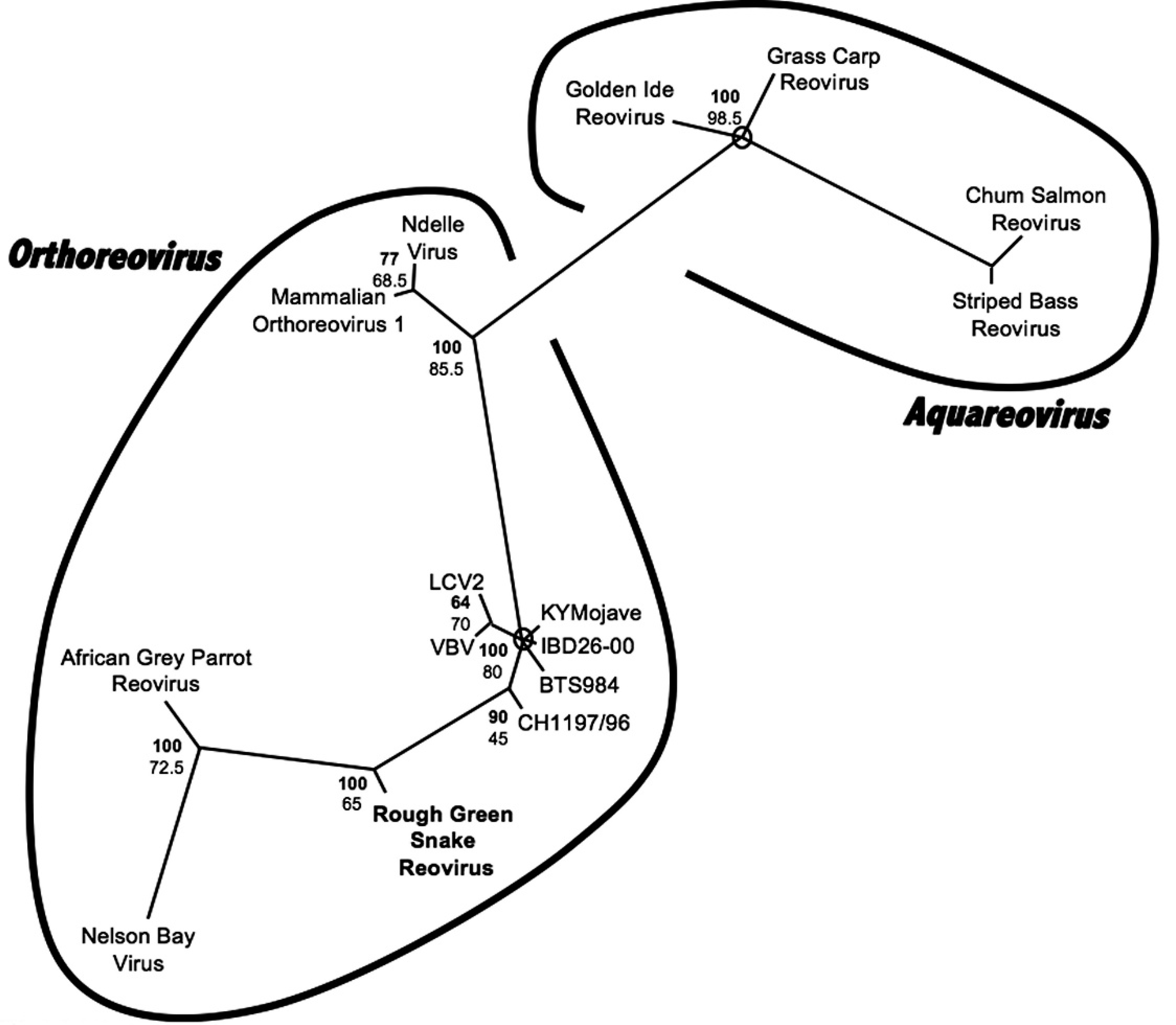
Bayesian phylogenetic tree of predicted 36 amino acid partial reoviral RNA-dependent RNA polymerase sequences. Bayesian posterior probabilities of branchings as percentages are in bold, and maximum likelihood (ML) bootstrap values for branchings based on 200 resamplings are given below. Striped bass reovirus (GenBank accession no. AAM93410) was designated as the outgroup. Brackets delineate virus genera. Branches with Bayesian posterior probabilities less than 60 are collapsed, and circles mark areas of multifurcation. Rough green snake reovirus is in bold. Sequences retrieved from GenBank include Chum salmon reovirus (GenBank accession no. AAL31497), Golden ide reovirus (AAM93415), Grass carp reovirus (AAG10436), Mammalian orthoreovirus serotype 1 (NP694626), Ndelle virus (AAL36027), Nelson Bay orthoreovirus (EU309694), African Grey parrot reovirus (EU309695), Reptile reovirus KYMojave (EU309696), Reptile reovirus CH1197/96 (EU309697), Reptile reovirus IBD26-00 (EU309701), Reptile reovirus LCV2 (EU309704), Reptile reovirus VBV (EU309705), and Reptile reovirus BTS984 (EU309706).
In all 5 snakes, concurrent disease processes, including parasitism and inanition, were present and contributed to clinical course and/or death. Intestinal cryptosporidiosis consistent with infection by a non–Cryptosporidium serpentis sp. has been previously documented in this species. 4,32 However, the gastric as well as intestinal tropism of the cryptosporidiosis noted in the current study was different from previous reports. Gastroenteric cryptosporidiosis, diagnosed at postmortem in both snakes 1 and 2, instigated gastric lavage of remaining live snakes. Euthanasia was elected in all cases for group health and management reasons. Pancreatic cryptosporidiosis was not considered a likely differential for chronic pancreatitis noted in snakes 3 and 4 because organisms were lacking in examined sections of pancreas. In fact, in all snakes, Cryptosporidium spp. infection was limited to the stomach and intestine.
The cases in the present study support the rationale for an extended quarantine period for newly acquired, and especially wild-caught, snakes. Quarantine programs should include complete physical examination and necessary diagnostic testing of new animals to ensure protection and maintenance of the health of the zoological collection as a whole. In particular, findings in the current study suggest that diagnostic gastric lavage for Cryptosporidium spp. infection surveillance in snakes may be an invaluable screening tool that could enhance the effectiveness of herptile quarantine programs. The results of the current report also highlight the utility and importance of complete gross and histopathologic evaluation of all deceased animals as a component of vigilant disease surveillance. Pathologic and associated ancillary diagnostic investigations described herein not only revealed a cause for illness in the quarantined snakes, but also identified additional subclinical infections that had group health implications. In light of these findings, introduction of disease into the zoo's existing captive herptile collection was prevented.
Acknowledgements
The authors would like to acknowledge Lou Ann Miller and the University of Illinois Center for Microscopic Imaging for assistance with electron microscopy. The authors also thank the University of Illinois histology laboratory for technical assistance.
Footnotes
a.
QIAquick® Gel Extraction Kit, Qiagen Inc., Valencia, CA.
b.
Big-Dye® Terminator Kit, Applied Biosystems, Foster City, CA.
c.
Applied Biosystems 3130 Genetic Analyzer, Applied Biosystems, Branchburg, NJ.
