Abstract
Several bacteria belonging to the family Pasteurellaceae are potential pathogens in rabbits. In particular, Pasteurella multocida is considered to be important, and outbreaks caused by this species result in considerable economic losses in rabbitries. However, Pasteurellaceae spp. isolated from rabbits are poorly characterized, and thus, proper identification of P. multocida isolates from these animals is problematic and often unsatisfactory, thereby hampering epidemiological investigations. Therefore, 228 isolates from rabbit populations originating from a breeding and fattening organization with group management and postmortem cases with pasteurellosis from individual owners were phenotypically and genotypically analyzed using biochemical tests and repetitive extragenic palindromic polymerase chain reaction (REP-PCR). Furthermore, 41 samples representing observed phenotypes were selected for phylogenetic analysis using 16S ribosomal RNA and rpoB genes. The REP-PCR typing and phylogenetic analyses correlated well and appeared to be distinct molecular methods for characterization of rabbit isolates. Phenotyping, however, diverged from molecular recognition, reflecting the problematic conventional diagnosis of these strains. The fermentation of sorbitol appeared to be an imprecise indicator for P. multocida subspecies classification. According to REP-PCR and sequencing results, 82% of the isolates were characterized as P. multocida subsp. multocida, 3% as P. multocida subsp. septica, and 5% as P. multocida. Further, 5% were identified as Pasteurella canis. The other 5% represented a homogeneous group of unknown species belonging to the Pasteurellaceae. Samples obtained from individual postmortem cases demonstrated a higher phenotypic and genetic heterogeneity than samples from group management rabbits.
Keywords
Introduction
Respiratory diseases, often induced by bacterial infections, are predominant causes of death in rabbits. 20 Pasteurella multocida is the most common pathogenic microorganism isolated from rabbits, and its prevalence has been reported to be between 7% and nearly 100%. 13,23,31 This bacterial species is considered an opportunist or secondary pathogen and can be found in the respiratory tract of healthy and diseased animals. 13 In rabbit colonies, it can emerge as a major pathogen causing upper respiratory tract infections, and pasteurellosis is the predominant bacterial disease in these animals, resulting in considerable economic losses in rabbitries. 12,14 Noninfected and resistant animals; chronically infected, healthy carriers; and animals with local infections (rhinitis, otitis media), pneumonia, and septicemia can be distinguished. 13,15,16 Besides P. multocida, little is known about the occurrence and importance of other members of the family Pasteu-rellaceae in rabbits.
The current identification of Pasteurellaceae, mainly based on complex phenotypic characterization, is time consuming and often imprecise because of possible differences in analytical methods and results interpretation, as well as variation between isolates from different hosts. 5 In particular, P. multocida isolates can be very heterogeneous in their biochemical activity, appearing as variations in a single key character and making the unambiguous assignment of this organism on the species level quite problematic. 8 Moreover, discrimination between subspecies of P. multocida, namely P. multocida subsp. multocida, P. multocida subsp. septica, and P. multocida subsp. gallicida, based on sorbitol, trehalose, and dulcitol fermentation reactions can be ambiguous. 6,7,17,21 A more feasible tool for diagnostic purposes is, therefore, DNA sequence-based identification. The comparative sequence analysis of the 16S ribosomal RNA (rRNA) coding gene as well as certain housekeeping genes have been successfully used for identification and for clarifying the phylogenetic relationship within the family Pasteurellaceae as well as on the P. multocida subspecies level. 10,25,26 However, separation of P. multocida into subspecies is rarely done but can be useful for epidemiological purposes. Interestingly, strains phylogenetically identified as P. multocida subsp. septica are clearly different from the 2 other indistinguishable subspecies, P. multocida subsp. multocida and P. multocida subsp. gallicida. 4,11,26
Repetitive extragenic palindromic polymerase chain reaction (REP-PCR) is a molecular technique that enables analysis of the distribution of REP elements in the entire bacterial genome. Fragments obtained after REP-PCR form a pattern that acts as a genomic fingerprint of species and even strains. 38 This method is known for its ease of application and interpretation, high discrimination power, good reproducibility, and the opportunity to perform PCR without extensive DNA extraction. 19 Repetitive extragenic palindromic PCR, which appears to be a suitable method for investigation of genetic relatedness between Pasteurellaceae strains, 1,18,36,37 has also been suggested as a useful approach for the description of genera, species, and subspecies of Pasteurellaceae. 9
In the present study, the phenotypic and genetic diversity within strains belonging to the Pasteurellaceae family obtained from rabbits was investigated. Whereas biochemical analysis showed high heterogeneity and in some cases provided unclear results, REP-PCR and comparative 16S rRNA and rpoB genes sequence analysis allowed precise genetic characterization of investigated strains.
Material and methods
Sources of isolates
During a 2-year period (2004–2006), 123 isolates from the nares or sinus of slaughtered group management rabbits were gathered. The animals came from different breeding and fattening farms belonging to a single Swiss rabbit-meat organization. In breeding farms, they were held indoors, in groups of 8 does, with 1 male and their offspring. The breeding material was provided by 1 main breeder. The offspring were distributed to fattening farms all over Switzerland where they were held indoors in groups up to 28 animals per cage. Clinical status of the animals was not available.
Another 105 strains were isolated from altered organs of postmortem cases with pasteurellosis, sent to the Institute of Veterinary Bacteriology (Zurich, Switzerland) by individual rabbit owners. The rabbits represented various breeds from different origins, were of all age groups, and showed pathologic findings, such as rhinitis, otitis media, pneumonia, pericarditis, pleuritis, conjunctivitis, phlegmon, abscesses, meningoencephalitis, mastitis, endometritis, salpingitis, or septicemia. Ninety of these isolates were collected during 1991 and 1992, and 15 were collected during 2004 and 2005.
Phenotyping
Affected organs or heart blood were investigated bacteriologically using routine procedures. Strains identified as species belonging to the family Pasteurellaceae were lyophilized and frozen using the Protect a system. For further characterization, all 228 isolates were subcultured from stock cultures on 5% sheep blood agar b and incubated under aerobic conditions at 37°C for 24 hr. The biochemical characterization was carried out as described previously. 28 Briefly, the genus Pasteurella was identified using the following criteria: independence of β-nicotinamide adenine dinucleotide (β-NAD), hemolysis, indole production, oxidase, catalase, and lysine decarboxylase. For identification at the species level, tests for the fermentation of dulcitol, sorbitol, mannitol, trehalose, and maltose and for the production of urease and ornithine decarboxylase (ODC) were performed. Reactions were observed after inoculation of 1–2 colonies of pure culture in 2 ml of growth medium with appropriate indicator and incubation at 37°C for 48 hr; sorbitol reactions were recorded up to 14 days.
Molecular characterization by REP-PCR
Preparation of cell lysate/DNA template. Three to 4 bacterial colonies from blood agar plates were transferred to 50 μl of nuclease-free water c and boiled in a heating block at 100°C for 10 min. After centrifugation at 10,000 × g for 5 min, the supernatant was collected and stored at 4°C until use.
REP-PCR. Fragments between the REP elements present in rabbit Pasteurellaceae isolates were generated in REP-PCR using the following degenerated primers: REP1R-IDt (5′-NNNNGCNGCNGTAGNCCG-3′) and REP2-IDt (5′-NCGNCTTATCNGGCCTAC-3′). 37 The PCR mixture in a 50-μl final volume contained 1 × PCR buffer with 4 mM MgCl2 and 1.25 U Taq DNA polymerase, 200 μM of each deoxyribonucleotide triphosphate (dNTP), c 50 pmol of each primer, and 2.5 μl of bacterial cell lysate as template. The REP-PCR was run in a thermal cycler d with an initial denaturation step at 95°C for 7 min, followed by 35 cycles of denaturation at 94°C for 1 min, annealing at 40°C for 1 min, and extension at 65°C for 8 min. The PCR was terminated with a final extension at 65°C for 16 min. 34 The amplified fragments were separated on a 1.5% agarose gel e with ethidium bromide in 1 × Tris-acetate-ethylenediamine tetraacetic acid (TAE) at 70 V for 240 min. The molecular size marker f ranged from 100 to 3,000 bp. Bands were visualized using ultraviolet transillumination, and the sizes of the fragments were analyzed with a computer-aided bioimage system. g The number and size of the obtained fragments provided a basis for delineation of the REP-PCR types. Profiles that differed in 1 or more bands were determined as a distinct type. To check the reproducibility of the method, a set of 40 strains was repeated.
Sequence analysis of the 16S rRNA and the rpoB genes
Based on REP-PCR results and phenotyping, 41 Pasteurella rabbit isolates were selected for sequencing and phylogenetic analysis of the 16S rRNA and rpoB genes. From the most frequently represented group, characterized as REP-PCR type I, 9 strains that differed in phenotype, representing both rabbit groups, were chosen for sequencing. From less-represented REP-PCR types II-IX, nearly all strains were included in the DNA sequence-based analysis. Sequences were determined as described previously. 25,27 Sequencing products were run on a genetic analyzer d followed by editing using the Sequencer soft-ware. h The sequences obtained were cross-compared against GenBank. Phylogenetic analysis of the proofread sequences and selected representatives of each described genus within the Pasteurellaceae family was performed using the tree-building neighbor-joining method with the BioNumerics program version 4.61. i All sequences were submitted to GenBank under the accession numbers ranging from EF579813 to EF579894.
Differentiation of rabbit Pasteurella isolates based on biochemical activity type and repetitive extragenic palindromic polymerase chain reaction (REP-PCR).*

+ = positive reaction; - = negative reaction for 14 days. Numbers in italics indicate the number of isolates from group management, and numbers in bold indicate the number of isolates from clinical cases.
Comparison of the methods
Comparison between 16S rRNA and rpoB gene sequence-derived clusters and REP-PCR types was done with BioNumerics program version 4.61 i using the congruence of methods feature based on Pearson correlation test.
Results
Phenotyping
All 228 isolates were β-NAD-independent, nonhemolytic, and oxidase and catalase positive and negative for lysine decarboxylase and the production of urease. Differences were observed in the production of indole; the fermentation of dulcitol, sorbitol, mannitol, trehalose, and maltose; and in the presence of ODC. Based on the biochemical activity of the strains, a total of 12 diverse biochemical types (BT) could be recognized (Table 1). Phenotypically, 209 rabbit isolates could be identified as P. multocida. Most of these strains (n = 116) were assigned to BT 2. They were positive for indole, sorbitol, mannitol, and ODC. Negative reactions were observed for dulcitol, trehalose, and maltose. The second largest group within the phenotypically delineated P. multocida strains was BT 7, represented by 68 isolates, which were positive for indole, mannitol, and ODC and negative for dulcitol, sorbitol, trehalose, and maltose. The remaining 25 P. multocida strains were distributed among the other BTs (1, 3–6, 8–10). Eleven isolates were identified as Pasteurella canis (BT 11), and 8 isolates were unidentifiable based on their phenotypic appearance (BT 12).
Molecular characterization by REP-PCR
Application of REP-PCR for all investigated strains resulted in 9 different patterns (Fig. 1; Table 2). Whereas REP-PCR types I-III as well as IV-VI demonstrated similar patterns, REP-PCR types VII-IX differed strongly from each other. Isolates obtained from group management rabbits in particular belonged to REP-PCR type I, 1 strain showed a pattern typical for type II, and 10 were assigned as type VII. Isolates from postmortem cases were much more heterogeneous, covering each of the 9 REP-PCR types (Tables 1, 2).
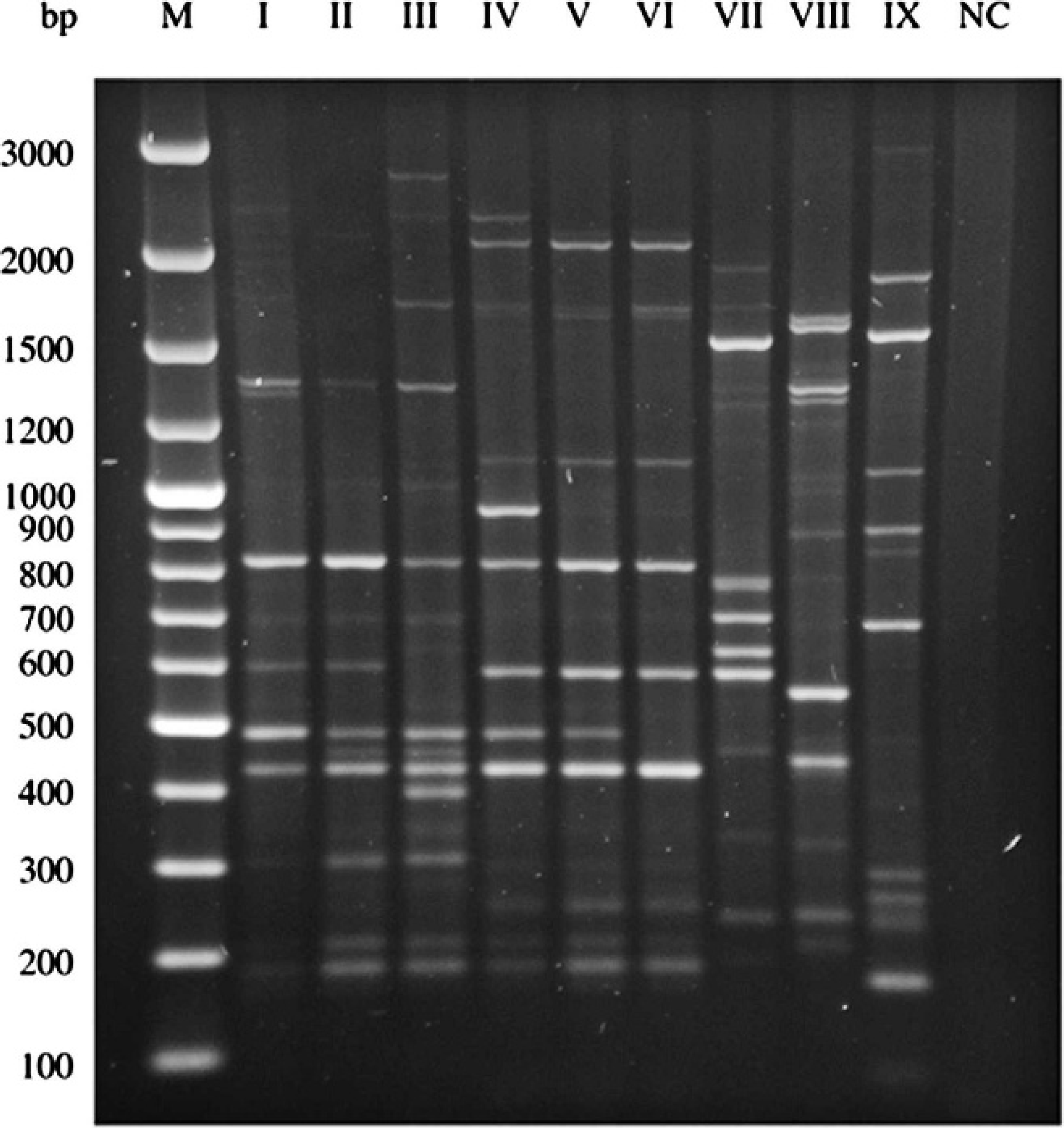
Whole-cell repetitive extragenic palindromic polymerase chain reaction (REP-PCR) profiles of Pasteurellaceae rabbit isolates. Lane 1: molecular weight standard e ; lanes 2–9: REP-PCR types I-IX; lane 10: negative control.
Sequence analysis of the 16S rRNA and rpoB genes
After truncation of primer sequences, the 16S rRNA gene sequencing in most cases resulted in a fragment of 1,364 bp; however, 10 samples gave a fragment of 1,356 bp. All rpoB sequences were the same length of 520 bp. The phylogenetic relationships of the 16S rRNA and the rpoB gene sequences of 41 isolates, representing all observed phenotypes and REP-PCR patterns, are illustrated in Figures 2 and 3, respectively. The investigated strains clustered in a very similar way in both trees.
In the 16S rRNA-based tree, they were grouped in 5 different clusters, A-E (Fig. 2). Most of the samples were located in the highly homogeneous cluster A, which contains the type strains of P. multocida subsp. multocida as well as P. multocida subsp. gallicida, and showed a very high similarity to both type strains with 99.79-99.85% and 99.93-100%, respectively. The rpoB gene provided a higher resolution than 16S rRNA and split the strains located in P. multocida subsp. multocida/P. multocida subsp. gallicida cluster A of the 16S rRNA-based tree in 3 rpoB clusters, A1-3 (Fig. 3). Nine strains were located in cluster A1 and showed 99.23-99.81% sequence similarity to the type strain of P. multocida subsp. multocida and only 96.5-97.3% sequence similarity to the type strain of P. multocida subsp. gallicida, which, together with another strain, formed cluster A2. A further 10 strains built cluster A3 within the rpoB-based tree. In both trees, the same 6 isolates formed cluster
Four isolates sharing identical 16S rRNA gene sequences were placed in cluster C together with the P. canis type strain and differed from it in 1 nucleotide. The same isolates showed identical rpoB sequences and also built cluster C together with the P. canis type strain from which they differed in 8 bp.
The isolates with shorter 16S rRNA gene sequences of 1,356 bp and 1 unique isolate, formed clusters D and E, respectively, which are distant from currently known species within the family Pasteurellaceae. However, the 16S rRNA sequence comparison of isolates from cluster D against GenBank showed 99% similarity to strain CCUG 28028 (accession nos. AF224303 and AF024531), which has not yet been described on the species level. The 16S rRNA gene sequence of the only isolate from cluster E (clinical case 29) matched 99% with Pasteurella sp. MCCM 02539 (accession no. AF224293). Analysis of the rpoB sequences provided a similar picture of a homogenous cluster D comprising the same 10 isolates as in the 16S rRNA-based tree. The strain clinical case 29 formed a monophyletic cluster E linked to Volucribacter psittacicida. No rpoB sequences similar to the strains from clusters D and E in the current study could be found in GenBank. Correlation values of r = 0.98 between REP-PCR types and rpoB gene sequence-based analysis and r = 0.57 for REP-PCR types and 16S rRNA gene sequence-based analysis were observed.
Discussion
To the authors' knowledge, this is the first extended study of phenotypic and genetic diversity within a collection of Pasteurella isolates from rabbits using biochemical tests and molecular methods including REP-PCR and sequencing of 16S rRNA and rpoB genes. Conventional phenotyping demonstrated a high heterogeneity within the strains, resulting in recognition of 12 BTs in the authors' large strain collection (Table 1). The biochemical activities shown in BTs 1–10 have been previously recognized as features characteristic of P. multocida, whereby sorbitol negativity and trehalose positivity is used to differentiate subsp. septica (BT 9) and dulcitol positivity indicates subsp. gallicida (BT 10). 30 High variability in the phenotype of P. multocida is well known and has also been reported within strains isolated from different sources like cattle, pig, poultry, turkey, dog, cat, and bird. 2,3,6,7,17,22,24,26,33,35 Rabbit isolates showing biochemical activity comparable to ODC-negative BTs 3 and 4 and sorbitolnegative BT 7 in the current study have been described previously. 2 Interestingly, the most widespread phenotype of isolates in the investigated animals from the group management breeding system was BT 2 (51%), which has never been observed before in rabbits, and BT 7 (30%). These 2 most distributed types appear to play a role in rabbit pasteurellosis because they are also the 2 most often found BTs from clinical cases. Biochemical type 11 represents P. canis biotype 1, which is usually associated with the oral cavity of dogs and cats or bite wounds from these animals and is rarely found in rabbits. 2 The group of isolates representing BT 12 could not be assigned to any known species; however, the phylogenetic analysis of the 16S rRNA and rpoB gene sequences clearly placed them within the family of Pasteurellaceae (Figs. 2, 3).
Fragment patterns of repetitive extragenic palindromic polymerase chain reaction (REP-PCR) types I-IX.

Strain identification based on fermentation patterns especially on the subspecies level partially diverged from results provided by genetic methods applied in the current study, indicating the problem of proper conventional identification of P. multocida and related species. However, results of 2 different molecular methods, REP-PCR and sequencing of rpoB, were in full accordance to each other and shed some light on the diversity of investigated isolates from rabbits. Similar results were obtained by 16S rRNA gene sequencing; however, the resolution was lower. Whereas all isolates that did not belong to P. multocida clearly separated genetically and phenotypically, strains of P. multocida showed heterogeneity with all approaches.
Eleven strains could clearly be assigned to P. canis, represented by BT 11 and REP-PCR type VII. The 4 strains selected for sequencing out of this group showed a 100% sequence match to each other for 16S rRNA as well as for rpoB genes and formed an identical cluster C in the phylogenetic trees (Figs. 2, 3). Moreover, they differed in 1 position within the 16S rRNA gene and in 8 of 520 within the rpoB from type strain CCUG12400T (originating from dog), suggesting that this variant of P. canis might be specific for rabbits. However, to clarify this statement, further investigation with larger numbers of strains is needed.
A further homogenous group of strains found in both phylogenetic trees was cluster D. All the strains represent REP-PCR type VIII not found in other clusters, and most were phenotypically characterized as BT 12, which was not found within other genotypes. However, 1 strain each of BTs 2 and 6 were present in this genetically homogenous cluster. These 2 BTs could also be found among P. multocida isolates (cluster A). Cluster D did not show significant sequence similarity to any known species of Pasteu-rellaceae. However, the 16S rRNA gene sequence cross-comparison against GenBank revealed that the isolates comprising cluster D are closely related to strain CCUG 28028 belonging to the Pasteurellaceae family, but not classified on the species level, which was obtained from a guinea pig (R. Mutters, personal communications, 2007). This comparison shows that these organisms are adapted to more than 1 host.
Another putative new species is represented by the single isolate of cluster E. Phenotypically, the isolate represents maltose-positive P. multocida (BT 5), but genetically, it is very different and shows the unique REP-PCR profile IX. The isolate clinical case 29 is related to strain MCCM 02539, which is supposed to represent a putative new genus within the family Pasteurellaceae. 32 These strains grouped in cluster D and E were obtained from diseased rabbits, which indicates that the strains are potential pathogens.
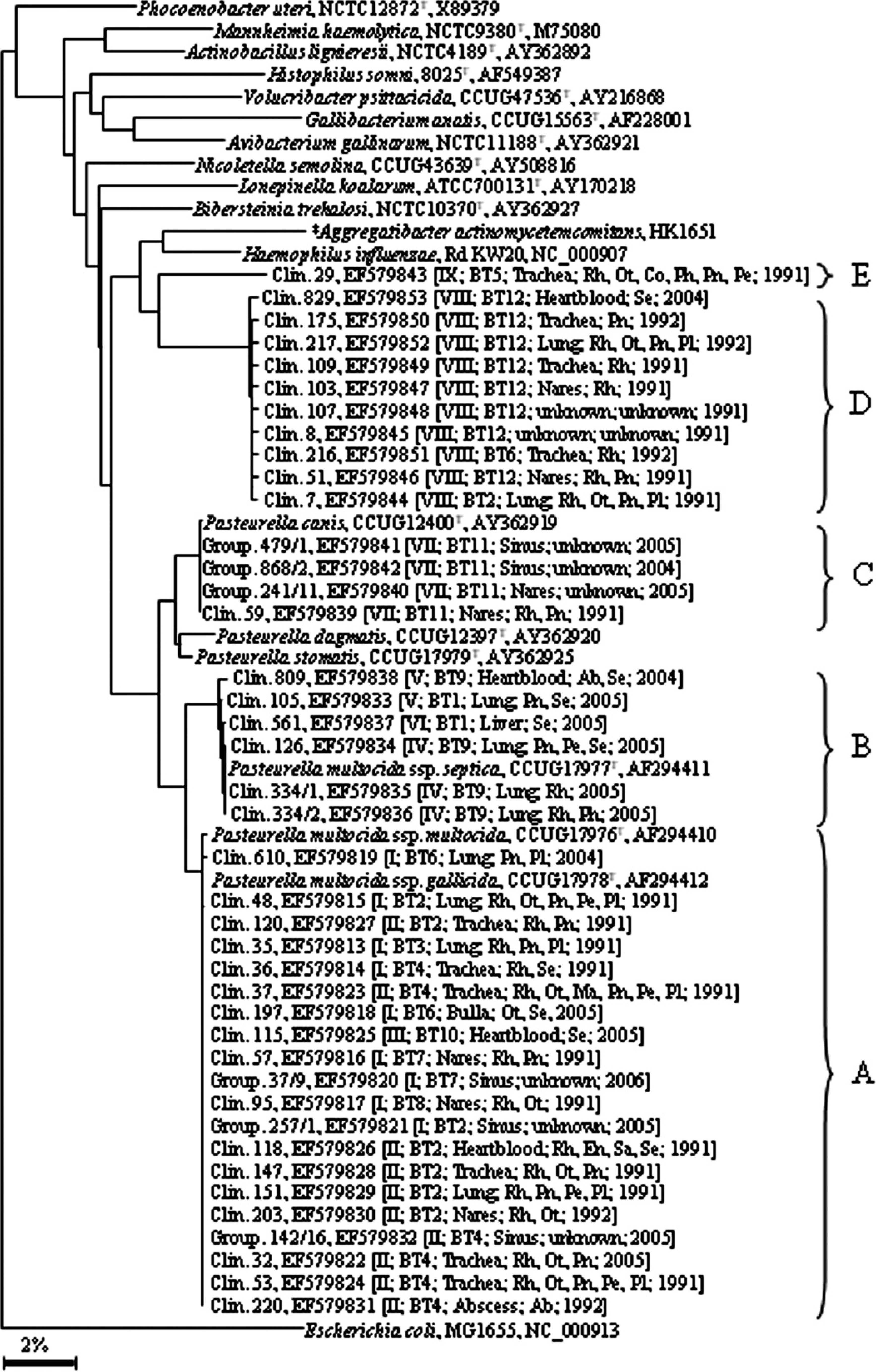
16S ribosomal RNA gene sequence-based phylogenetic tree. Sequences of 41 representative rabbit isolates and GenBank entries of type species of all currently known genera within the family Pasteurellaceae were used (www.oralgene.com). The species names, strain numbers, and accession numbers are listed, and clusters are indicated from A to E. Escherichia coli was used as an outgroup. The scale bar represents percentage of sequence divergence. Clin. = clinical cases; Group. = group management; REP-PCR = repetitive extragenic palindromic polymerase chain reaction types I-IX; BT = biochemical types 1-12; Rh = rhinitis; Ot = otitis media; Pn = pneumonia; Pe = pericarditis; Pl = pleuritis; Co = conjunctivitis; Ph = phlegmon; Ab = abscess; Me = meningoencephalitis; Ma = mastitis; En = endometritis; Sa = salpingitis; Se = septicemia.
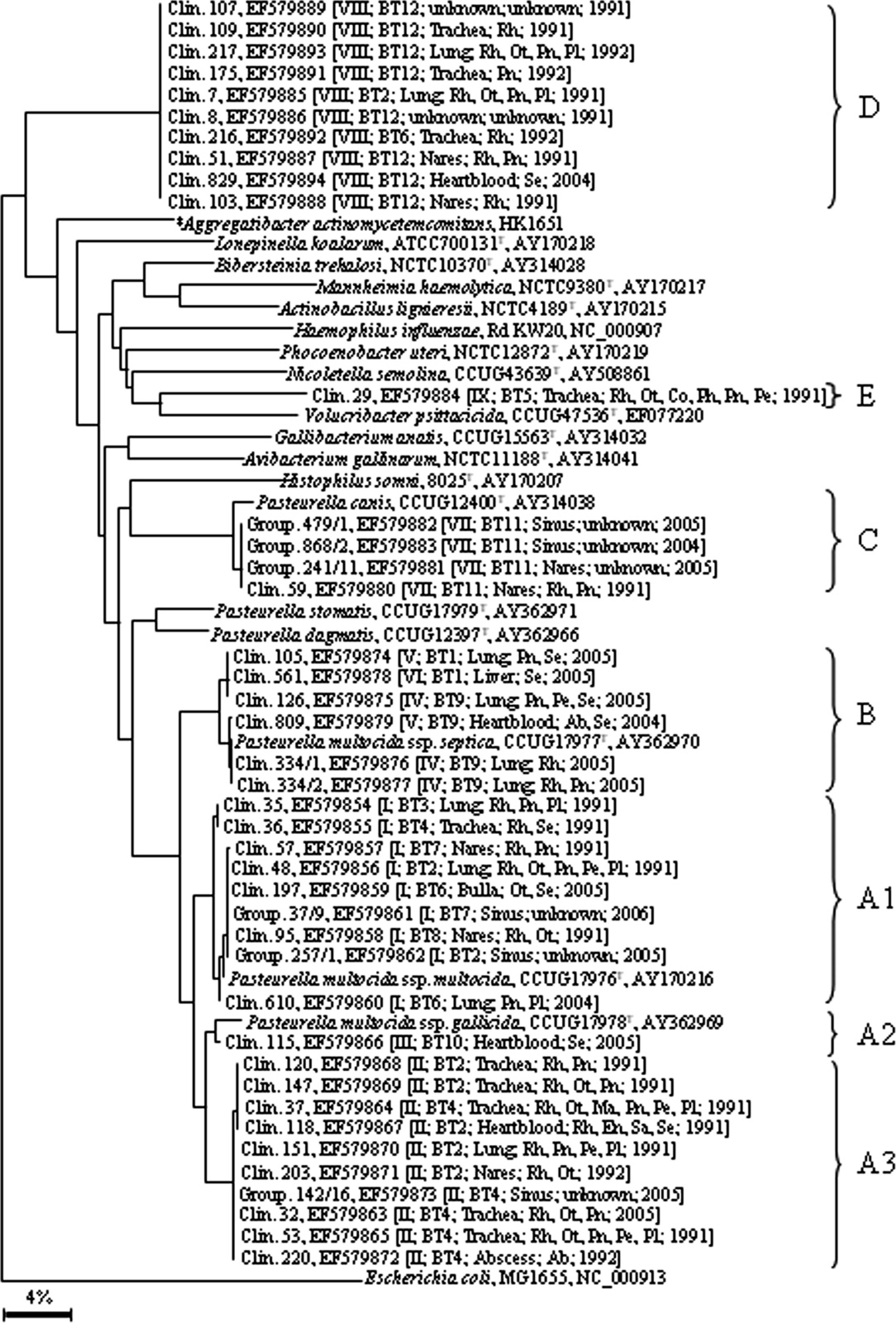
rpoB sequence-based phylogenetic tree. Sequences of 41 representative rabbit isolates and GenBank entries of type species of all currently known genera within the family Pasteurellaceae were used (www.oralgene.com). The species names, strain numbers, and accession numbers are listed, and clusters are indicated from A to E. Escherichia coli was used as an outgroup. The scale bar represents percentage of sequence divergence. Clin. = clinical cases; Group. = group management; REP-PCR = repetitive extragenic palindromic polymerase chain reaction types I-IX; BT = biochemical types 1-12; Rh = rhinitis; Ot = otitis media; Pn = pneumonia; Pe = pericarditis; Pl = pleuritis; Co = conjunctivitis; Ph = phlegmon; Ab = abscess; Me = meningoencephalitis; Ma = mastitis; En = endometritis; Sa = salpingitis; Se = septicemia.
All other strains analyzed phylogenetically clustered around the 3 P. multocida subspecies. There was no congruence between BT and genotype. However, REP-PCR types correlated perfectly with results of phylogenetic analysis of rpoB and to a lesser extent with 16S rRNA gene fragments. In the 16S rRNA-based tree, 20 rabbit isolates belonging to REP-PCR types I-III (Fig. 1) together with the type strain of P. multocida subsp. multocida and P. multocida subsp. gallicida formed the very homogenous cluster A (Fig. 2). In the rpoB-based tree, the same set of subsp. multocida/gallicida strains was split into 3 clusters A1, A2, and A3 (Fig. 3). In cluster A1, 9 rabbit isolates appeared to be very closely related to the type strain of subsp. multocida. Remarkably, they have all been assigned to REP-PCR type I, suggesting that this type is characteristic for subsp. multocida from a rabbit source. One strain identified as typical dulcitol-positive P. multocida subsp. gallicida showed the highest rpoB gene fragment similarity of 98.27% to the type strain of this subspecies. Further, 10 strains that were phenotypically indistinguishable from subsp. multocida belonged to cluster A3, which was clearly separated from all type strains of P. multocida subspecies. The strains are most likely variants of P. multocida that mainly occur in rabbits and might represent a putative subspecies. Moreover, there was again agreement between clustering and REP-PCR types II and III.
The 16S rRNA gene and the rpoB gene fragment analyses allowed classification of sorbitol-positive rabbit isolates, previously phenotypically identified as subsp. multocida (BT 1), and the sorbitol-negative strains representing subsp. septica (BT 9), to subsp. septica with a high sequence identity to the type strain ranging from 99.7% to 100% and 99.0% to 100%, respectively. Phylogenetic analysis confirmed the previous finding that P. multocida subsp. septica forms a distinct cluster from P. multocida subsp. multocida and P. multocida subsp. gallicida, which cannot be separated based on 16S rRNA gene sequences.
26
The REP-PCR patterns IV-VI, which are very similar to each other, were only found in cluster
Interestingly, REP-PCR demonstrated the presence of 1 particular genotype in an integrated organization of breeding and fattening rabbits. With 112 of 123 isolates (91%), REP-PCR type I was widely spread throughout the Swiss population of group management rabbits. The top-down practice with a single breeder delivering animals to all other farmers presumably led to the distribution of this particular strain to different rabbitries. The identification of such a genotype can clarify presumptive route of infection and ease vaccine production. On the other hand, this indicates the normal flora status of rabbits, and once established in an animal, the strains are passed down to the offspring. The isolates from postmortem cases showed a high heterogeneity. Only 75 of the 105 isolates (72%) were grouped as P. multocida subsp. multocida (REP-PCR type I). The involvement of 9 different genotypes from clinical material collected within 2 distinct time periods documents the heterogeneity of Pasteurella isolates within a small population of 300,000 animals kept by 30,000 owners in Switzerland.
In conclusion, P. multocida is the most common member of the family Pasteurellaceae in the population of Swiss rabbits examined in the current study, but other bacteria belonging to the family of Pasteurellaceae have also been found in clinical material. With 82% of all characterized isolates, P. multocida subsp. multocida was the largest group of all found Pasteurellaceae. The P. multocida subsp. septica strains represented a small number of the isolates (3%). Further, 5% of the isolates represent atypical subspecies of P. multocida. In addition, 11 isolates, which could not be assigned to a known species, were identified and might represent a rabbit pathogen. The role of P. canis (5%) in rabbits should be further investigated.
The phenotypical characterization of Pasteurella strains is the most common but not the most suitable tool for identification of P. multocida isolates from rabbits. DNA sequence-based identification using the 16S rRNA and rpoB marker genes clarifies the phylogenetic relationships of the isolates and is more appropriate for identification. Both genes showed a high congruence. Interestingly, REP-PCR typing reflects the sequencing results, and therefore, represents a suitable tool for genetic characterization of rabbit Pasteurellaceae isolates while also allowing epidemiological investigations.
Acknowledgements
The authors are very grateful to L. Bigler, F. Näf, and U. Wullschleger for their cooperation and assistance in providing part of the research material, and to B. Zimmermann for the preparation of the biochemical reagents. The authors also appreciate the help and support of R. Keller, M. Kreienbühl, C. Neff, C. Nievergelt, M. Rusch, and O. Schenker (Institute of Veterinary Bacteriology, NRGK, University of Zurich).
Footnotes
a.
Technical Service Consultants, Heywood, UK.
b.
Oxoid Ltd., Cambridge, UK.
c.
TaqBead™ Hot Start Polymerase, Promega Corp., Madison, WI.
d.
DNA 2720 Thermal Cycler, ABI Prism 3130xl Genetic Analyser; Applied Biosystems, Foster City, CA.
e.
Eurobio, Courtaboeuf, France.
f.
peqGold 100-bp DNA-Leiter plus, PEQLAB Bioechnologie GMBH, Erlangen, Germany.
g.
Alpha Innotech Corp., San Leandro, CA.
h.
GeneCodes Corp., Ann Arbor, MI.
i.
Applied Maths NV, Sint-Martens-Latem, Belgium.
