Abstract
The nested polymerase chain reaction (nPCR) using frozen feather pulp is useful for detecting fowl glioma-inducing virus (FGV), which belongs to the Avian leukosis virus family, and it has recently been suggested that FGV has spread to ornamental chickens kept in Japanese zoological gardens. In the current study, the practicality of using DNA and RNA from dried feather shafts as PCR samples was examined to establish a simple method for tissue preservation. Feather shafts were collected from 7 FGV-positive chickens and stored at room temperature for 30 days. DNA and RNA were extracted from these dried materials. All DNA and complementary DNA (cDNA) prepared from the RNA showed positive results for chicken β-actin and FGV, respectively. Screening for FGV was performed on Japanese fowls kept in zoological garden N. Of the feather shafts collected from 57 birds, 1 sample tested positive for FGV according to PCR of DNA and cDNA samples from the dried feather shafts. This positive bird originated from zoological garden A and had brain lesions suggestive of fowl glioma. The results suggest that DNA and RNA from dried feather shafts can be used in nPCR to detect the FGV genome.
Fowl glioma is histologically characterized by multiple nodular growths of astrocytes with nonsuppurative encephalitis 17 and has been reported in Germany, 2 South Africa, 11 the United States, 12 Denmark, 3 the United Kingdom, 19 and Australia 15 from the 1930s to the 1980s. In 1996, fowl glioma was found in Japanese bantams kept in a zoological garden in Japan. 14 The causal agent of this disease is fowl glioma-inducing virus (FGV), a strain of Avian leukosis virus (ALV; family Retroviridae, subfamily Orthoretrovirinae, genus Alpharetrovirus) subgroup A. 10,18 Avian leukosis virus usually causes lymphoid leukosis in chickens and can be vertically or horizontally transmitted. 5 Currently, there is no treatment for this viral infection. To eradicate ALV infection, chicken flocks including infected birds must be culled. However, this strategy is inappropriate for Japanese fowl for the following reasons: 17 breeds of Japanese fowl are designated as protected species by the Japanese government, and it is laborious to maintain Japanese fowl because of the strict standards for each breed. Because the number of Japanese fowl in Japan is decreasing each year, the detection and elimination of infected birds are the best ways to control FGV infection in Japanese fowl at present.
In a previous report by the authors, 9 a nested polymerase chain reaction (nPCR) using frozen feather pulp was established as a detection assay for FGV, and an etiological study was performed on 131 Japanese fowl in zoological garden A in Japan. The results indicated that 52 chickens (39.7%), including Japanese bantams, were infected with FGV. As zoological garden A is currently a major supplier of Japanese fowl to other domestic zoological gardens, it is suspected that FGV infection has spread to chickens in other zoological gardens in Japan. Although it is necessary to collect samples from many birds in other zoological gardens to screen for and control FGV infection, in general, tissue for nucleic acid extraction should be stored at −80°C as soon as possible after sampling. To collect field samples from widely disparate regions, a simple procedure for tissue preservation is desired. In the current study, to simplify the process of sample preservation, the usefulness of dried feather shafts for nucleic acid extraction was examined.
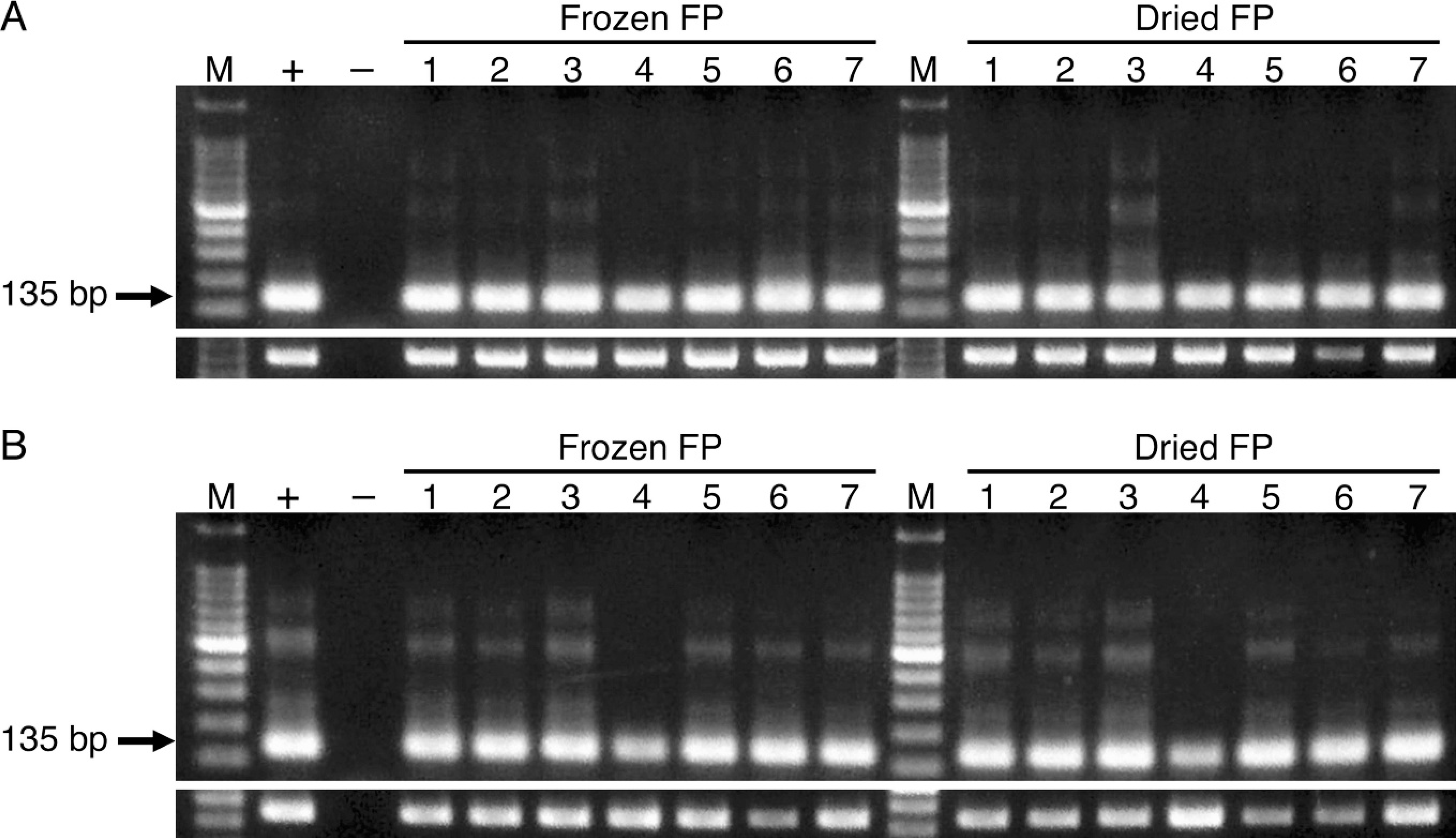
Polymerase chain reaction (PCR) using DNA and RNA from dried feather pulps. A, PCR using DNA samples:
At first, feather shafts containing feather pulp were collected from 7 FGV-positive chickens from zoological garden B, which was surveyed for FGV infection, 8 and were stored at room temperature or −80°C. After 30 days, the frozen feather pulp was squeezed from the shafts, and the dried feather shafts were incubated in typical DNA extraction buffer at 37°C for 16 hours. DNA samples were prepared with standard phenol-chloroform extraction and ethanol precipitation.
The quality of all DNA samples was confirmed with PCR to detect the chicken β-actin gene. The PCR was performed according to a previous method. 8 Briefly, the PCR mixture contained a 10-μl reaction volume including 1 μl of 10× Ex Taq Buffer, a 0.8μl of deoxyribonucleotide triphosphate a (2.5 mM each), 5 pmol of each primer, 0.25 U of TaKaRa Ex Taq HS, a and 10 ng of DNA. The PCR conditions involved denaturation at 98°C for 1 min, followed by 30 cycles of 98°C for 10 sec, 51°C for 15 sec, and 72°C for 45 sec. The PCR products were analyzed by 2.0% agarose gel electrophoresis.
For the RNA extraction, dried feather shafts were incubated in extraction reagent b for 12 hr at room temperature. Frozen feather pulp was squeezed from the shafts and handled in the same manner as the other tissues. Total RNA was extracted according to the manufacturer's instructions. The extracted RNA was treated with DNase. c Reverse transcription (RT) was performed with random hexamers a and reverse transcriptase. d As an internal control for RNA extraction and complementary DNA (cDNA) synthesis, PCR was performed for all cDNA samples using chicken β-actin-specific primers (β-actin 5′: 5′-TATCCGTAAGGATCTGTATG-3′, β-actin 3′: 5′-ATCTCGTCTTGTTTTATGCG-3′). These primers were designed to anneal to sequences in 2 exons on opposite sites of an intron. When template nucleic acid contains only cDNA, a single product sized 338 bp will be observed. On the other hand, 692 bp–sized bands will also be generated when genomic DNA contaminates RT samples.
For detection of FGV, specific nPCR was performed using an established procedure. 7 The first round of the nPCR was performed using the following external primer set: EnvFwd3 (5′-ACGTTGCCGGAATGTGTTGT-3′) and L7Krev (5′-CTATACACACCCTAGTCCC-3′). The internal primers, 5FGOG (5′-TGTAGGCGGGCTCTTGTATT-3′) and 3FGOG (5′-TTGTCGGTCAAGCCTTGCCTT-3′), were designed based on the 3′ untranslated region of the FGV genome and were used in the second round. A 135-bp fragment was amplified as the target sequence. The composition of the first-round PCR mixture was the same as that for chicken β-actin described above, except for the primers. In the second round, 0.2 μl of undiluted first-round PCR product was added to 9.8 μl of the reaction mixture. The first round of the PCR was performed with the following profile: 98°C for 1 min, followed by 25 cycles of 98°C for 10 sec, 60°C for 15 sec, and 72°C for 30 sec using a commercial PCR system e in Max ramp speed mode. In the second round, the number of reaction cycles and the extension time were changed to 30 cycles and 15 sec, respectively. The PCR products were analyzed by 2.0% agarose gel electrophoresis. All PCR reactions included a positive control with template DNA isolated from FGV-infected DF-1 cells, a cell line that originated from chicken embryo fibroblast.
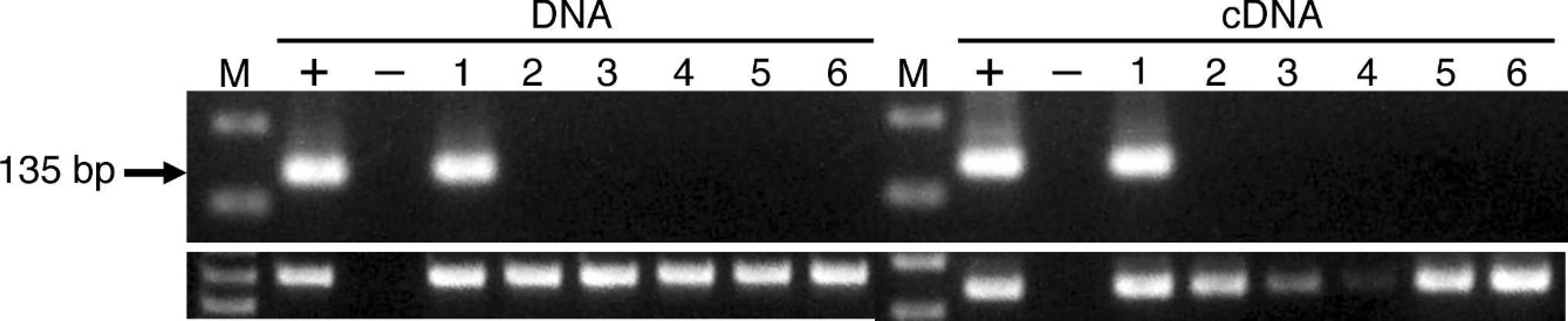
The fowl glioma-inducing virus (FGV) surveillance of chickens kept in zoological garden N.
With PCR using chicken β-actin primers, single bands sized 692 bp were obtained from all DNA samples from the frozen and dried feather shafts of 7 chickens of zoological garden B (Fig. 1A). The extracted RNA samples from the dried feather shafts of sample no. 5 and the frozen and dried feather shafts of sample no. 6 were colored black or brown due to melanin pigment. At first, the cDNA samples made from the stained RNA were negative for chicken β-actin. However, melanin is known to inhibit PCR and/or RT. 6,7 According to the method of a previous report, 6 1 μg of bovine serum albumin a was added to each RT mixture, including the stained RNA, to avoid inhibition of RT. All cDNA samples showed 338 bp–sized single bands by PCR using β-actin primers and positive results for FGV (Fig. 1B).
To confirm whether Japanese fowls kept in zoological garden N, which obtained some Japanese fowls from zoological garden A, were infected with FGV, they were examined by nPCR. Feather shafts collected from 57 birds were stored and transported at room temperature for 3 days. The sample handling and PCR were performed with the same procedures described above. One chicken was positive by nPCR using DNA and cDNA, respectively (sample no. 1; Fig. 2). This PCR-positive bird from zoological garden N had originated from zoological garden A. The bird died after the examination and was necropsied. Nested PCR performed using the brain of the bird also yielded positive results for both β-actin and FGV. The nucleotide sequences of the nPCR products obtained using DNA and cDNA from the feather shafts and DNA from the brain were compared. The sequences of all the 3 samples showed high identity (94%) with each other and the corresponding sequence of the FGV genome (Fig. 3). In addition, a pathological examination revealed that the chicken had brain lesions consistent with fowl glioma (Fig. 4).

The nucleotide sequences of the nested polymerase chain reaction detection region. “DNA” and “cDNA” indicate DNA and complementary DNA from dried feather pulp, respectively. “Brain” denotes the sequence obtained using DNA from the brain. “FGV” indicates the corresponding region of fowl glioma-inducing virus (GenBank accession no. AB112960).
According to a previous report, 1 genomic DNA from feathers that had been kept for 2 weeks at room temperature or for up to 1 month at 4°C were useable in PCR and Southern blotting assays for sex determination and paternity testing of the birds. Marek's disease was detected by PCR using the feather tips of commercial chickens 4 and wild geese. 13 In addition, the provirus of ALV subgroup J was detected by PCR using fresh chicken feathers. 16 The results of the current study reveal that DNA and RNA are well conserved for at least 30 days in dried feather shafts stored at room temperature and suggest that DNA and RNA extracted from dried feather pulp are viable for detection of FGV by nPCR. In conclusion, feather shafts can be collected from chickens in disparate locations without special equipment, reagents, or procedures and used for PCR testing for FGV.
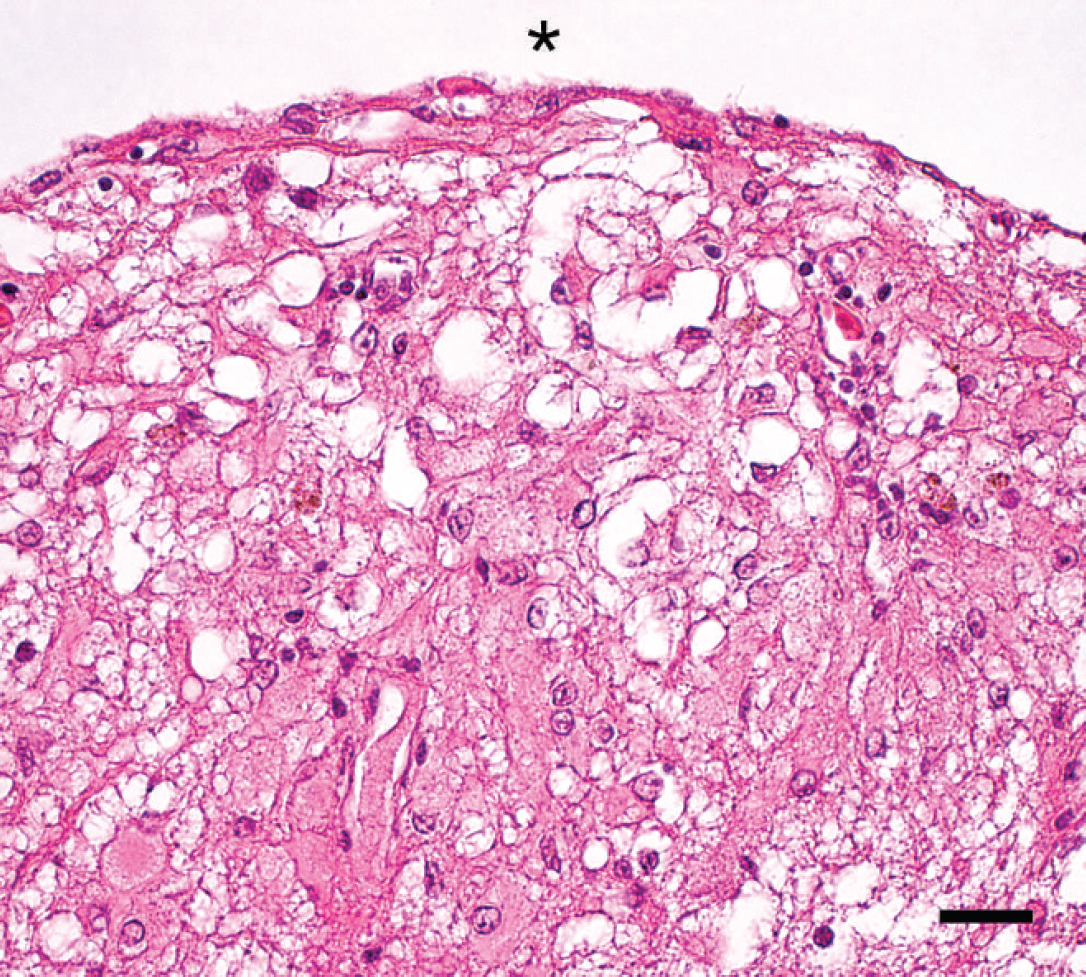
The histopathology of Japanese fowl that was positive for fowl glioma-inducing virus by nested polymerase chain reaction. Focal proliferation of astrocytes under the ependyma. The asterisk shows the lateral ventricle. Hematoxylin and eosin. Bar = 20 μm.
Acknowledgements. This work was supported in part by Grants-in-Aid for Scientific Research (B: 18380179 and B: 20405039) from the Ministry of Education, Culture, Sports, Science, and Technology of Japan, and Grants-in-Aid from the Japan Society for the Promotion of Science Fellows (19004098).
Footnotes
a.
Takara Bio Inc., Otsu, Shiga, Japan.
b.
TRIzol® Reagent, Invitrogen Japan K.K., Tokyo, Japan.
c.
DNA-free™ Kit, Applied Biosystems, Foster City, CA.
d.
SuperScript® II RNase H− Reverse Transcriptase, Invitrogen Japan K.K., Tokyo, Japan.
e.
GeneAmp® PCR System 9700, Applied Biosystems, Foster City, CA.
