Abstract
Generalized tularemia was diagnosed in a vervet monkey (Chlorocebus aethiops) and a patas monkey (Erythrocebus patas), both of which died suddenly in the Szeged Zoo, Szeged, Hungary. Macroscopic lesions in each animal included disseminated, grayish-white foci in the lungs, lymph nodes, spleen, liver, and kidney. All focal lesions were characterized microscopically as purulent to pyogranulomatous to granulomatous inflammation with necrosis. Francisella tularensis subsp. holarctica strains were isolated from tissue samples on modified Francis agar after mouse passage and identified by a commercial carbon-source utilization test and polymerase chain reaction-based amplification and sequencing of a portion of the 16S ribosomal RNA gene.
Tularemia is a life-threatening zoonotic disease caused by the bacterium Francisella tularensis. 5 As few as 10 bacteria are sufficient to cause severe disease and death in susceptible hosts. 4 There are 3 recognized subspecies of F. tularensis: F. tularensis subsp. tularensis, F. tularensis subsp. holarctica, and F. tularensis subsp. mediasiatica (German Collection of Microorganisms and Cell Cultures: 2008.
Available at http://www.dsmz.de/microorganisms/bacterial_nomenclature.php. Accessed September 2008). Although all F. tularensis subspecies are capable of infecting many animal species, their true reservoir is not clearly established. Natural infections with F. tularensis have been reported in a range of vertebrates, including mammals, birds, amphibians, and fish, and in certain invertebrates. 8 The 2 known cycles of tularemia are the terrestrial and aquatic cycles. 5,8,10 In the terrestrial cycle, hares, rabbits, and rodents are the most important mammalian hosts, although ticks, mosquitoes, and horseflies may play a role as arthropod vectors. In the aquatic cycle, beaver, muskrat, and voles serve as main hosts and could shed live bacteria into the environment. Recently, a species of protozoan (Acanthamoeba castellanii) has been found that might be an important environmental reservoir of F. tularensis and might possibly play an additional role in the aquatic cycle. 1 Francisella tularensis can persist for a long time in watercourses and at low temperatures in the terrestrial environment.
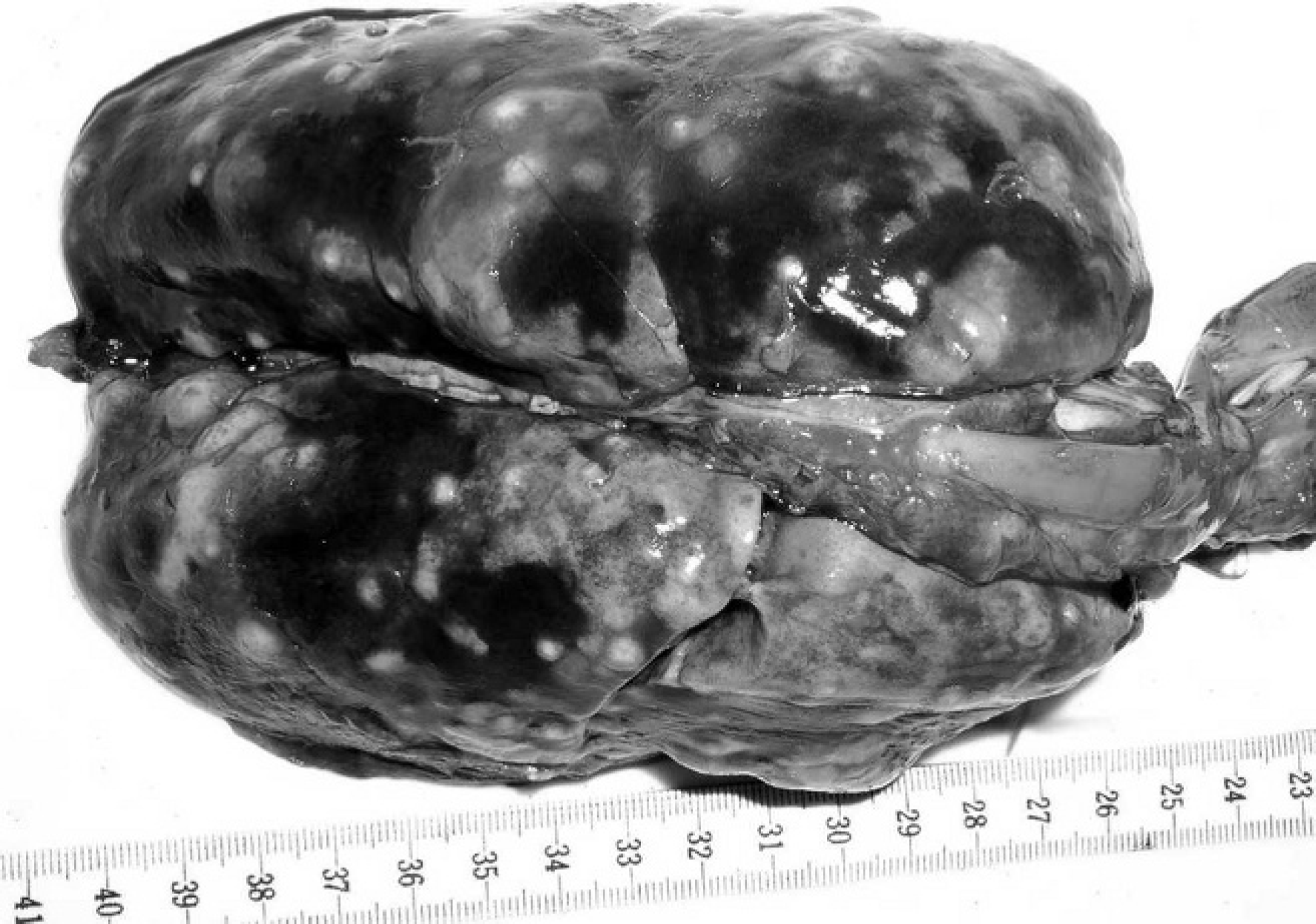
Lung from a vervet monkey (Chlorocebus aethiops) containing numerous granulomas.
Humans are highly susceptible to F. tularensis. The virulence of the organism, dose, and route of exposure are all important factors influencing the clinical form and severity of the disease. 4 Humans can be infected with F. tularensis by several routes, such as being bitten by infected arthropods; handling infectious animal tissues or fluids; having direct contact with or ingesting contaminated water, food, or soil; and inhaling infective aerosols. Generally, the course of clinical disease includes sudden onset of fever, generalized aches, inflammation of the upper respiratory tract with nasal discharge, vomiting, malaise, and anorexia. 4 Seven clinico-pathological forms of tularemia have been described in human medicine: ulceroglandular, glandular, oculoglandular, oropharyngeal, pneumonic, typhoidal, and septicemic. 3,4 The first human tularemia cases were diagnosed in Hungary in 1951, and since then, cases have been appearing yearly. In most cases, there was a history of tick bite and close contact with brown hares, hamsters, or rats.
Tularemia of nonhuman primates and other species is uncommon in European zoos. An outbreak of tularemia affected 18 cynomolgus monkeys (Macaca fascicularis) in the German Primate Center in 2005, 7 and solitary cases were recorded in a golden-headed lion tamarin (Leontopithecus chrysomelas) 6 and a common marmoset (Callithrix jacchus) 10 in Switzerland. The present study summarizes the postmortem lesions and the results of the bacteriologic examination of a vervet monkey (Chlorocebus aethiops) and a patas monkey (Erythrocebus patas) that died suddenly due to tularemia at Szeged Zoo in October 2003.
An adult, female vervet monkey and a 2-year-old male patas monkey were kept in the same large open-air enclosure with a closed shelter at the Szeged Zoo (Csongrád County, Hungary). A total of 4 patas monkeys and 6 vervet monkeys were kept in this enclosure at the time. The 2 affected animals were born and raised in Szeged Zoo. After showing clinical signs of depression for 2 days, the vervet monkey died at the end of October 2003. The patas monkey died unexpectedly 10 days later and exhibited no prior clinical signs. The animals were not treated because of the mild nonspecific clinical signs or lack of clinical signs followed by sudden death. Other animals and in-contact humans did not show any clinical signs of concurrent illness and did not receive any prophylactic treatment.
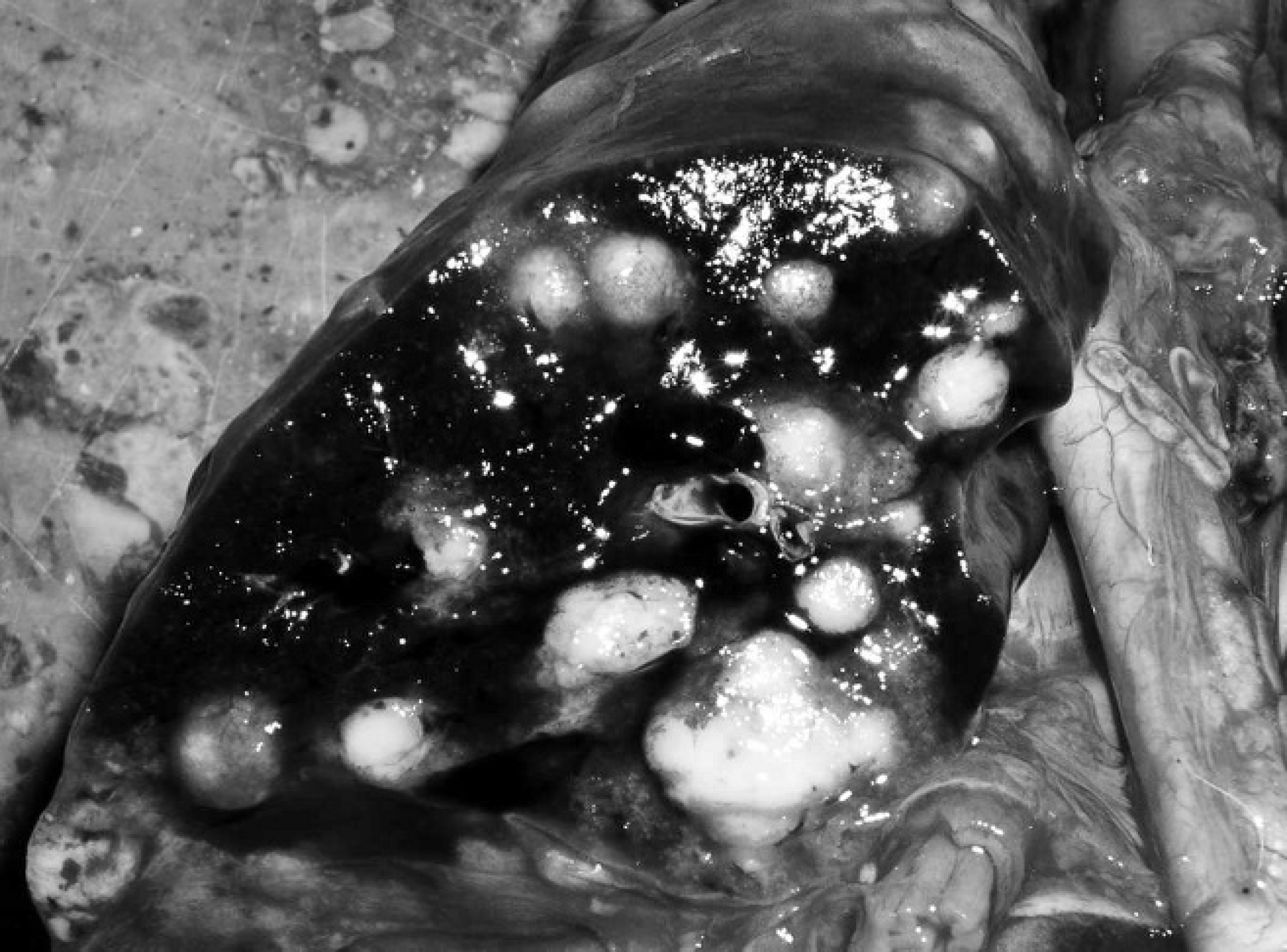
Spleen from a patas monkey (Erythrocebus patas) with necrotic foci.
The carcasses were necropsied under appropriate bio-safety conditions. Tissue samples were fixed in 10% neutral buffered formalin, processed routinely, and embedded in paraffin. Sections were stained with hematoxylin and eosin, coverslipped, and examined by light microscopy.
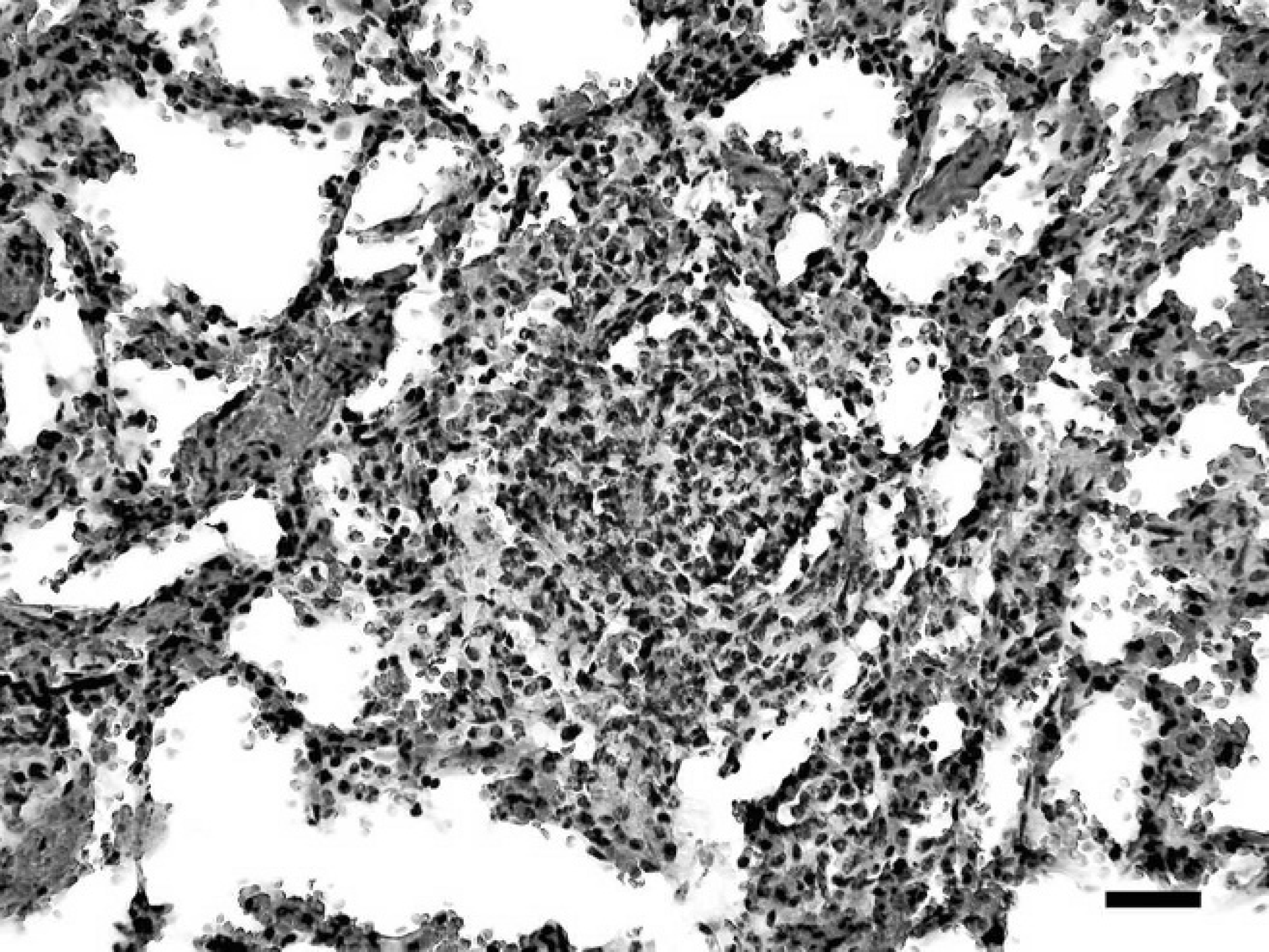
Lung from a vervet monkey (Chlorocebus aethiops) with focal neutrophilic inflammation surrounded peripherally by fibroblasts and mononuclear cells. Hematoxylin and eosin stain. Bar = 50 μm.
Bacteriologic culture was performed from the lungs, spleens, and livers of both animals on blood agar and MacConkey agar. Tissue samples (approximately 1 g) excised from lesions of both monkeys were homogenized individually and suspended in 2 ml of sterile, normal saline. Specific pathogen-free laboratory mice (Naval Medical Research Institute mouse, approximately 20 g) were injected subcutaneously with 1 ml of suspension. The mice were euthanized when symptoms of tularemia appeared. Heart, blood, and bone marrow samples were spread on modified Francis agar plates (chocolate agar plate 2 containing 1% glucose and 0.1% cysteine). Culture plates were incubated at 37°C in 6.5% CO2 for 5 days and checked daily for growth of F. tularensis. Each strain was subcultured 3 times to obtain pure cultures. Culture, morphologic, and biochemical characteristics were examined using standard methods. 2 The utilization of 95 carbon sources was examined using a commercial carbon-source utilization test a following the manufacturer's instructions with minor modifications. All procedures were performed under biosafety level 3 conditions.
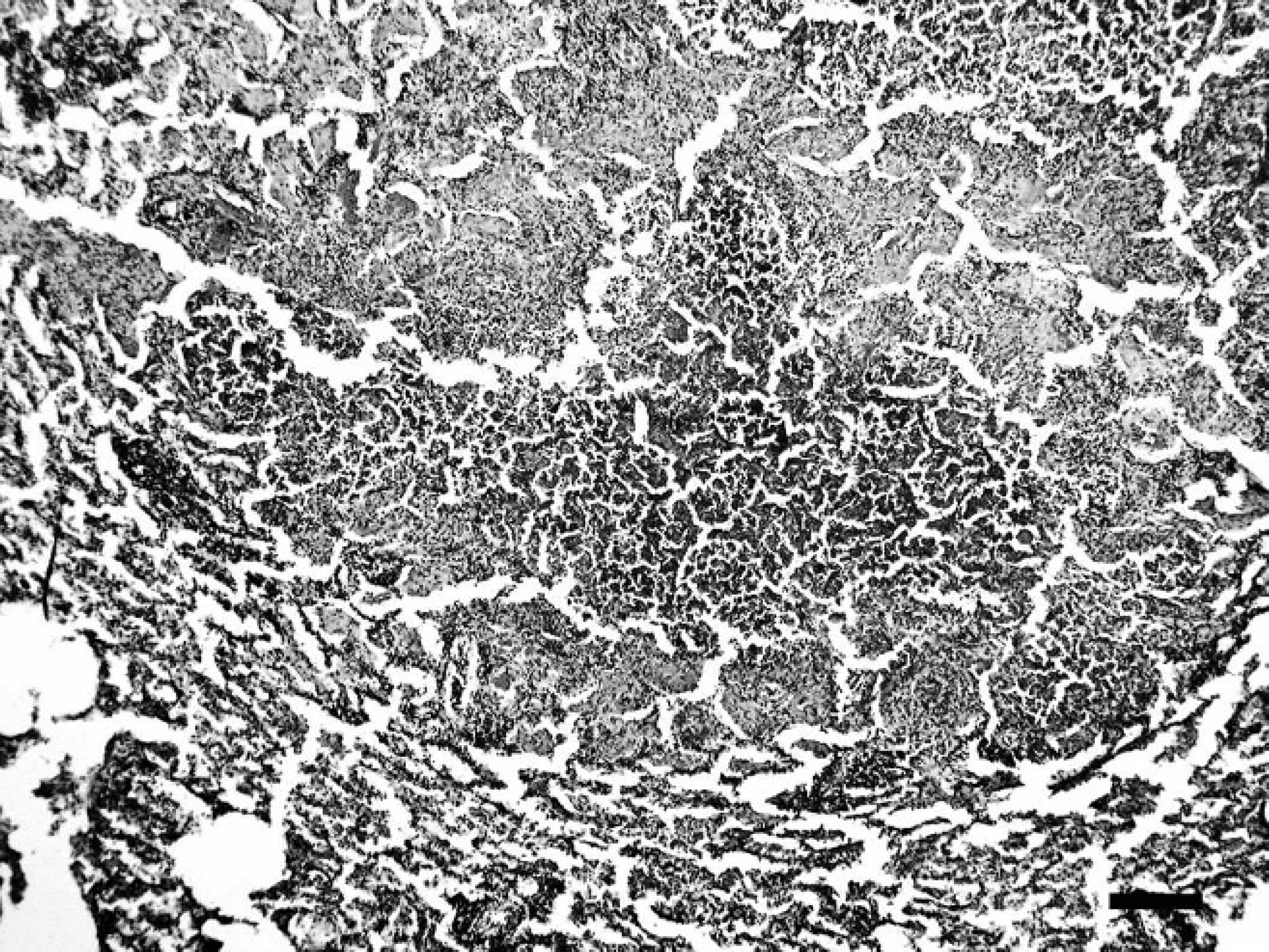
A typical granuloma in the lung of a vervet monkey (Chlorocebus aethiops) with a central area of inflammatory cells and caseous necrosis. Hematoxylin and eosin stain. Bar = 250 μm.
DNA was extracted from 1 distinct colony of each strain with a commercial DNA isolation kit b according to the manufacturer's recommendations for Gram-negative bacteria. A part of the 16S ribosomal RNA (rRNA) gene was amplified using a polymerase chain reaction (PCR) system with the following primer pair: 91E: 5′-TCAAAGGAATT-GACGGGG-3′ and 13B: 5′-CCGGGAACGTATT-CACCG-3′. 11 All PCR reactions were performed in 50 μl total volume, containing 10 to 100 ng target DNA, 1 μl commercially purified c water, 5 μl Taq buffer (NH4/2SO4), d 4 μl MgCl2 (25 mmol), d 5 μl deoxyribonucleotide triphosphate (dNTP; 2 mmol), d 1 μl of each primer (25 pmol/μl), and 0.33 μl Taq polymerase (5 unit/μl). d The PCR reaction was performed in a commercial thermal cycler. e The PCR consisted of initial denaturation for 3 min at 95°C followed by 39 amplification cycles of denaturation for 30 sec at 93°C, primer annealing at 49°C for 30 sec, and extension at 72°C for 1 min. The final extension step was performed for 5 min at 72°C. After amplification, 10 μl of each reaction mixture was subjected to electrophoresis in a 1% agarose gel, and the amplified gene products were visualized with ultraviolet light after ethidium bromide staining. The PCR products were isolated from the agarose gel with a commercial gel extraction kit, b and direct cycle sequencing was performed on an automated DNA sequencer in both directions using the same primers used in the PCR. f
The diagnosis of tularemia was confirmed by the recovery of a strain showing typical cell and colony morphology, biochemical characteristics, carbon utilization pattern, PCR amplification, and F. tularensis sequence data of the resulting amplicons. In both monkeys, macroscopic postmortem findings included numerous, purulent, 1–5 mm, grayish-white foci surrounded by dark, hyperemic areas throughout the lung parenchyma (Fig. 1). The lymph nodes were enlarged and contained several grayish-white foci measuring 1 mm in diameter. The spleens were dark, enlarged, and contained multifocal necrotic foci (Fig. 2). The livers were friable and mildly enlarged. In addition, the liver of the vervet monkey also contained multiple grayish foci measuring 1 mm in diameter. Similar foci were found in the kidneys of the patas monkey. Histopathological examination of the lung lesions revealed a severe, multi-focal, purulent, pyogranulomatous to granulomatous pneumonia throughout the lung parenchyma (Figs. 3, 4). Typical pyogranulomas were demarcated by connective tissue and consisted of a central necrotic area surrounded by hemorrhages and large numbers of histiocytes and fewer neutrophils (Fig. 4). Spleens and lymph nodes of both monkeys had granulomatous inflammation with additional foci of necrosis in the lymph nodes. Granulomatous inflammation was also present in the liver of the vervet monkey, while a multifocal pyogranulomatous tubulonephritis was observed in the kidney of the patas monkey.
Bacteria showing F. tularensis-like colony morphology were isolated in pure culture from both monkeys. Grayish-white colonies, 1 mm in diameter, could be seen after 4 days of incubation. These bacteria were Gram-negative, small (<1 μm), nonmotile coccobacilli. They were oxidase negative, had weak positive catalase activity, and were not able to use glycerol. Both strains were identified as F. tularensis with the commercial carbon-source utilization test. Because these organisms were unable to use glycerol as a sole carbon source, the strains were identified as F. tularensis subsp. holarctica. The bacteriologic diagnosis was confirmed by PCR amplification and sequencing of the partial 16S rRNA gene amplicons. The resulting sequences showed 100% identity with the F. tularensis subsp. holarctica (GenBank accession no. CP000803.1) and 99% identity with the F. tularensis subsp. tularensis (GenBank accession no. CP000608.1) 16S rRNA sequences deposited in GenBank.
The presence of tularemia in a vervet monkey and a patas monkey in a zoo was an unusual finding. Although tularemia is not an uncommon disease in Hungary, reservoirs of infection are found in the free-ranging wildlife population. During the past decade (1998–2008), the number of reported human cases of tularemia have ranged between 2 and 5 per year in Csongrád County (∼∼425,000 inhabitants; http://www.oek.hu/).
Although lesions were present in multiple organ systems, the most severe lesions involved the lungs and regional lymph nodes. This pathological pattern suggests that the most likely route of infection may have been through inhalation. Zoo primates regularly hunt and consume small prey species, such as rodents and birds that enter their enclosures. This could be a potential source for contracting tularemia from freeranging wildlife. Furthermore, the heavily forested territory of Szeged Zoo may increase the risk of harboring infected wildlife. The narrow time interval between the 2 deaths suggests that there could have been a common source of infection; however, no further evidence of tularemia has been seen at the zoo since this outbreak. Finally, the current report indicates that infected nonhuman primates may be potential sources of zoonotic disease for animal keepers and visitors, and appropriate hygienic measures should be taken.
Footnotes
a.
MicroLog™ MicroStation™ System, GN2 Microplates; Bio-log Inc., Hayward, CA.
b.
QIAamp DNA Mini Kit, QIAquick Gel Extraction Kit; Qiagen Inc., Valencia, CA.
c.
Milli-Q, Millipore Corp., Billerica, MA.
d.
Fermentas Inc., Burlington, Canada.
e.
Biometra TPersonal Thermal Cycler, Biometra GmbH, Göttingen, Germany.
f.
ABI 373A, Applied Biosystems, Foster City, CA.
