Abstract
A total of 745 slaughtered pigs were examined during routine meat inspection for suspected tuberculous lesions. Specimens from suspected lesions were collected for conventional mycobacteriologic examinations. Suspected mycobacterial colonies were subjected to molecular typing based on the Mycobacterium species-specific intergenic spacer (IGS) target. The study resulted in detection of suspected lesions in 110 (14.8%) carcasses, from which only 67 specimens produced suspected mycobacterial colonies. Conventional examination was only able to identify 56 isolates as Mycobacterium species, which was confirmed by polymerase chain reaction amplification of the IGS target. Interestingly, out of these, 18 and 12 isolates were Mycobacterium tuberculosis and Mycobacterium bovis, respectively. Sequence analysis of IGS resolved the identities of 10 of the 11 conventionally unidentified isolates as being 4 different nontuberculous Mycobacterium species. The last isolate was proposed as a non-Mycobacterium species and was confirmed by its identification as Rhodococcus equi based on the 16S ribosomal DNA sequence analysis. The study described the isolation of Mycobacterium tuberculosis from pigs and revealed high burden of infection with both tuberculous and nontuberculous mycobacterial species among pigs in Egypt. In addition, the study showed the usefulness of IGS sequence analysis as a reliable molecular tool that would be useful for further epidemiologic and public health studies.
Keywords
Introduction
Economic losses due to swine mycobacteriosis, erroneously described as swine tuberculosis, are not only restricted to the condemnation of pig meat, head, and visceral organs at abattoir inspection points due to tuberculous lesions but also involve losses among cattle due to bovine tuberculosis caused by dissemination of Mycobacterium bovis in the environment by infected pigs. 13,21 In addition, the dissemination of nontuberculous Mycobacterium species can sensitize cattle, with the subsequent nonspecific reaction to tuberculin skin test and losses from destruction of false-positive reactors. 16,20,23
On the other hand, the increasing reports of several types of mycobacterial infections among Human immunodeficiency virus–infected and immunocompromised patients 2,12,27 has drawn attention to animals as the source of these Mycobacterium species to humans. Previous studies have shown that pigs are among the possible sources of mycobacterial infections to humans and animals. 1,11,21 Nevertheless, scarce data are available about the types and nature of mycobacterial infection in pigs.
Therefore, detection and identification of the causative agent of swine mycobacteriosis is an important asset for veterinary public health and epidemiologic studies of zoonotic mycobacteriosis. Phenotyping, based on acid-fast staining and standard culture and biochemical analysis, represents the main tool for differential identification of members of genus Mycobacterium in most clinical laboratories. 8 However, these conventional methods are time consuming and are difficult to assimilate into precise identification of closely related species and difficult to identify species. 10,26 Genotypic assays that use specific genetic elements have shown promise as alternative diagnostic tools. 10,28 Molecular identification of Mycobacterium species has 2 primary advantages compared with phenotypic identification: rapid turnaround time and improved accuracy. 4,26 Sequence-dependent identification has been shown to be an especially effective molecular tool that provides rapid and accurate differential identification of Mycobacterium species. 3,18 Most molecular approaches have focused on the conserved 16S small subunit ribosomal DNA (rDNA) sequence. 26 The 16S rDNA gene has been used extensively for identification and phylogenetic studies of Mycobacterium species. 5 However, the fact that the product of the 16S rDNA gene serves a vital function in the bacteria makes the frequency of permissible mutations in this gene inherently limited. This limitation results in the presence of identical or highly homogenous 16S rDNA sequences among some of the Mycobacterium species, making the differentiation of some closely related species difficult. 9 Alternatively, sequence analysis of the intergenic spacer (IGS) between the small (16S) and large (23S) subunit rRNA genes has been successfully used to differentiate among closely related Mycobacterium species. 18,24
The current study was undertaken to identify different Mycobacterium species associated with swine mycobacteriosis in pigs raised under poor hygienic conditions that could represent a potential public health concern. Genotypic assay using IGS-based sequence analysis was implemented to detect mycobacterial infections in suspected specimens and to identify the difficult to identify isolates.
Material and methods
Animals and postmortem examination
During the routine meat inspection of slaughtered pigs at Basatin Abattoir, the central abattoir for pig slaughter in Egypt, 745 pigs were randomly selected during the period of April 2004 to October 2005 and examined for suspected tuberculous lesions. During postmortem examination, retropharyngeal, submaxillary, parotid, bronchial, mediastinal, mesenteric, and prescapular lymph nodes were examined visually and by palpation in situ. Any abnormalities in size, shape and texture of examined lymph nodes were considered suspected lesions and were collected aseptically for further investigation.
Conventional bacteriologic examination
Tissue specimens from suspected lymph nodes were homogenized in a tissue-grinding mortar with 5 ml of sterile saline for isolation of Mycobacterium species. Briefly, tissue homogenates were centrifuged at 3,000 × g for 15 min. The supernatant fluid was discarded, and the sediment was treated with 2 ml of 4% sulfuric acid for 15 min, and then washed twice with sterile saline and centrifuged at 3,000 × g for 15 min. The sediment was then neutralized with 4% NaOH containing phenol red indicator and immediately inoculated onto Lowenstein-Jensen (L-J) media a and incubated at 37°C for up to 8 weeks. Smears of suspected sediment were stained with Ziehl-Neelsen stain to assess acid fastness and morphology of bacteria. Colony morphology and pigment production under dark and light conditions were examined. Conventional biochemical tests were performed as previously described 8 and included niacin accumulation, nitrate reduction, Tween 80 hydrolysis, urease activity, iron uptake, tolerance to 5% sodium chloride, growth on thiophene-2-carboxylic acid hydrazide (T2H), and ability to grow on MacConkey agar.
Molecular identification
DNA extraction and target amplification. Crude DNA was extracted from suspected colonies as described previously 22 and purified using the QIAamp Blood Kit b according to the manufacturer's recommendations. The hypervariable region of the IGS target was amplified from suspected isolates using the previously described panmycobacterium primer set ITS-A1 and ITS-A6. 18 For suspected nonmycobacterial isolates, the hypervariable 500 base pair bp segment of the 16S rDNA was amplified using the previously described primer set. 7 In the polymerase chain reaction (PCR) assay, 5 μl of template DNA (10 ng/μl) in a total reaction volume of 50 μlto include PCR buffer [20 mmol Tris-HCL (pH 8.4) and 50 mmol KCl], 0.1 mmol each of dNTP (deoxyri-bonucleotide triphosphate), 1.5 mmol of MgCl2, 0.3 μmol of each primer, and 1.5 U of RED Taq DNA polymerase c were used. The PCR amplification was performed in a Techne thermocycler model TC-312 d starting with an initial denaturation step at 95°C for 10 min, followed by 35 cycles where each cycle consisted of a denaturation at 95°C for 1 min, an annealing at 64°C for 30 sec, and an extension step at 72°C for 1 min. The PCR products were loaded on 2% agarose gel and visualized using a UVP gel documentation system e according to the manufacturer's instructions.
Sequence analysis. The PCR products of selected isolates were purified from gel using the OMEGA gel extraction kit. f Purified PCR products were sequenced for either IGS or 16S rDNA sequences using the above forward and reverse amplification primers at the Molecular Biology Core Laboratory, Egyptian Institute for Biological Products and Vaccine Production (Cairo, Egypt). Sequence analysis was carried out using both the custom MycoAlign database 18 and BLAST (Basic Local Alignment Search Tool; http://www.ncbi.nlm.nih.gov/BLAST/) using the GenBank database.
Results
Detection of mycobacterial infection
Suspected tuberculous lesions were detected in 110 (14.8%) of 745 examined swine carcasses. Of these, suspected mycobacterial isolates were detected in 49 (44.5%), 67 (61%), and 66 (60%) cases based on positive acid-fast bacillus (AFB), L-J cultures, and PCR-based genotyping methods, respectively.
Phenotypic versus genotypic typing of suspected mycobacterial infection
Of the 67 suspected mycobacterial colonies, conventional phenotyping was able to identify only 56 isolates as Mycobacterium species; 11 isolates were difficult to identify and were described as unidentifiable Mycobacterium species. Interestingly, out of the conventionally identified species, 18 and 12 isolates were Mycobacterium tuberculosis and M. bovis, respectively (Table 1). Genotypically, based on PCR amplification of the Mycobacterium species-specific IGS target, 66 isolates were typed as Mycobacterium species, including 10 of the 11 conventionally unidentified isolates.
. Conventional identification of mycobacterial isolates from suspected tuberculous lesions of slaughtered pigs.
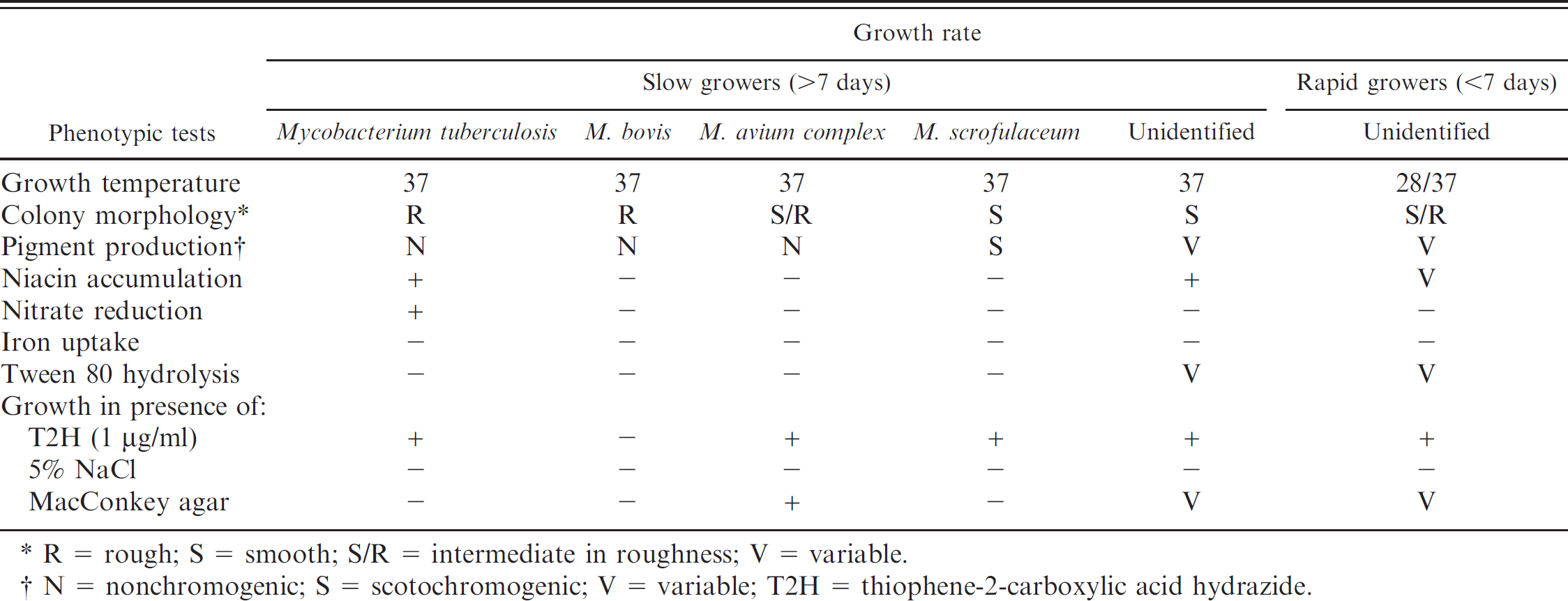
R = rough; S = smooth; S/R = intermediate in roughness; V = variable.
nonchromogenic; S = scotochromogenic; V = variable; T2H = thiophene-2-carboxylic acid hydrazide.
Molecular identification of conventionally unidentifiable isolates
Of the 11 conventionally unidentified isolates, 10 isolates were positive for the IGS target and typed as Mycobacterium species, and 1 isolate was negative for IGS target and proposed as non-Mycobacterium species (Fig. 1). Sequence analysis of the IGS target from suspected mycobacterial isolates identified them as M. chelonae (n = 5), M. porcinum (n = 2), M. gallinarum (n = 2), and M. simiae (n = 1) using both MycoAlign and BLAST tools. Sequence analysis of the universal 16S rDNA target of the suspected nonmycobacterial isolate identified it as Rhodococcus equi using the BLAST search tool based on the GenBank database (Table 2).
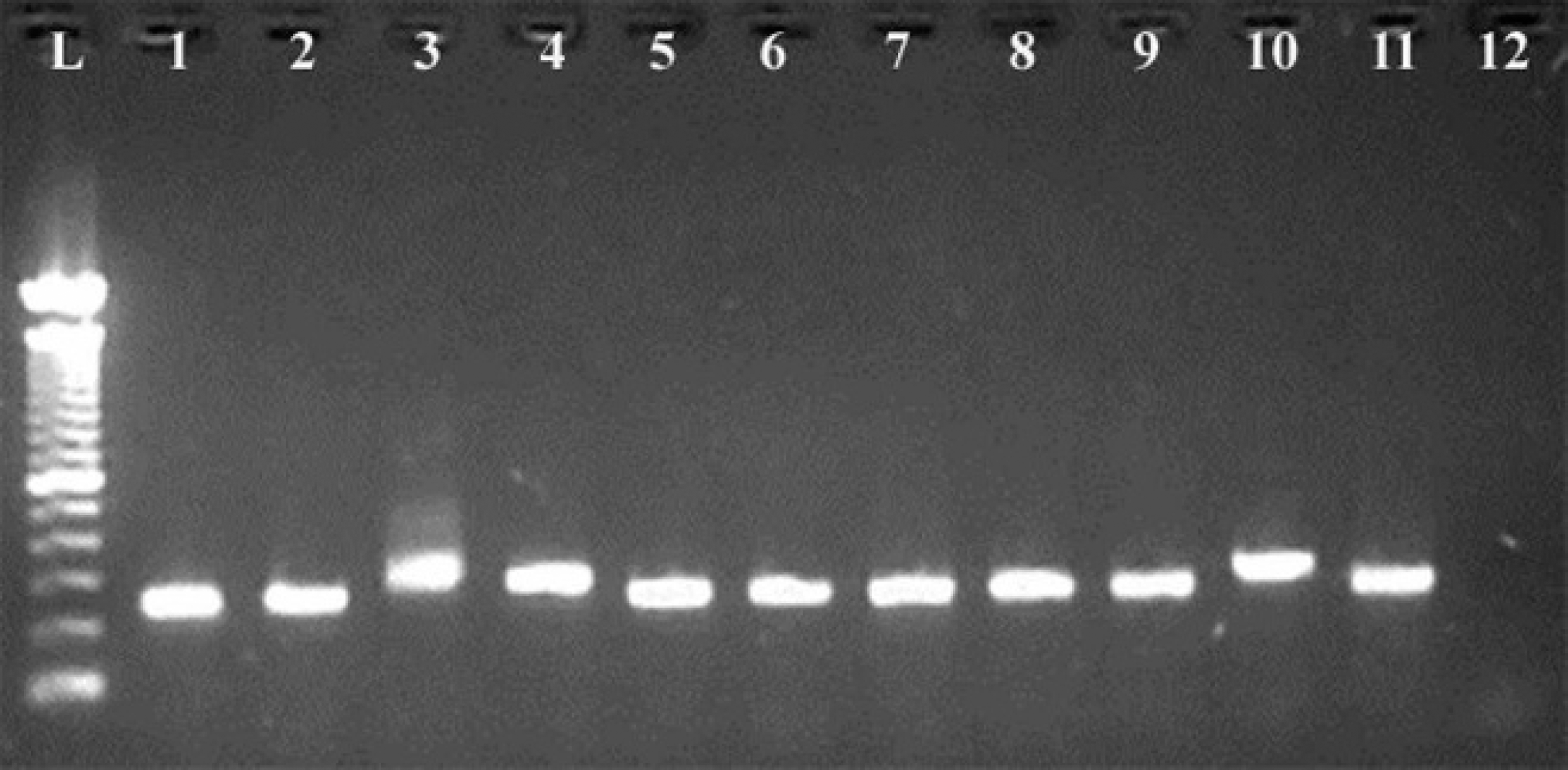
Amplification of intergenic spacer (IGS) target from the 11 conventionally unidentified isolates. Lane L: 100 base pair DNA ladder; lane 1: Mycobacterium bovis AN-5 positive control; lanes 2–11: confirmed Mycobacterium species by positive Mycobacterium species-specific IGS amplicons (230–320); lane 12: confirmed non-Mycobacterium species by absence of the Mycobacterium species-specific IGS amplicon.
Discussion
Although recognition of acid-fast bacilli in smears is the most rapid and cost-effective screening method, it lacks the required sensitivity; therefore, subsequent culture of suspected cases is required. 26 This was confirmed by the increase in the number of suspected mycobacterial infections from 44.5% (49/110), based on acid-fast staining, to 61% (67/110) by culturing from suspected lesions.
Of the 67 suspected mycobacterial colonies, conventional phenotyping confirmed and identified only 56 cases as Mycobacterium species (Table 2), whereas 11 cases were classified as unidentifiable Mycobacterium species. The increasing number of newly defined mycobacterial species and the difficult to identify variants of known species represent a significant challenge for conventional approaches 6,17 and explain the inability of the conventional phenotypic tests used in the current study to reach definite identification of a number of suspected isolates. The use of more conventional phenotypic tests could have resulted in more mycobacterial species being identified. The identities of the 11 conventionally unidentified isolates were resolved by using a sequence analysis-based differential identification assay based on the Mycobacterium species-specific IGS target from suspected mycobacterial isolates and the universal 16s rDNA sequence from the suspected non-Mycobacterium species. Although not a common finding, the isolation of Rhodococcus equi from pigs, as documented in the current study, has been reported previously as one of the nonmycobacterial agents found in soil that frequently infects pigs and results in similar localized lesions in the submaxillary lymph nodes. 14,15
. Conventional and molecular identification of suspected colonies.*
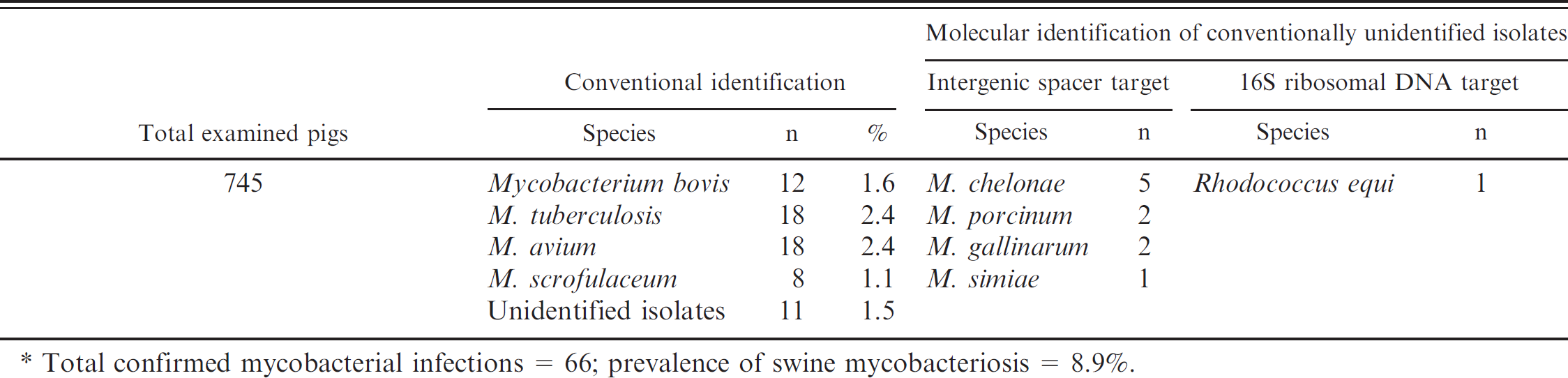
Total confirmed mycobacterial infections = 66; prevalence of swine mycobacteriosis = 8.9%.
Previous studies have identified pigs as a possible potential reservoir for many mycobacterial infections to other farm animals and humans. 11,16 This assumption would intensify the importance of the high burden of swine mycobacterial infection in Egypt (8.9%) as revealed in the current study. This prevalence is unexpectedly high compared with the 0.5% prevalence in Europe, 11 considering the fact that it is based on the isolation of Mycobacterium species only from pigs with suspected tuberculous lesions. The prevalence could be higher if all examined animals were subjected to bacteriologic evaluation. Previous studies have documented the isolation of different pathogenic mycobacterial species from healthy slaughtered swine without any apparent lymph node or tissue lesions. 16,25,29 An interesting finding of current study was the multiplicity of the identified mycobacterial species and particularly the high incidence of M. tuberculosis (18/66), M. bovis (12/66), and M. avium complex (18/66) compared with the Europe study, 11 where the vast majority (91/92) of the agents isolated and genotyped were M. avium. Animal infection with M. tuberculosis, although not common, has been described among domestic and wild animal species most frequently living in close, prolonged contact with humans. 19 The high prevalence of both tuberculous and nontuberculous mycobacterial infections among pigs, as revealed in the current study, could be attributed to the poor hygienic standards of pig farming in Egypt, where feeding pigs on household and hospital waste as well as waste from animal and chicken farms is a common practice.
In conclusion, although the findings of the current study may indicate pigs as a possible source of infection for other hosts, including humans, the evidence for this is weak at best. In addition, achievement of definite identification of suspected isolates only after implementing molecular assays denotes the usefulness of this tool as an important asset for epidemiologic and veterinary public health studies.
Footnotes
a.
Biolife Italiana Srl, Milano, Italy.
b.
Qiagen Inc., Valencia, CA.
c.
Sigma-Aldrich, St. Louis, MO.
d.
Techne, Duxford, Cambridge, UK.
e.
UVP LLC, Upland, CA.
f.
OMEGA BIO-TEK, Doraville, GA.
