Abstract
Fimbriae and enterotoxins are major virulence factors associated with enterotoxigenic Escherichia coli (ETEC). In this study, 3 sets of multiplex polymerase chain reaction (mPCR) assays targeting fimbriae, enterotoxins, and other adherence factors were developed for detecting ETEC. A total number of 188 E. coli field isolates were examined, and percentages of E. coli strains carrying each virulence factors were as follows: F4 (7.45%), F5 (29.79%), F6 (6.38%), F18 (15.43%), F41 (3.72%), STa (10.11%), STb (20.74%), LT (9.57%), Stx2e (2.13%), EAST1 (42.02%), F1 (67.55%), AIDA-I (2.66%), and pAA (7.45%). Of the 188 E. coli field isolates examined, 25.53% were found to be pathogenic ETEC, having both fimbriae and enterotoxins. However, the ratio increased to 44.68% when the presence of other adhesins was considered as criteria for virulence. Among the adherence factors, F1 was found to be the most prevalent. AIDA-I and pAA were also found with similar ratio as compared with other virulence factors. In addition, virulence patterns carrying these alternate adhesive genes with enterotoxins were detected with significant ratio. Therefore, it is desirable that alternate adhesins be considered as markers for diagnosis of ETEC.
Enterotoxigenic Escherichia coli (ETEC) are known to be the causative agents of colibacillosis in piglets with diarrhea during suckling and weaning periods. 15 Neonatal diarrhea has always been one of the major problems facing the pig industry. In neonatal and weaned piglets, E. coli induce acute and watery diarrhea that may be followed by terminal bacteremia, which remains an important cause of economic loss for swine producers. 7
To develop a disease, ETEC strains need 2 kinds of virulence factors, fimbrial adhesins and enterotoxins. 15 Through fimbriae, ETEC attach to microvilli of intestinal epithelial cells and colonize the mucosal surface of the small intestine. Then, they produce 1 or more enterotoxins and cause diarrhea by changing the water and electrolyte balance within the small intestine. 15,16 There are diverse types of fimbriae and enterotoxins associated with ETEC in different host species. The common fimbriae associated with ETEC in swine are F4 (K88), F5 (K99), F6 (987P), F18, and F41. 7,15 The common enterotoxins are heat-stable (STa, STb) and heat-labile (LT). 15,16 Stx2e of shiga toxigenic E. coli (STEC) 14 and enteroaggregative E. coli heat-stable enterotoxin 1 (EAST1) of enteroaggregative E. coli (EAEC) are also closely related to ETEC. 22
Recently, while ETEC strains lacking recognized fimbriae are becoming more common, most ETEC strains have no known adherence factor. This may be related to the widespread use of vaccines targeting common virulence factors. 21 Thus, other adhesins from ETEC strains of swine have been intensively studied and include F1 (type 1 fimbriae), adhesin involved in diffuse adherence (AIDA-I), and porcine attaching and effacing-associated factor (pAA), F42, 23 F165, 8 and CS1541 antigen. 5 Among these, the F1 pilus is known to be expressed in the majority of E. coli strains from human and animal origin. 13 In human isolates, the F1 pilus has been proven to be a crucial adhesin of urinary tract infections 6 and inflammatory bowel disease, known as Crohn's disease. 3 It is also believed to affect the initial phase of the infectious process in poultry by allowing bacteria to attach to the tracheal epithelia. 19 However, its role in colibacillosis of piglets has been controversial. 1,4,9,21 It has been reported that the F1 pilus facilitates SIgA-mediated biofilm formation, and this result suggests that the F1 pilus may be a potential agent to colonize the gut. Therefore, it can be inferred that F1 fimbriae expressed in bacteria may play an important role in colibacillosis of pigs. 2 In addition, AIDA-I of diffuse-adhering E. coli (DAEC) 17 and pAA of EAEC have been considered as adherent molecules of porcine ETEC. 12
Several diagnostic methods, including the polymerase chain reaction (PCR), have been developed for detecting ETEC. However, multiplex (m)PCR assays targeting the genes encoding various virulence factors have not been well developed, and to the authors' knowledge, there is no report establishing mPCR to detect F1 pilus in ETEC strains from pigs. In the present study, 3 sets of mPCR assays were developed for detecting virulence factors of ETEC, which included the detection of the F1 pilus. The prevalence of virulence factors were then investigated within field isolates from piglets.
Between 2002 and 2007, 188 E. coli strains were isolated from piglets of suckling and weaning periods with diarrhea on different farms in South Korea. After incubation at 37°C for 18 hr on MacConkey's and eosin methylene blue (EMB) agar plates, single colonies were selected and identified as E. coli by IMViC (indole, methyl red, Voges–Proskauer, and citrate) test. Final confirmation was performed using the Vitek system. a These field isolates were grown overnight in trypticase soy broth (TSB) and, after addition of glycerol to 12.5%v/v, stored at −80°C until further use. The following E. coli strains were used as controls: S-99(LT), S-125 (STb and LT), S-126 (STa, STb, and LT), S-127 (STa, STb, and LT), E-94 (F1 and F5), and E-103 (F1 and F41). b , c For PCR amplification, E. coli isolates and reference strains were grown on Luria–Bertani (LB) agar plates, and a single colony from each bacterium was suspended in 50 μl of distilled water and boiled for 10 min. After chilling on ice for 5 min, the suspensions were centrifuged at 11,830 X g for 2 min, and the supernatants were used as DNA templates for PCR assays.
The polymerase chain reaction primers used in the present study.
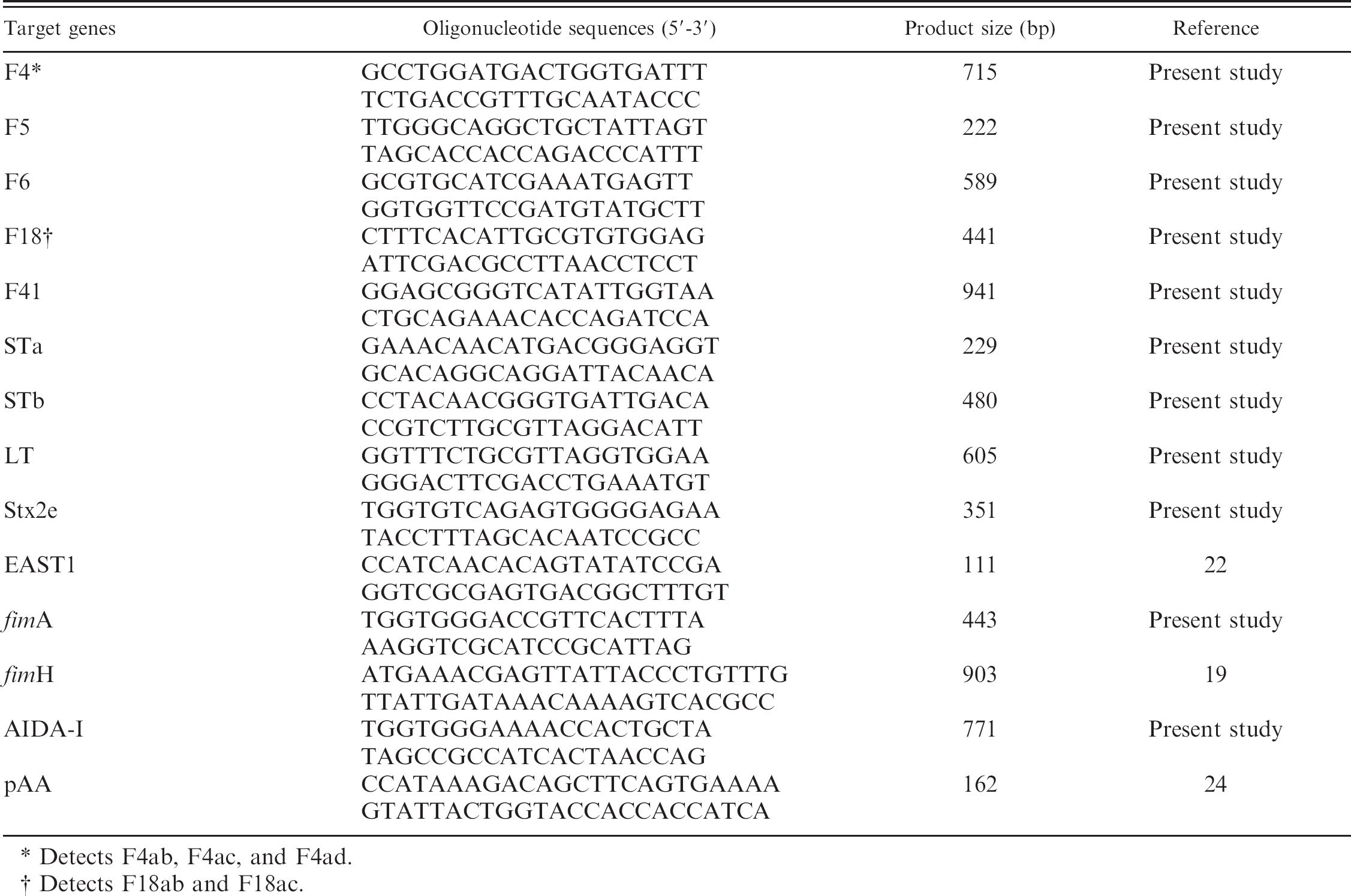

Detects F4ab, F4ac, and F4ad.
Detects F18ab and F18ac.
Fourteen sets of oligonucleotide primers d were used to detect the genes encoding for the following 13 different virulence factors of ETEC: the genes of fimbriae (F4, F5, F6, F18, and F41), the genes of enterotoxins (STa, STb, LT, Stx2e, and EAST1), and the genes of alternate adhesins (F1, AIDA-I, and pAA). Primers for EAST1, fimH of the F1, and pAA were selected according to previous reports. 19,22,24 Other primers were designed based on sequence data from GenBank using Primer 3 software. e The base sequences and predicted sizes of amplicons used in the current study are shown in Table 1.
Using these primers, 3 sets of mPCR assays were developed for diagnosis of ETEC. The reactions targeted the genes encoding the fimbriae, enterotoxins, and adhesins. The mPCR assays were carried out within 50 μl of reaction mixture containing 10X PCR buffer, f 1 mM of dNTPs, f 2.5 U of Taq polymerase, f 3 μl of the DNA template, and the PCR primers. The concentrations of primers were 0.4 pmole except EAST1 and pAA, which were 0.8 pmole.
Polymerase chain reaction of each set was carried out using the following steps: after initial denaturation at 95°C for 5 min, 30 cycles of amplification were performed (denaturation at 95°C for 30 sec, annealing at 61°C, 57°C, and 56°C for fimbriae, enterotoxins, and alternate adhesins, respectively, for 30 sec, extension at 72°C for 1 min), and final extension was at 72°C for 7 min. This condition was used for each set of PCR analysis. The PCR products were analyzed by electrophoresis on 2.0% agarose gel, and the amplicons were visualized under ultraviolet light after ethidium bromide staining.
The Mantel–Haenszel χ2 test was used to examine differences in prevalence of virulence genes of ETEC from piglets with diarrhea. The statistical analysis was performed using the Statistical Package for the Social Sciences (SPSS) program. g
Three sets of mPCR assays for detecting virulence factors of ETEC were established. To evaluate the specificity, each mPCR was optimized using the DNA templates from the reference E. coli strains. All of the PCR product amplicons were with expected sizes, and nonspecific bands were not observed (Fig. 1). There were no amplicons in PCR reactions with other members of the Enterobacteriaceae family and the E. coli DNA template, neither of which have virulence factors targeted in the PCR assays.
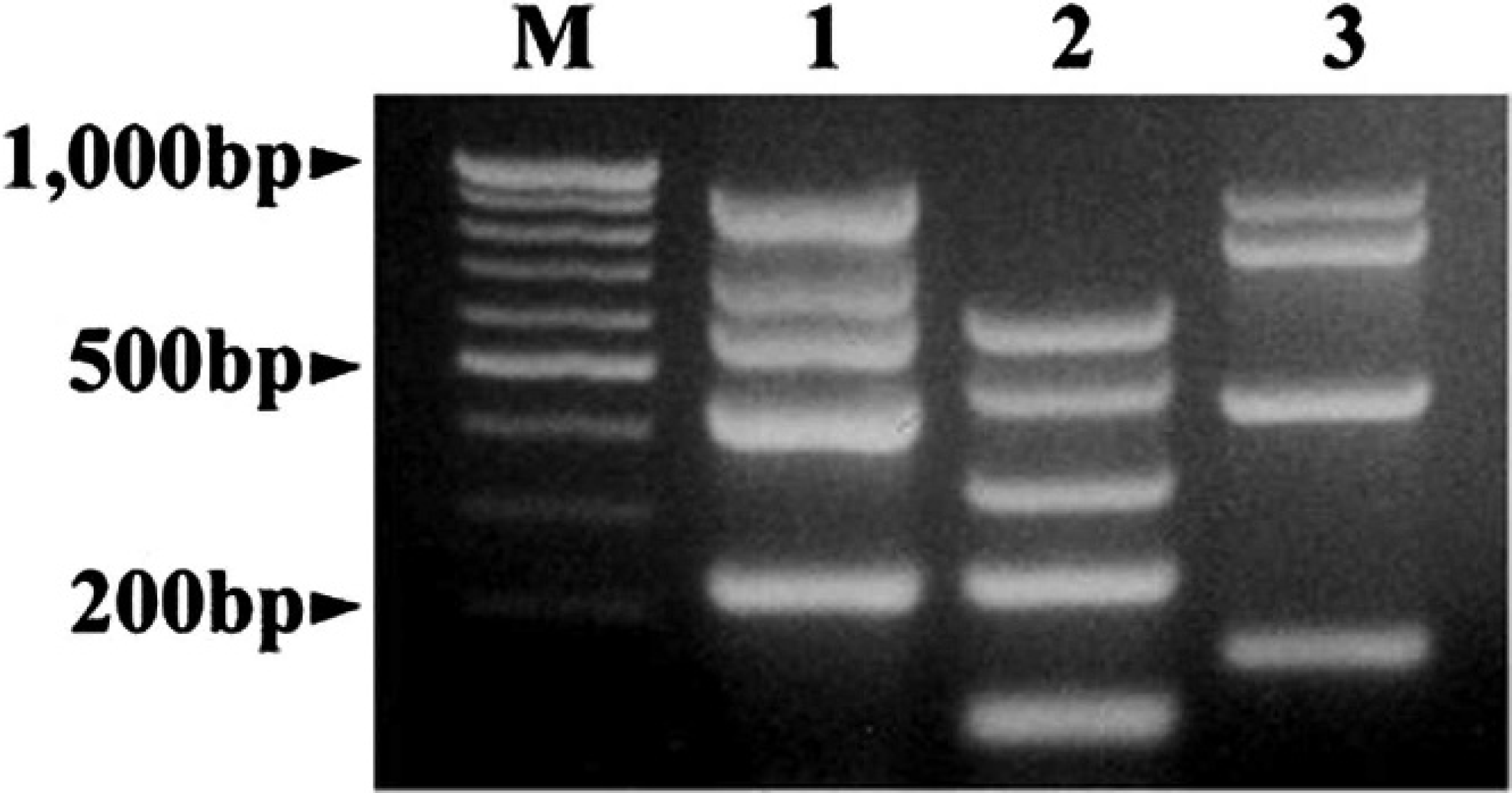
Three sets of multiplex polymerase chain reaction (mPCR) assays for detection of virulence genes for enterotoxigenic Escherichia coli. Lane M: molecular weight marker (100-bp ladder); lane 1: PCR targeting the genes of fimbriae (F4, F5, F6, F18, and F41); lane 2: PCR targeting the genes of enterotoxins (STa, STb, LT, Stx2e, and EAST1); lane 3: PCR targeting the genes of alternate adhesive factors (fimA and fimH of F1, AIDA-I, and pAA).
Out of 188 E. coli strains, 48 isolates (25.53%) were found to be pathogenic ETEC carrying both fimbriae and enterotoxins. When all adhesins were considered as criteria for virulence, 91 isolates (48.40%) were found to be pathogenic ETEC, whereas only 17 isolates (9.04%) did not carry any virulence gene. Percentages of E. coli strains carrying genes of the adherence factors and enterotoxins were as follows: F4 (7.45%), F5 (29.79%), F6 (6.38%), F18 (15.43%), F41 (3.72%), STa (10.11%), STb (20.74%), LT (9.57%), Stx2e (2.13%), EAST1 (42.02%), F1 (67.55%), AIDA-I (2.66%), and pAA (7.45%). Prevalence of E. coli strains carrying genes of each virulence factor in suckling and weaned piglets is shown in Table 2.
A presence of diverse patterns of virulence profiles was observed (Table 3). In the PCR to detect the genes of fimbriae, bacteria harboring the genes encoding more than one of the fimbriae were found in 74 isolates (39.36% of total isolates). Among fimbrial genes, F5 was the most prevalent (56/74; 75.68%; P < 0.05). Of these 56 isolates, 30 isolates had the gene encoding F5 only, and 15 isolates had the genes encoding both F5 and F18. The second most prevalent fimbrial gene was F18 (29/74; 39.19%; P < 0.05), which was primarily associated with F5 in 21 isolates, whereas 6 of these isolates had F18 alone. Using PCR assay to detect the genes of enterotoxins, 107 isolates (56.91% of total isolates) were confirmed to have the genes encoding for at least one of the enterotoxins. Among enterotoxin genes, EAST1 was the most prevalent gene (79/107; 73.83%; P < 0.05) and STb was second (39/107; 36.45%; P < 0.05). The pattern carrying only EAST1 was prevalent (50/107; 46.73%), and the pattern carrying both EAST1 and STb was second (28/107, 26.17%). In the PCR assay to detect genes of alternate adhesins, the genes encoding at least one of the adhesins were detected in 140 isolates (74.47%). Among them, both alternate adhesin and fimbriae were found in 59 isolates (31.18% of total isolates), and presence of alternate adhesins without fimbriae were found in 81 isolates (43.08% of total isolates). Of the isolates carrying adhesins, F1 was the most prevalent (127/140; 67.55%; P < 0.05).
In the present study, 3 sets of mPCR assays were developed to detect genes encoding different virulence factors of ETEC. Each set was optimized to detect fimbriae, enterotoxins, and alternate adhesins, respectively. All primers used in the current study, except for EAST1, fimH, and pAA, were newly designed, and the primer for F4 (used to detect F4ab, F4ac, and F4ad) and F18 (used to detect F18ab and F18ac) were designed to detect these genes at one time. Thus, convenient and efficient diagnosis of ETEC might be possible by selecting each sorted PCR assay on a case-by-case basis.
Using the mPCR assays, the authors detected clinical isolates of ETEC from suckling and weaned piglets from different regions of South Korea. There was no significant difference in the ratio of ETEC between suckling and weaning periods. However, the distribution of fimbriae and enterotoxins was indicative of having some differences. The ratio of fimbriae was higher during the weaning period (45.57%) than during the suckling period (34.86%). In contrast, enterotoxins were higher during the suckling period (60.55%) than during the weaning period (51.90%). Escherichia coli strains with EAST1, which was isolated during suckling periods, was more prevalent than other enterotoxins, and the prevalence in suckling period was higher compared with weaning periods. The prevalence of E. coli carrying F4, F5, F6, F18, and F41 was slightly higher in weaned piglets than suckling piglets. Several studies have demonstrated that F4 pili are common, representing 15.9–70.2% of the E. coli strains causing diarrhea in piglets. 15,20 However, F5 pili (29.79%) and F18 pili (15.43%) were more common than F4 pili (7.45%) in the present study.
Prevalence of Escherichia coli strains carrying genes of virulence factors of enterotoxigenic E. coli in suckling and weaned piglets.
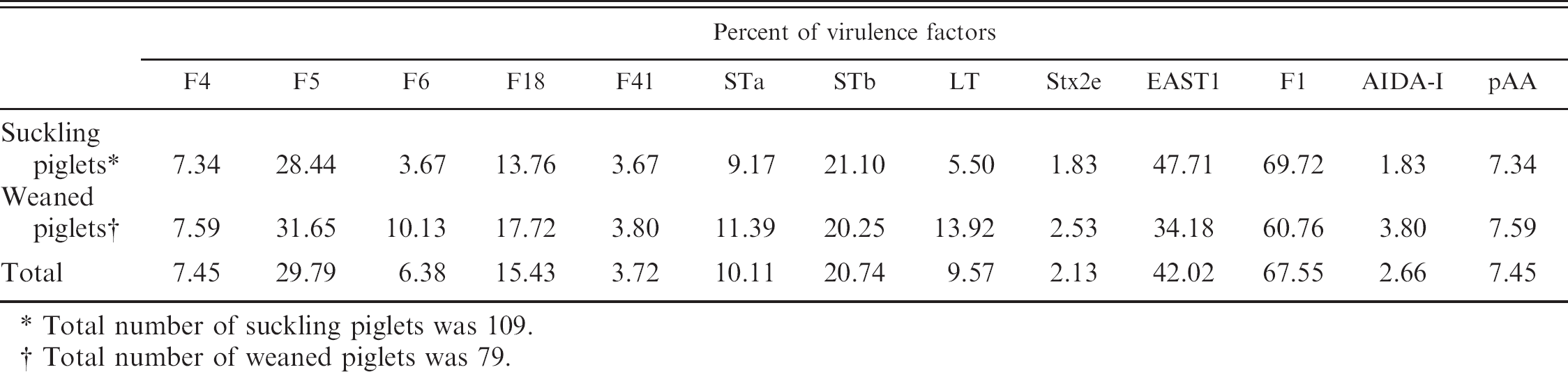
Total number of suckling piglets was 109.
Total number of weaned piglets was 79.
Distribution of genes for adherence factors and enterotoxins among 188 Escherichia coli strains isolated from piglet with diarrhea during suckling and weaning periods.
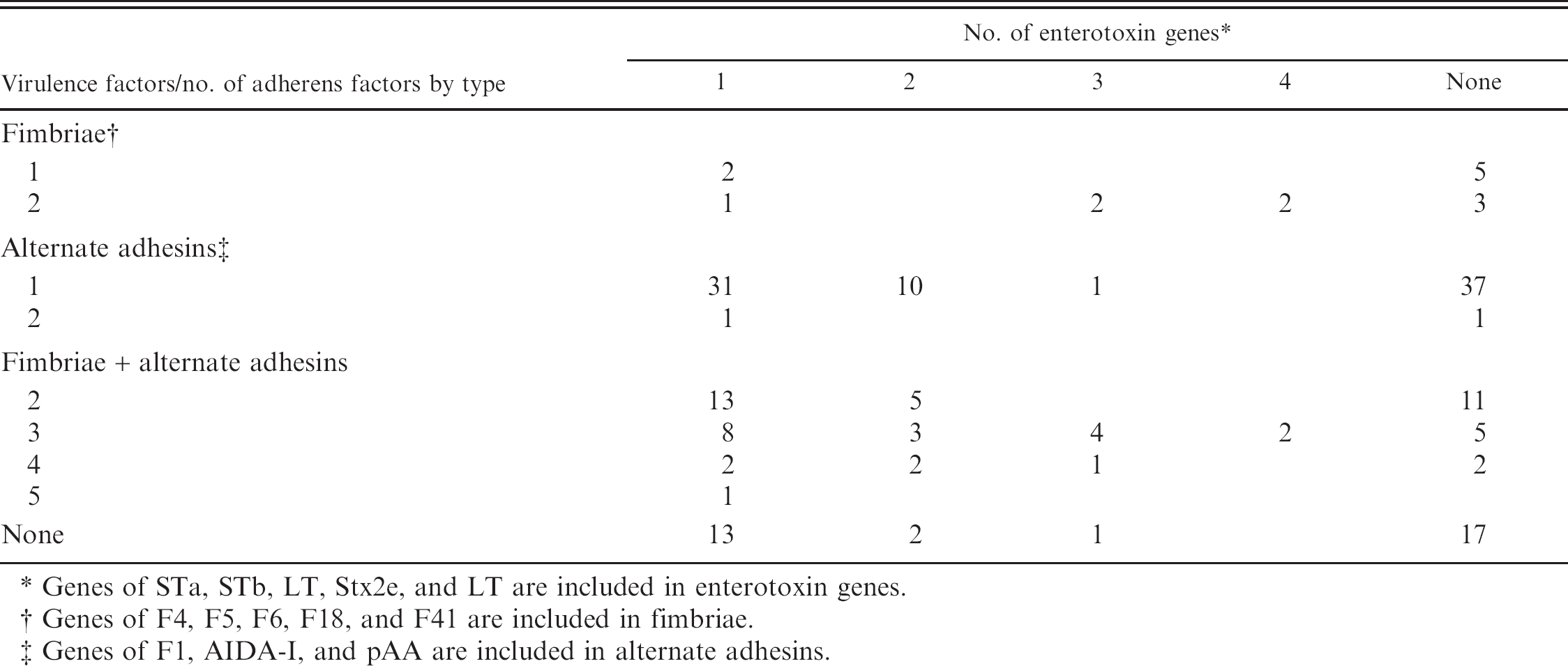
Genes of STa, STb, LT, Stx2e, and LT are included in enterotoxin genes.
Genes of F4, F5, F6, F18, and F41 are included in fimbriae.
Genes of F1, AIDA-I, and pAA are included in alternate adhesins.
The isolates of E. coli that harbored genes for fimbrial adhesins or enterotoxins are regarded as pathogenic ETEC. 15 The prevalence of pathogenic ETEC among E. coli strains from field isolates was 25.53% in the present study. However, when alternate adhesins were considered as virulence factors, the ratio was increased to 48.40%. The prevalence of the alternate adhesins appeared to be high in this examination. Of the 84 isolates (44.68% of a total of 188 isolates) that had alternate adhesins with enterotoxins, 43 isolates (22.87% of a total 188 isolates) were not found to carry any fimbria. Although the role of alternate adhesins in the pathogenesis of ETEC has been unclear, the distribution of the genes in ETEC has been examined in many studies. 21,24 Prevalence of F1 pili was high (66.33%). A previous study showed that 81% of E. coli isolates from neonatal piglets with diarrhea expressed the F1 pili. 4 Thus, even though heterogeneity between other serotypes is considered, the F1 pili on some E. coli strains can be regarded as pathogenic or at least having a potential role in the adherence to pig gastrointestinal tract. The F1 pilus is composed of a repeating major subunit, FimA and several minor subunits, FimF, FimG, and FimH. 11 The FimH is an adhesive protein and localizes at the lateral ends of the fimbriae. 10,11 Thus, detection of both FimA and FimH is required to detect F1 pili accurately and the mPCR assay was designed to target these 2 genes in the present study. Additionally, AIDA-I (2.66%) and pAA (7.45%) were also detected and were considered as contributing to ETEC diarrhea.
There were isolates recovered that do not carry any virulence factors of ETEC. These isolates may possess ETEC carrying new or other virulence factors that were not examined in this study, or other pathogens known as causative agents of diarrhea in piglets, such as Porcine epidemic diarrhea virus (PEDV), Transmissible gastroenteritis virus (TGEV), and type-B rotavirus. 18
To detect the virulence genes of ETEC, 3 sets of mPCR assays were developed resulting in an efficient and practical method for the detection of ETEC isolated from piglets with diarrhea. Of the 188 isolates of E. coli, 48 isolates (25.53%) containing the genes for fimbriae and enterotoxins were classified as pathogenic ETEC. However, including alternate adhesins as criteria for virulence increased the percent virulence isolates to 48.40%. Thus, it is suggested that alternate adhesins be considered as markers for diagnosis of ETEC.
Acknowledgements. This study was supported through Biogreen 21 (20070401034009) RDA, BK21 Program for Veterinary Science, KRF (2006–005–J502901) and the Research Institute for Veterinary Science, Seoul National University.
Footnotes
a.
bioMérieux Inc., Durham, NC.
b.
National Veterinary Research and Quarantine Service, Anyang, South Korea.
c.
Escherichia coli strains carrying F4, F5, F6, F18, F41, Stx2e, EAST1, AIDA-I, and pAA supplied by Professor C. Chae, Seoul National University, Seoul, South Korea.
d.
Bioneer, Daejon, South Korea.
e.
National Human Genome Research Institute Inc., Bethesda, MD.
f.
iNtRON Biotechnology Inc., Seongnam, South Korea.
g.
SPSS Inc., Chicago, IL.
