Abstract
Thirty-seven fluoroquinolone-resistant Escherichia coli strains from ruminants (according to Clinical and Laboratory Standards Institute guidelines) were screened by molecular methods for mutations in the quinolone resistance-determining region (QRDR) of the gyrA and parC genes and for the presence of the qnrA gene. One of the strains studied was an enterohemorrhagic E. coli (EHEC) strain potentially pathogenic for humans. Three E. coli strains resistant to enrofloxacin (minimal inhibitory concentration [MIC] = 2 μg/ml) but not to ciprofloxacin (MIC = 1 μg/ml) presented single mutations in the gyrA and parC genes, while 34 strains resistant to both fluoroquinolones presented double and single mutations in gyrA and parC, respectively (31 strains), or double mutations in gyrA and parC (3 strains). The EHEC strain presented a double amino acid substitution in the GyrA protein (Ser-83→Leu and Asp-87→Gly) and a double amino acid substitution in the ParC protein (Gly-78→Cys and Ser-80→Arg), one of which has not been previously described. The present study shows that most of the mutations in the QRDR of the gyrA and parC genes of fluoroquinolone-resistant E. coli strains from ruminants are the same as those seen in E. coli strains from other animal species and humans and that there are no differences in mutation patterns in the QRDR of E. coli strains from healthy ruminants and those with diarrhea. No strains carried qnrA, which indicates that this gene does not play an important role in the selection of fluoroquinolone-resistant E. coli strains from ruminants.
Introduction
Fluoroquinolones are broad-spectrum antimicrobials used in medicine and veterinary practice in Europe for treatment of infectious diseases caused by enteric bacteria such as Escherichia coli. Before the early 1990s, fluoroquinolone resistance was rarely found in human and animal E. coli; however, since then the frequency has significantly increased worldwide. 3,4,6,11–13,16,21,22
The most frequent mechanism of quinolone resistance in E. coli includes alterations in genes that encode subunits of the quinolone targets DNA gyrAse (in the gyrA and gyrB genes) and topoisomerase IV (in the parC and parE genes). These alterations involve mainly mutations located in the quinolone resistance-determining region (QRDR) of the gyrA gene and its homologous region of the parC gene. In contrast, mutations in the gyrB and parE genes are of minor importance and are rare contributors to quinolone resistance. 3–6,16,18,21,22 Plasmids carrying qnr genes have been found to transmit quinolone resistance. These genes encode pentapeptide repeat proteins that block the action of quinolones on bacterial DNA gyrAse and topoisomerase IV, resulting in low-level quinolone resistance. 14 This reduced susceptibility is most likely important for facilitating the selection of mutants with higher-level resistance. 14 Although the qnrA and qnrB genes seem to be uncommon among human clinical isolates of E. coli from the United States and Europe, 10,14 qnrA is prevalent in fluoroquinolone-resistant E. coli from Asia. 17,19 Other mechanisms of resistance, such as decreased expression of outer membrane porins or overexpression of multidrug efflux pumps, can decrease susceptibility to quinolones. 3,6,16
Amino acid changes in the quinolone resistance-determining region (QRDR) of GyrA and ParC proteins in 37 fluoroquinolone-resistant Escherichia coli strains from ruminants and the corresponding enrofloxacin and ciprofloxacin minimal inhibitory concentrations (MICs).
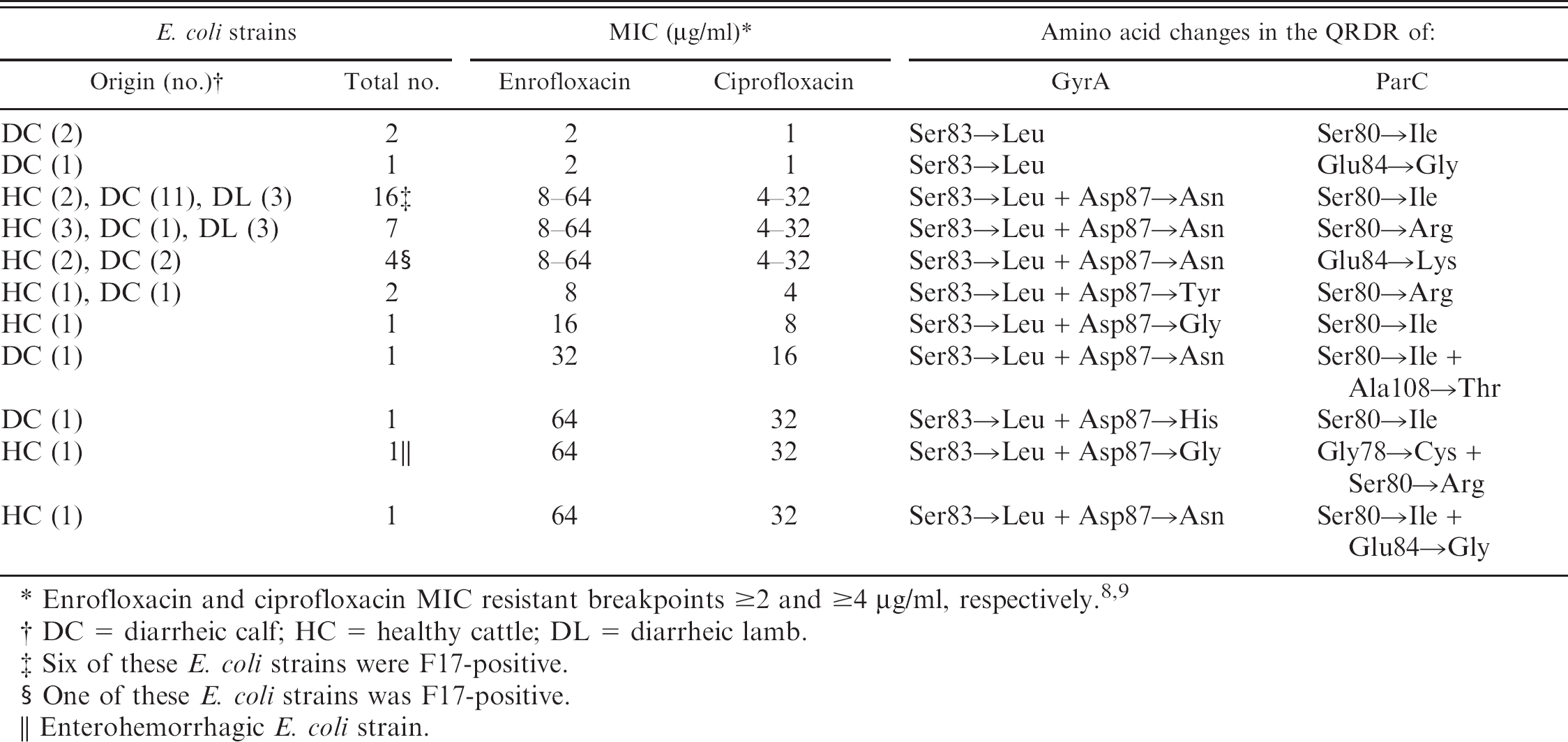
Enrofloxacin and ciprofloxacin MIC resistant breakpoints ≥2 and ≥4 μg/ml, respectively. 8,9
DC = diarrheic calf; HC = healthy cattle; DL = diarrheic lamb.
Six of these E. coli strains were F17-positive.
One of these E. coli strains was F17-positive.
Enterohemorrhagic E. coli strain.
It is important to characterize the mechanisms of resistance to fluoroquinolones. 20 Most of the studies on mechanisms of quinolone resistance in E. coli have been performed in human clinical strains, and, to the authors' knowledge, the study of these mechanisms in fluoroquinolone-resistant E. coli strains from ruminants has not been performed. In the current study, 37 fluoroquinolone-resistant E. coli strains from cattle and sheep were screened for mutations in the QRDR of the gyrA and parC genes and for the prevalence of the qnrA gene.
A total of 37 E. coli strains isolated from diarrheic calves and lambs (20 and 6 animals, respectively) and from healthy cattle (11 animals) were used in the present study. The minimal inhibitory concentrations (MICs) of quinolones were determined in accordance with Clinical and Laboratory Standards Institute (CLSI, previously the National Committee for Clinical Laboratory Standards) guidelines. 8,9 All strains were isolated between 1993 and 1998 from farms located in Spain and were resistant to nalidixic acid (MIC > 512 μg/ml; MIC resistant breakpoint ≥ 32 μg/ml) and enrofloxacin (MIC = 4–64 μg/ml; MIC resistant breakpoint ≥ 2 μg/ml), according to CLSI guidelines. 8 In addition, all strains, with the exception of 3 of them, were resistant to ciprofloxacin (MIC = 1–32 μg/ml; MIC resistant breakpoint ≥ 4 μg/ml), also according to CLSI guidelines. 9 One of these strains was an enterohemorrhagic E. coli (EHEC) strain potentially pathogenic for humans. 2 In addition to the EHEC strain, 7 strains produced the F17 fimbria and are, thus, potentially pathogenic for ruminants. 7 All these strains, with the exception of 1 strain, have been described previously. 11–13
Detection of mutations in the QRDR of the gyrA gene, as well as in the analogous region of the parC gene, was performed in the 37 fluoroquinolone-resistant E. coli strains by polymerase chain reaction (PCR). 3 Amplified fragments were purified, a and both strands were automatically sequenced b using the same set of primers as for the PCR. The qnrA gene was detected by PCR, as previously described. 15 Plasmid pMG252 was used as positive control.
Table 1 shows the enrofloxacin and ciprofloxacin MICs for the 37 fluoroquinolone-resistant E. coli strains and the deduced amino acid changes in the QRDR of the GyrA and ParC proteins. None of the E. coli strains tested carried the qnrA gene. Three E. coli strains resistant to enrofloxacin (MIC = 2 μg/ml) but not to ciprofloxacin (MIC = 1 μg/ml), according to CLSI guidelines, 8,9 showed single mutations in the gyrA and parC genes, while 34 strains resistant to both fluoroquinolones showed double and single mutations in gyrA and parC, respectively (31 strains), or double mutations in gyrA and parC (3 strains). Four mutations (2 in gyrA and 2 in parC) were associated with high levels of resistance to enrofloxacin and ciprofloxacin (32–64 and 16–32 μg/ml, respectively). However, some E. coli strains with 3 mutations showed the same range of enrofloxacin and ciprofloxacin MICs as those showed by strains with 4 mutations. It is possible that in addition to mutations in gyrA and parC, some E. coli strains display other mechanisms of resistance to fluoroquinolones, especially those strains with a high level of resistance. There were no differences in mutation patterns in the QRDR of E. coli strains from healthy ruminants and from those with diarrhea.
All E. coli strains resistant to fluoroquinolones exhibited the Ser83→Leu substitution in the GyrA protein. Nucleotide substitutions at codon 83 are the most common gyrA mutations found in clinical, veterinary, and laboratory strains of E. coli, and this serine residue is most commonly substituted with a leucine. 3,4,6,16,21,22 The 34 strains resistant to ciprofloxacin exhibited an additional substitution in the GyrA protein of Asp-87 for Asn (29 strains), Gly (2 strains), Tyr (2 strains), or His (1 strain). Not all mutations in gyrA confer the same level of quinolone resistance. Thus, mutations at codons Ser83 and Asp87 confer higher levels of quinolone resistance than mutations at other codons within the QRDR. 6
As in other E. coli strains previously described, 3,4,6,16,21,22 mutations in parC were always found together with mutations in gyrA. This is due to topoisomerase IV being a secondary target for quinolones in E. coli. 6 However, mutations in parC play an important role in the formation of highly resistant strains. 6 A single amino acid change in the QRDR of the ParC protein was found in 34 strains. In addition, double amino acid changes in the ParC protein were detected in 3 strains. The amino acid substitutions were as follows: Gly-78→Cys (1 strain), Ser-80→Arg (10 strains), Ser-80→Ile (22 strains), Glu-84→Lys (4 strains), Glu-84→Gly (2 strains), and Ala−108→Thr (1 strain). Mutations in parC at codon 80 have been the most frequently identified, with Ser-80→Ile being the most common substitution. 3,4,6,16,22 To the authors' knowledge, this is the first report of the Gly-78→Cys and Ala−108→Thr substitutions in the ParC protein of E. coli strains. With respect to amino acid substitutions in the ParC protein, the ability to form hydrogen bonds is reduced when Ser-80 is substituted for Ile or Arg, whereas the change of Glu for Lys or Gly at position 84 involves the loss of a negative charge. 16
One of the strains resistant to fluoroquinolones was an EHEC strain (positive for verotoxin and eae gene) isolated from a healthy cow. This strain was the first fluoroquinolone-resistant EHEC described (enrofloxacin and ciprofloxacin MICs of 64 and 32 μg/ml, respectively) 11 and presented a double amino acid substitution in the GyrA protein (Ser-83→Leu and Asp-87→Gly) and a double amino acid substitution in the ParC protein (Gly-78→Cys and Ser-80→Arg), one of which, as described above, has been reported for the first time in this work. To the authors' knowledge, this is the first fluoroquinolone-resistant EHEC strain in which the mutations of gyrA and parC have been studied. The resistance to fluoroquinolones of the EHEC strain is especially relevant since that strain belongs to one of the serogroups (O103) most frequently isolated from children with severe diarrhea and hemolytic-uremic syndrome. 1 Fluoroquinolones, although not recommended for treating gastroenteritis or diarrhea, are indicated for the treatment of invasive illness. 6
The 7 F17-positive E. coli strains were isolated from diarrheic calves, 12 and their enrofloxacin and ciprofloxacin MICs ranged from 8–32 and 4–16 μg/ml, respectively. All these strains exhibited the Ser-83→Leu and Asp-87→Asn substitutions in the GyrA protein and an additional substitution of Ser-80→Ile (6 strains) or Glu-84→Lys (1 strain) in the ParC protein.
It has been suggested that the qnrA gene may contribute to the increase in quinolone resistance in human E. coli. 19 However, the absence of that gene in the E. coli strains tested in the present study indicates that qnrA does not play an important role in the selection of fluoroquinolone-resistant E. coli strains from ruminants.
In conclusion, the present study shows that most of the mutations in the QRDR of the gyrA and parC genes of fluoroquinolone-resistant E. coli strains from ruminants are the same as those seen in E. coli strains from other animal species and humans and that there are no differences in mutation patterns in the QRDR of E. coli strains from healthy ruminants and from those with diarrhea. The results of this study also indicate that fluoroquinolone resistance mediated by qnrA in E. coli from ruminants is rare.
Acknowledgements. This study was supported by grants from the Dirección General de Investigación (AGL2004–08139) and the Universidad Complutense-Comunidad de Madrid (920338). The authors thank J. M. Rodríguez-Martínez (Hospital Virgen Macarena, Seville, Spain) for generously providing the positive control of qnrA.
Footnotes
a.
SpinPrepTM PCR Clean-up kit, Novagen, Merck KGaA, Darmstadt, Germany.
b.
Secugen SL, Madrid, Spain.
