Abstract
We investigated in vitro activity of a novel veterinary fluoroquinolone, pradofloxacin, against methicillin-resistant Staphylococcus pseudintermedius (MRSP) isolates and compared with other fluoroquinolones. A total of 38 MRSP isolates were subjected to agar disk diffusion tests for sensitivity to pradofloxacin, orbifloxacin, marbofloxacin, enrofloxacin, and ciprofloxacin. The minimal inhibitory concentration (MIC) values of pradofloxacin, ciprofloxacin, and enrofloxacin were determined. Mutations in the genes encoding DNA gyrase subunit A (GyrA) and topoisomerase IV (GrlA) proteins associated with fluoroquinolone resistance were studied by an analysis of partial sequences of the genes encoding these proteins. Two MRSP isolates were susceptible in disk diffusion and microdilution test to all fluoroquinolones tested, including pradofloxacin. Based on the results of the disk diffusion testing, 33 of 38 isolates showed resistance to pradofloxacin and 3 were intermediate, whereas, by pradofloxacin MIC testing, 35 isolates were classified as resistant and 1 as intermediate. Single alterations in GyrA and GrlA proteins were observed in the 35 resistant isolates and the 1 intermediate isolate (MIC results). These same 36 isolates were also resistant to the other tested fluoroquinolones. The results of the current study showed that MRSP isolates are usually resistant to all fluoroquinolones, including pradofloxacin. Therefore, in routine susceptibility testing to pradofloxacin by disk diffusion, the results should be carefully interpreted for MRSP isolates, especially those resistant to other fluoroquinolones and, in questionable cases, the pradofloxacin MIC should be determined to confirm the susceptibility testing results.
Introduction
Fluoroquinolones are synthetic antimicrobials that have been used in human and veterinary medicine for many years. Pradofloxacin is a novel third-generation veterinary fluoroquinolone approved in Europe and Canada for treatment of bacterial infections in dogs and cats. In the United States, pradofloxacin is approved for the treatment of bacterial infections in cats.4,18,27 In comparison with fluoroquinolones of earlier generations, pradofloxacin shows an extended spectrum of activity against gram-positive and gram-negative bacteria, including anaerobic pathogens.11,18,26,28 The first fluoroquinolone developed for veterinary application was enrofloxacin. 21 Other veterinary antimicrobials of this group are orbifloxacin and marbofloxacin. Although ciprofloxacin is mainly used in human medicine and not licensed for use in animals, it is occasionally used off-label for veterinary purposes.3,21,22
The activity of pradofloxacin, like other fluoroquinolones, is a result of the inhibition of 2 bacterial enzymes involved in the DNA replication process, the DNA gyrase (topoisomerase II) and topoisomerase IV.15,18,21 Fluoroquinolones of earlier generations also inhibit these 2 enzymes, but DNA gyrase is the primary target for gram-negative bacteria, and topoisomerase IV is the primary target for gram-positive bacteria. However, the older fluoroquinolones have lower affinity for topoisomerase IV than DNA gyrase. In contrast, pradofloxacin inhibits both enzymes equally. 12 Mutations in the DNA gyrase (gyrA) or topoisomerase IV (grlA) genes cause changes in encoded proteins leading to resistance to fluoroquinolones.12,21 The mutations responsible for resistance more frequently occur in specific regions of each enzyme subunit called the quinolone-resistance determining regions (QRDRs). Alterations in QRDRs involved in fluoroquinolone resistance have been described in several bacterial species including gram-positive species such as Staphylococcus aureus, Staphylococcus pseudintermedius, and Staphylococcus epidermidis.1,16,26
Dogs are considered natural hosts of S. pseudintermedius, which is one of the common opportunistic pathogens of these animals, often associated with skin, ear, or wound infections. During the past decade, methicillin-resistant S. pseudintermedius (MRSP) has emerged as an important pathogen in veterinary medicine.5,24 As MRSP isolates are often also resistant to other antimicrobials, choosing an effective antimicrobial agent for treatment of infections caused by these pathogens can be difficult.10,23 Most MRSP strains are resistant to fluoroquinolones of earlier generations.8,10,17,24
It has been reported that pradofloxacin is characterized by low minimal inhibitory concentration (MIC) and mutant prevention concentration (MPC) values against target bacteria pathogenic for dogs and cats, including S. pseudintermedius.18,26,27 So far, the available data on susceptibility of canine S. pseudintermedius to pradofloxacin do not differentiate between methicillin-susceptible S. pseudintermedius (MSSP) and MRSP. The purpose of our work was to assess the in vitro activity of pradofloxacin, and to compare that activity with that of orbifloxacin, marbofloxacin, enrofloxacin, and ciprofloxacin against canine MRSP strains. As well, the study aimed to establish the relationship between alterations in QRDRs and resistance of MRSP to pradofloxacin.
Materials and methods
The canine MRSP isolates (n = 38) were obtained from clinical specimens submitted to the Microbiological Diagnostic Laboratory, Faculty of Veterinary Medicine, Warsaw University of Life Sciences-SGGW, from January 2007 to December 2011. Ten isolates were obtained from the skin and its appendages, 9 from wounds or abscesses, 7 from urine samples, 6 from the respiratory tract, 4 from internal organs, and 2 from the external auditory canal. Some of these strains were characterized previously.5,17 Identification of S. pseudintermedius strains was based on standard bacteriologic methods 19 and confirmed by a restriction fragment length polymorphism–PCR (RFLP-PCR) method 2 and the nuc multiplex PCR. 25 Methicillin resistance was determined by mecA gene amplification. 29 Three reference strains, Staphylococcus aureus ATCC 25923, a methicillin-susceptible Staphylococcus aureus ATCC 29213, a and methicillin-resistant Staphylococcus aureus Microbank 14.002, b were included in the study as controls for susceptibility testing and PCR.
The MIC of pradofloxacin c was determined by broth microdilution according to Clinical and Laboratory Standards Institute (CLSI) standards, 6 within the concentration range 0.008–16 µg/mL. The MICs of enrofloxacin and ciprofloxacin were determined using commercial strips d on Mueller–Hinton 2 agar e inoculated with a bacterial suspension (density of 0.5 McFarland) and incubated for 24 h at 35°C. The reference strain S. aureus ATCC 25923 a was used as a quality control for MIC determination with the commercial strips, according to the manufacturer’s instructions.
All 38 isolates were additionally subjected to antibiotic susceptibility testing by disk diffusion 6 for the following fluoroquinolones: orbifloxacin (10 µg), f marbofloxacin (5 µg), g enrofloxacin (5 µg), f ciprofloxacin (5 µg), f and pradofloxacin (5 µg). h The isolates were suspended in sterile saline, adjusted to the density of a 0.5 McFarland standard, spread plated on Mueller–Hinton agar, e and the disks applied. After the standard incubation, diameters of growth inhibition zones were measured. The scattergram for pradofloxacin was constructed by plotting zone diameters against MICs. 20
The isolates were classified as susceptible, intermediate, or resistant to the tested fluoroquinolones according to the criteria for veterinary pathogens approved by CLSI. 7 The guidelines based on human breakpoints recommended by the European Committee on Antimicrobial Susceptibility Testing (EUCAST, 2015 Breakpoint tables for interpretation of MICs and zone diameters, version 5.0, http://www.eucast.org) were used for ciprofloxacin.
All MRSP isolates were characterized by analysis of the QRDR sequences. The partial fragments of the gyrA and grlA genes were amplified by PCR, using primers and reaction conditions described previously.14,26 The obtained amplicons were sequenced using the forward PCR primers and a commercial sequencer. i The sequencing files were analyzed using a commercial software package, j and nucleotide sequences were translated into amino acid sequences with ExPASy Translate Tool (http://web.expasy.org/translate). Finally, amino acid sequences were aligned with the sequences of GyrA and GrlA proteins of the fluoroquinolone-susceptible reference strain, Staphylococcus pseudintermedius CCUG49543 (accessions CAK18884 and CAK18890), on the National Center for Biotechnology Information (NCBI) website (http://blast.ncbi.nlm.nih.gov) with Standard Protein BLAST. The multiple-sequence alignment was conducted with the CLUSTAL W program (http://www.clustal.org/) implemented in MEGA version 5.0.
Results
The MIC values of the tested fluoroquinolones for the canine MRSP isolates are presented in Table 1. The results of the disk diffusion tests and the MIC values of pradofloxacin are presented in Table 2. The MIC for the reference S. aureus ATCC 25923 strain determined with commercial strips b was 0.25 µg/mL for both enrofloxacin and ciprofloxacin, which is within the range specified by the manufacturer. Two MRSP isolates had MICs of enrofloxacin, ciprofloxacin, and pradofloxacin greatly below the breakpoints, and showed wide growth inhibition zones (34 mm in diameter for pradofloxacin disk), thus they were classified as susceptible to these antimicrobial agents. Based on disk diffusion testing results, the remaining 36 MRSP isolates were classified according to available interpretative criteria as resistant to ciprofloxacin, enrofloxacin, marbofloxacin, and orbifloxacin. The scattergram that resulted from plotting zone diameter against MICs for pradofloxacin revealed that 2 isolates were classified as susceptible and 33 as resistant by both methods (Table 2). However, 3 isolates were intermediate by disk diffusion test, 2 of which were classified as resistant according to microdilution test results. The remaining isolate exhibited intermediate resistance in both susceptibility testing methods.
The minimal inhibitory concentration (MIC) values for enrofloxacin, ciprofloxacin, and pradofloxacin against 38 canine methicillin-resistant Staphylococcus pseudintermedius isolates.
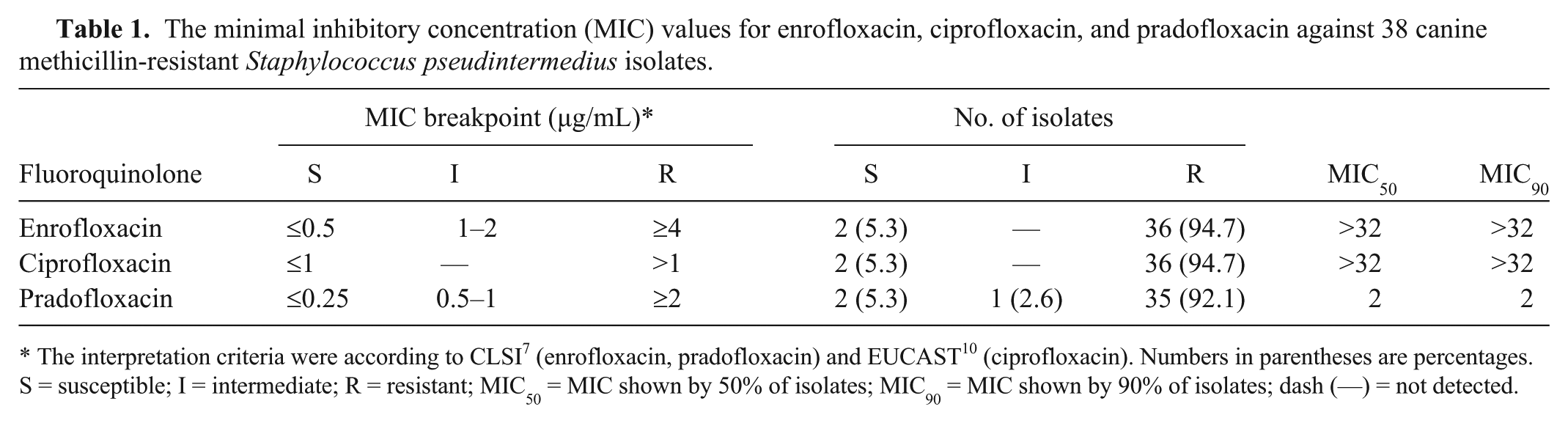
The interpretation criteria were according to CLSI 7 (enrofloxacin, pradofloxacin) and EUCAST 10 (ciprofloxacin). Numbers in parentheses are percentages. S = susceptible; I = intermediate; R = resistant; MIC50 = MIC shown by 50% of isolates; MIC90 = MIC shown by 90% of isolates; dash (—) = not detected.
Correlation between pradofloxacin minimal inhibitory concentrations (MICs) and zone diameters for 38 canine methicillin-resistant Staphylococcus pseudintermedius isolates.*
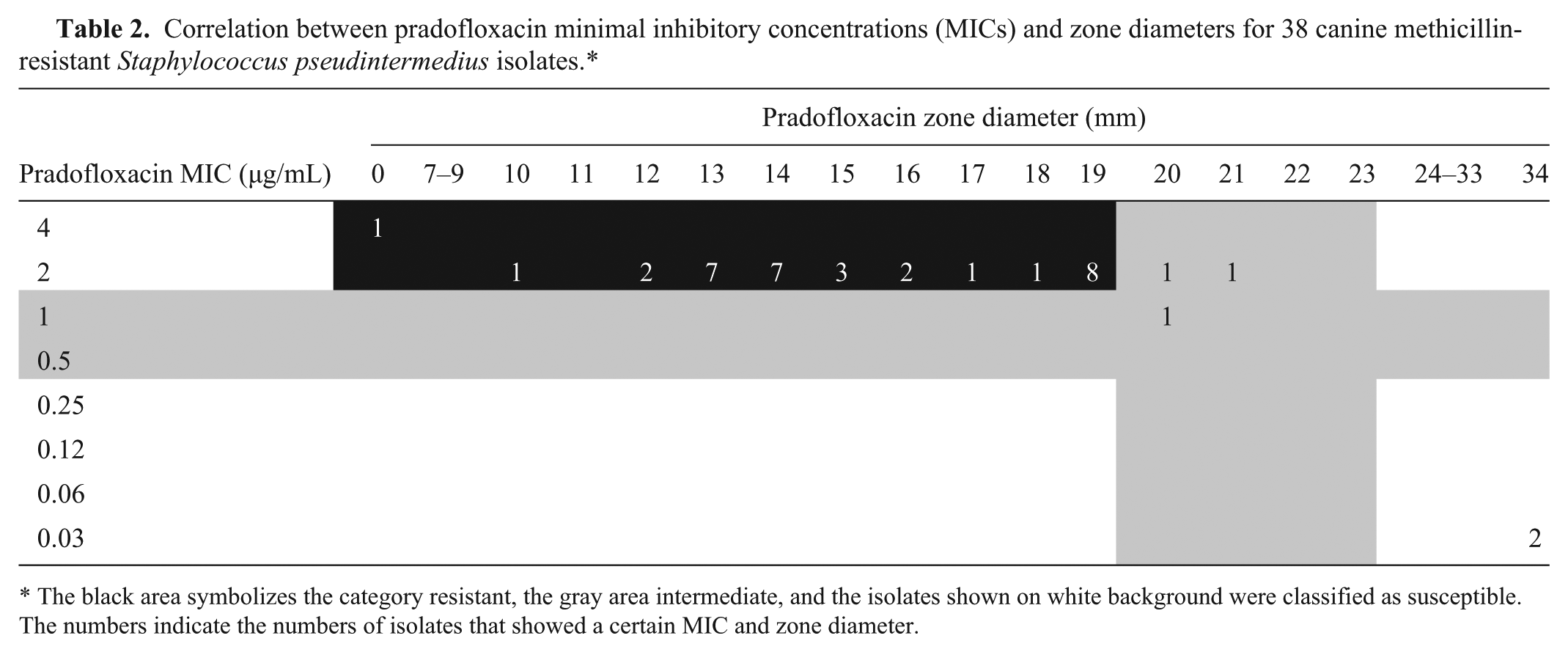

The black area symbolizes the category resistant, the gray area intermediate, and the isolates shown on white background were classified as susceptible. The numbers indicate the numbers of isolates that showed a certain MIC and zone diameter.
The alterations in the amino acid sequence of the GyrA and GrlA proteins of 38 MRSP isolates are presented in Table 3. A single alteration in the GyrA protein (serine to leucine at position 84) and a single alteration in the GrlA protein (serine to isoleucine at position 80) were observed to occur together in all MRSP isolates resistant to all fluoroquinolones according to MICs, and also in 1 intermediate isolate. A single mutation in GrlA, conversion of serine to isoleucine, was observed in 1 isolate susceptible to fluoroquinolones. The other isolate susceptible to all fluoroquinolones did not show any alterations in the amino acid sequence of either the GyrA or GrlA proteins.
The alterations in amino acid sequences of GyrA and GrlA proteins of 38 methicillin-resistant Staphylococcus pseudintermedius isolates.*
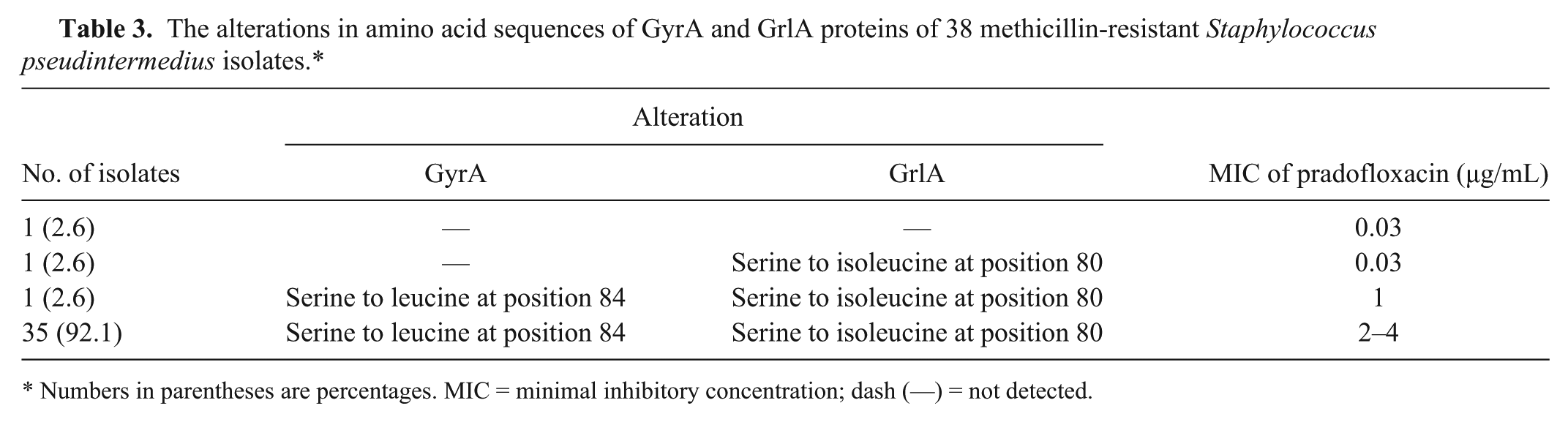

Numbers in parentheses are percentages. MIC = minimal inhibitory concentration; dash (—) = not detected.
Discussion
Pradofloxacin, a novel third-generation veterinary fluoroquinolone, is characterized by lower MIC values in comparison to other antimicrobials of this group.18,26,27 However, the available data concerning MIC values of S. pseudintermedius were based only on the results obtained for MSSP isolates. The occurrence of fluoroquinolone resistance among MSSP is relatively rare, in contrast to MRSP isolates.10,17,24 Data presented in this study clarified the currently available data regarding in vitro activity of pradofloxacin.
The majority of the studied MRSP isolates (94.7%, n = 36) did not show growth inhibition zones around disks of ciprofloxacin, enrofloxacin, marbofloxacin, or orbifloxacin. These isolates also exhibited high MIC values of enrofloxacin and ciprofloxacin; therefore, they were classified as resistant to these fluoroquinolones. According to the pradofloxacin disk diffusion interpretive criteria, 33 of these 36 MRSP isolates were identified as resistant to pradofloxacin. Two of the remaining 3 MRSP isolates were intermediate in disk diffusion test and resistant by MIC values. One isolate was intermediate by disk diffusion test, and also by microdilution test. However, the analysis of amino acid sequence of QRDRs showed the presence of mutations associated with resistance to fluoroquinolones in all 36 isolates showing resistance or intermediate resistance to pradofloxacin by MIC. The questionable identification of pradofloxacin resistance of 3 MRSP isolates was also proved on the scattergram, where these 3 isolates fell in the very minor error category (classified resistant by one method and intermediate by the other method). The other 33 isolates showed a MIC of pradofloxacin ≥2 µg/mL, so they were considered resistant, in accordance with the disk diffusion testing results. 20
Partial nucleotide sequences of gyrA and grlA were analyzed to detect the presence of mutations that are associated with fluoroquinolone resistance. The majority of examined MRSP isolates (94.7%) showed 1 mutation in gyrA and 1 in grlA, resulting in the following amino acid sequence changes: leucine to serine at position 84 in the GyrA protein and isoleucine to serine at position 80 in the GrlA protein, respectively. The same alterations in the QRDRs were also observed among fluoroquinolone-resistant MSSP, S. aureus, and coagulase-negative staphylococci.9,14,21,23 Additionally, 1 isolate in this study showed a single alteration in the GrlA protein, but exhibited low MICs for all fluoroquinolones, including pradofloxacin. A level of resistance in staphylococci isolates with a single substitution in 1 of 2 target enzymes may be affected by the affinity of a particular fluoroquinolone for these enzymes. Therefore, for “dual targeting” (targeting both DNA gyrase and topoisomerase IV) pradofloxacin, the susceptibility of bacteria is only slightly reduced when single alterations in GrlA or GyrA proteins are observed. However, any additional alterations may generate high MIC values. 12 This mechanism has been described in the case of S. aureus resistant to other newer generation fluoroquinolones such as sitafloxacin and moxifloxacin.13,30
Low MICs of pradofloxacin for various veterinary pathogens, including S. pseudintermedius, have been reported previously. A 2013 study 26 confirmed significantly lower MIC values for pradofloxacin as compared to the MICs of 6 other fluoroquinolones for S. pseudintermedius of canine and feline origin, but only MSSP strains were studied. Moreover, only 6 MSSP isolates out of 177 tested exhibited MIC for pradofloxacin of ≥2 µg/mL as well as mutations in gyrA and grlA genes. 26
The results of our study demonstrated the in vitro resistance of canine MRSP isolates to pradofloxacin. This study also showed that single MRSP isolates may be susceptible to pradofloxacin, as well as to other fluoroquinolones. It is remarkable that some MRSP isolates with confirmed alterations in amino acids sequences of QRDRs of both GyrA and GrlA proteins may be recognized as intermediate resistant to pradofloxacin by disk diffusion test and by MIC values. In conclusion, our study indicated an excellent correlation between the results of in vitro pradofloxacin activity testing obtained by disk diffusion and by broth dilution method. The “intermediate” result to pradofloxacin for MRSP isolates resistant to other fluoroquinolones should be interpreted cautiously.
Footnotes
Acknowledgements
We thank Carolin Ludwig for help in the susceptibility testing to pradofloxacin. We are grateful to Barbara Chojnacka and Alicja Grzechnik for excellent technical assistance and to Artur Zalewski for his support in the preparation of this study.
Authors’ contributions
All authors contributed to conception and design of the study; contributed to acquisition, analysis, and interpretation of data; drafted the manuscript; critically revised the manuscript; gave final approval; and agreed to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
a.
American Type Culture Collection, Manassas, VA.
b.
Microbank bacterial and fungal preservation system (14.002), Pro-Lab Diagnostics Inc., Toronto, Ontario, Canada.
c.
Bayer Animal Health GmbH, Leverkusen, Germany.
d.
E-test, bioMérieux Sverige AB, Askim, Sweden.
e.
Mueller Hinton 2 agar, bioMérieux SA, Mercy l’Etoile, France.
f.
BBL Sensi-Disc, BD, Franklin Lakes, NJ.
g.
Marbofloxacin, Marbocyl, Vetoquinol S.A., Veterinary Pharmaceuticals, Lure, France.
h.
Mastdiscs, Mast Diagnostics, Mast Group Ltd., Merseyside, United Kingdom.
i.
3730xl DNA analyzer, Applied Biosystems, Foster City, CA.
j.
Chromas Lite version 2.1.1 program, Technelysium Pty Ltd., Brisbane, Queensland, Australia.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: All susceptibility testing to pradofloxacin were performed and funded by Bayer Animal Health GmbH.
