Abstract
A Mycoplasma iowae real-time polymerase chain reaction (PCR) assay
using primers and probes targeting the 16S rRNA gene was developed and
field-validated in this study. The assay specifically identified M.
iowae with a detection limit of 80 colony-forming units (cfu) per turkey
cloacal swab sample (3.2 cfu per PCR reaction). It was validated by testing 154 field
turkey cloacal swab samples in parallel with culture isolation. The diagnostic
sensitivity of the
Mycoplasma iowae can cause mortality in chicken and turkey embryos, and airsacculitis in turkeys, 9 although some strains may not cause significant clinical problems (Arpad Ferencz, personal communication, May 8, 2007). To establish and maintain M. iowae-free stock or to eradicate the pathogen from a flock rapid and reliable diagnostic methods are required. The traditional culture isolation of M. iowae is time-consuming and complicated, requiring 2–3 weeks to complete because of the slow-growing nature of M. iowae, and it can be hindered by the overgrowth of background bacteria. It has been difficult to develop serologic tests for M. iowae because of its antigenic heterogeneity (there are 6 serotypes: I, J, K, N, Q, and R) and its phenotypic variation. 4 There has been no serologic test available commercially. In the past decade, gel-based polymerase chain reaction (PCR) assays have been developed for rapid detection of M. iowae. 1,5,6,11 Recently, real-time PCR has been used widely to detect bacterial agents. Compared with the traditional gel-based PCR, real-time PCR can be more rapid, more specific, and less prone to contamination. This technology has not been described for the detecting M. iowae.
All bacteria used in this study were obtained from the American Type Culture Collection or were isolated and characterized at the Animal Health Laboratory (AHL) at the University of Guelph (Table 1). The Mycoplasma strains were stored at −70°C in Hayflick's broth a , 8 or modified Friis broth 7 containing pig serum (20%), nicotinamide adenine dinucleotide (NAD, 2 mg/100 ml), and other bacteria in brain heart infusion broth containing 30% glycerol. b Before use, the Mycoplasma strains were propagated in Hayflick's broth a , 8 containing 20% pig serum; other bacterial strains were cultured in trypticase soy broth. a
In 2006 and 2007, 154 cloacal swabs were collected from turkey flocks suspected of having M. iowae infection. The samples were delivered to the AHL in modified in viral transportation medium (31 swabs) or Ortmayer broth (123 swabs), a Mycoplasma culture broth similar to Hayflick's broth except that it contained no glucose and contained NAD (10 mg/liter). The same swabs were used for DNA extraction and mycoplasma culture set up within 48 hr after sample collection. Cloacal swabs were inoculated to Friis agar plates directly and Friis broth containing 20% pig serum for enrichment. The broths were incubated at 35°C aerobically for 21 days. Subcultures were made at 48 and 96 hr. Thereafter, if color change was observed, 10 μl of the broth was inoculated onto a Friis agar plate. Agar plates were incubated at 35°C in 5% CO2 and 90% humidity, and were examined for growth every 48 hr. If the samples had bacterial overgrowth, the inoculated broth was filtered through a 0.45-μm filter to reduce bacterial contamination, and then the filtrate was used to inoculate broth and agar plates. The quantity of Mycoplasma in the samples was determined as “occasional” if an average of 1–2 colonies was observed in each 40 × magnification field, “few” if 2–10 colonies, “moderate” if 11–50 colonies, and “large” if >50 colonies, or “positive on enrichment” if M. iowae was isolated only after enrichment in broth culture. If no Mycoplasma colonies were observed in original and enrichment cultures, the sample was considered as M. iowae not isolated. If Mycoplasma colonies were observed, the species in the positive samples was identified by an indirect fluorescent antibody test (IFAT) 8 using antisera specific for M. gallisepticum, M. meleagridis, M. synoviae, and M. iowae serotype I (all produced at the AHL), and M. iowae serotypes J, K N, Q, and R (provided by Dr. Erno, Institute of Medical Microbiology, University of Aarhus, Denmark).
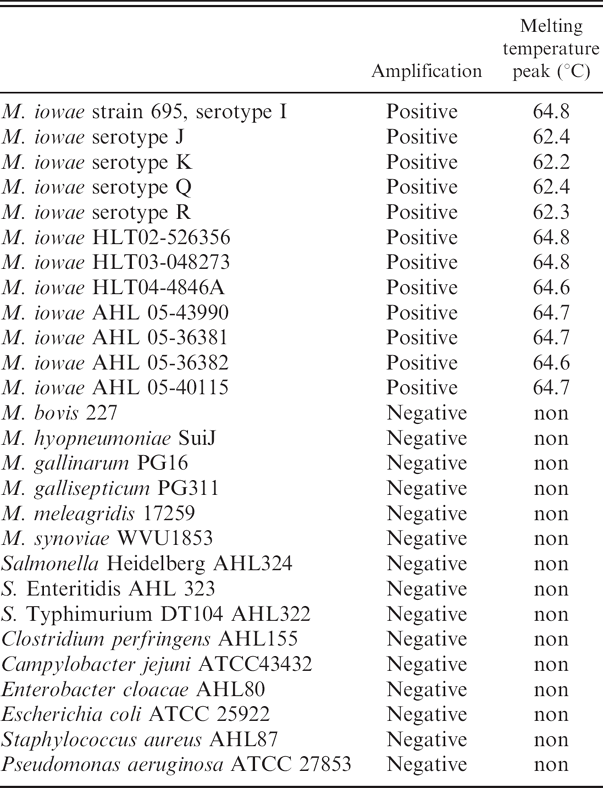
Testing bacterial pure cultures using Mycoplasma iowae real-time polymerase chain reaction.
DNA was extracted from broth cultures of control organisms and from cloacal swabs (after streaking on a plate for culture) using a DNA extraction kit c according to the manufacturer's instructions with some modifications for cloacal swab samples. Buffer ATL c (360 μl) was added to the tube containing the swab, then vortexed for 30 sec. The liquid was transferred to a microcentrifuge tube containing 20 μl of proteinase K, and incubated at 55°C for 30 min, after which the contents were transferred to a spin column, washed twice with washing buffer, c and eluted in a final volume of 50 μl. For larger numbers of samples, the tube containing the swabs and transport medium were vortexed for 30 sec and 200 μl was removed for DNA extraction using an automated extraction instrument d with an elution volume of 100 μl. Both DNA extraction methods had similar detection limits determined by testing the same serial dilution of M. iowae spiked on cloacal swabs (data not shown).
To select primers and probes, the sequences of the partial 16S rRNA gene around 1,500 base pair (bp) of 3 AHL M. iowae isolates were obtained as described previously (EF447271, EF44722, and EF447273). 2 These sequences, together with mycoplasmal and other bacterial 16S rRNA gene sequences retrieved from the GenBank were analyzed using a sequence multiple alignment software, e and primers and probes were selected using real-time PCR primer and probe design software. f PCR primers (MI-F 5′-TGACGG TATCCTTTGAATAAGTATCG and MI-R 5′-ACTCTA GATTTGCAGTTTCCAATG) and a pair of hybridization probes or fluorescence resonance energy transfer probes 3 (MI-FLp 5′-GAGAGCGCGGGCGGA-flurorescein and MI-LCp 5′-LC Red640–TGCAAGTCTGGTGTGAAA TACAACTGCTTAACGG-phosphate) were designed to amplify and identify a 189-bp fragment specific for the M. iowae 16S rRNA gene in variable region 4 (Fig. 1). The specificity of primers and probes was verified by Blast search against the sequences deposited in GenBank. The PCR was performed with a real-time PCR instrument g in a 20.0-μl reaction volume containing 2.75 mM of MgCl2, 2.0 μM of forward primer MI-F, 2.0 μM of reverse primer MI-R, 0.2 μM each of fluorescein probe MI-FLp and LCRed 640 probe MI-LCp, 2.0 μl of LC DNA Master Hyb Probes, 2.0 μl of DNA template, and 9.9 μl of PCR-grade water. All reagents were purchased from a real-time PCR reagent manufacturer, h and the primers and probes were manufactured commercially. i The PCR reaction started with a DNA denaturing and polymerase activating temperature of 95°C for 10 min, followed by 45 cycles of denaturing at 95°C for 1 sec, annealing at 60°C for 6 sec. and extension at 72°C for 8 sec. Temperature transition was 20°C/sec for denaturing and annealing, and 2°C/sec for extension. Fluorescence was acquired at the end of annealing (single mode) using a channel setting of 640 nm/530 nm. The melting temperature of the probe and PCR product hybrid was measured using a single cycle of 95°C for 120 sec (20°C/sec), 65°C for 60 sec (20°C/sec), 50°C for 60 sec (20°C/sec), and 80°C for 0 sec (0.1°C/sec). The fluorescence for the melting temperature (TM) measurement was acquired with a continuous mode using the same channel setting. Quality, quantity, and TM were calculated by the analytic software of the real-time PCR machine (automatic mode). Statistics on field validation data was performed using commercial software. j
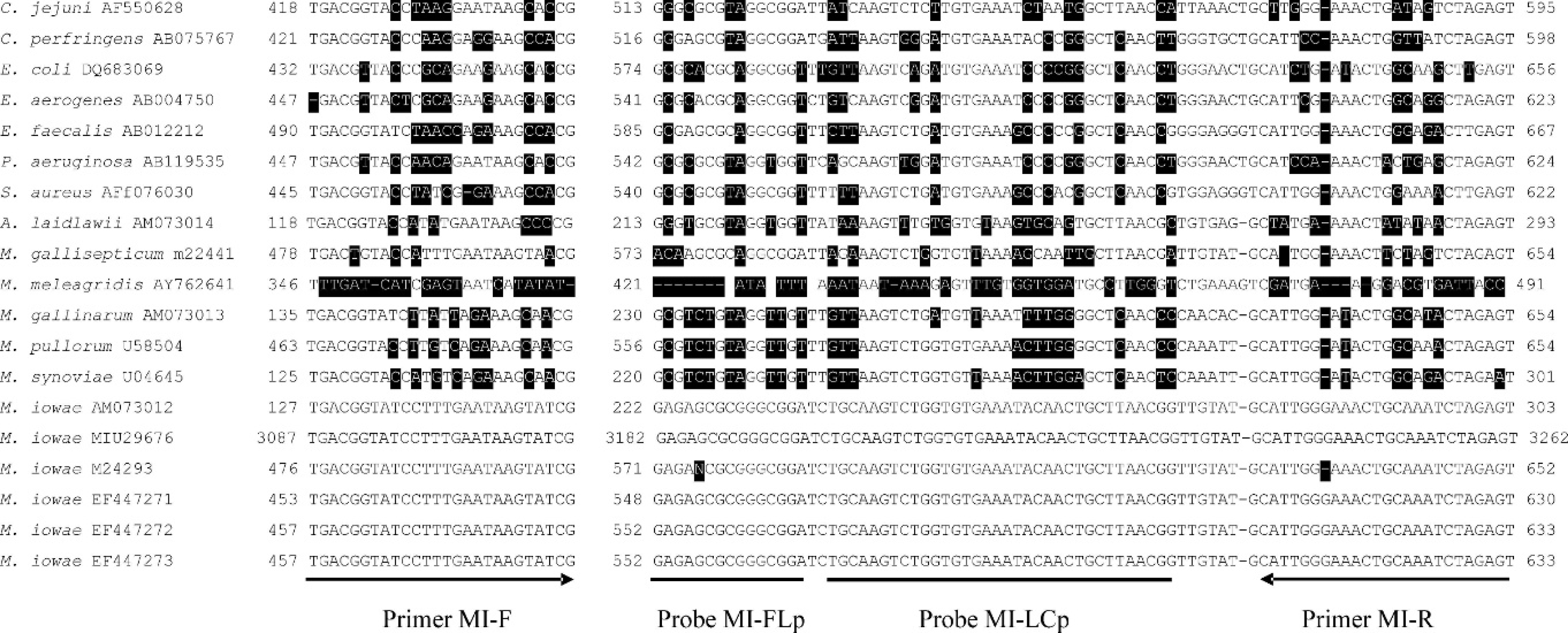
Mycoplasma iowae real-time polymerase chain reaction primers and probes. Multiple sequence alignment was performed on 16S rRNA gene sequences of M. iowae and other avian mycoplasma and bacterial pathogens. Each line consists of species, GenBank accession number, location of first base pair (bp) of the forward primer, sequences aligned to the forward primer, the location of the first bp of the probe sets, the sequences aligned to the probe sets and the reverse primer, and the location of 5′ end of reverse primer. Locations of primers and probes are displayed under the 16S rRNA gene sequence of M. iowae Animal Health Laboratory isolate (GenBank accession number: EF447273). Nonsimilar bases are highlighted.
The specificity of the M. iowae real-time PCR was validated by testing pure cultures of M iowae serotypes I, J. K, Q, and R; different field isolates of M. iowae; and other Mycoplasma and avian bacterial species. All M. iowae strains tested in the present study were real-time PCR positive. In the early stage of method development, the PCR products were analyzed by DNA sequencing and found to have the expected sequences (data not shown). All other mycoplasmal and other bacterial species tested were negative by M. iowae real-time PCR (Table 1). Serial dilutions (8 × 1 to 8 × 106 cfu/swab) of M. iowae were spiked onto cloacal swabs. Swabs were then tested by realtime PCR. The crossing points, the point where the sample's fluorescence curve turns sharply upward, indicating exponential amplification, were automatically determined by the software of the real-time PCR instrument as 20.1 (8 × 106), 22.7 (8 × 105), 26.4 (8 × 104), 30.1 (8 × 103), 33.3 (8 × 102), and 38.6 (8 × 101). The efficiency of the PCR assay was 1.97, very close to 2 (perfect PCR amplification; Fig. 2). Therefore, the detection limit of the real-time PCR assay was determined to be 80 colony-forming units (cfu) per cloacal swab (3.2 cfu per PCR reaction). The standard curve generated from the aforementioned data by the real-time PCR software was used for quantitation of M. iowae concentration in field cloacal samples. The efficiency of swab DNA extraction may vary because of the variation of fecal bearing volume and background bacterial load on the swabs. Therefore, the standard curve may not be perfectly linear and quantitation results may not precisely reflect the actual Mycoplasma concentration. A standard curve generated from serial dilutions of pure DNA, instead of from extracts of culture spiked onto swabs in this study, may be more linear, but it may not reflect the real situation of field swab samples.
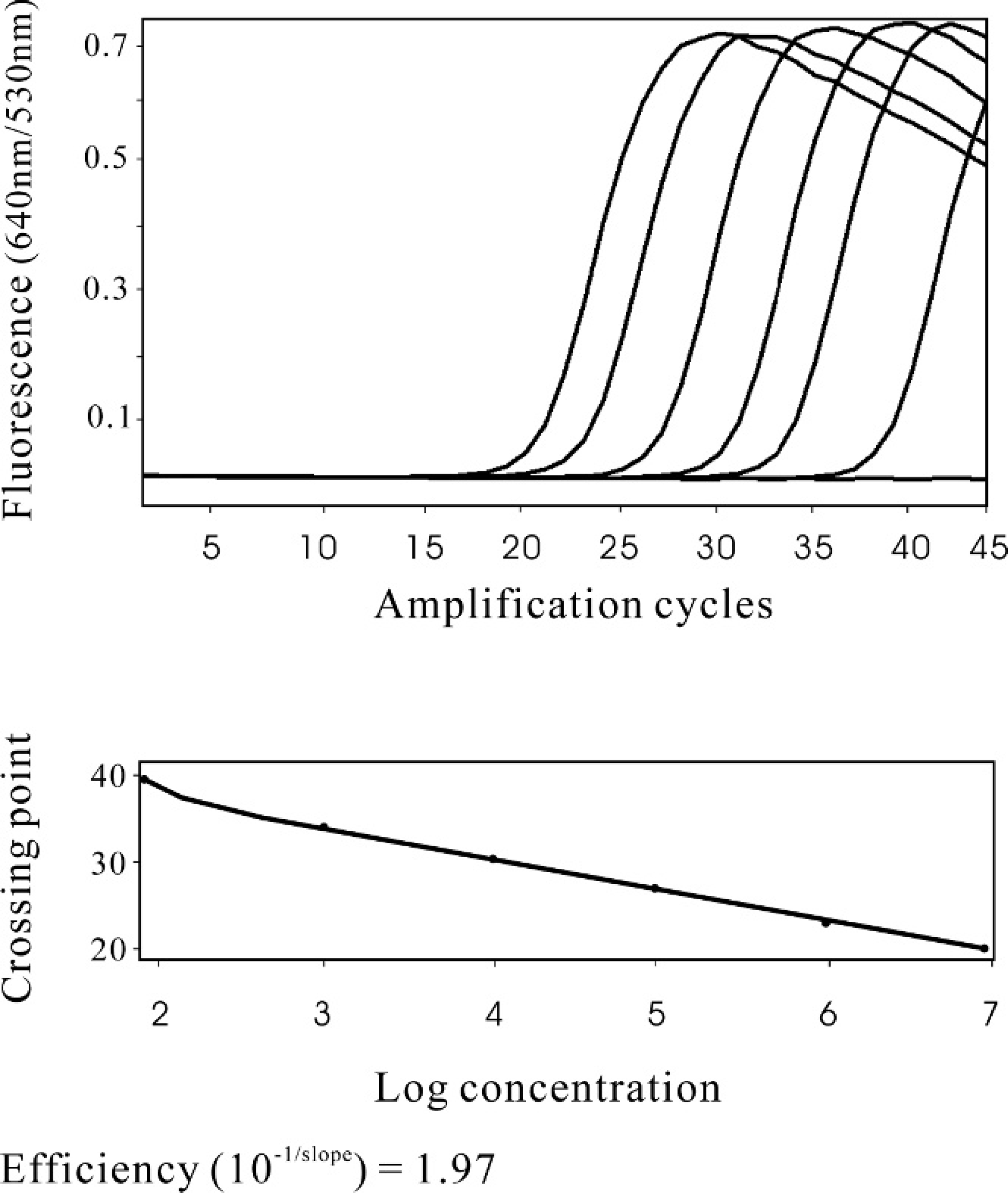
Standard curve of cloacal swab Mycoplasma iowae real-time polymerase chain reaction (real-time PCR). Serial dilutions of M. iowae in cloacal swab samples were tested by the real-time PCR. Upper panel: from left to right, the different amplification fluorescence curves represented the serial dilution of 8 × 106, 8 × 105, 8 × 104, 8 × 103, 8 × 102, and 8 × 101 colony-forming units of M. iowae per swab, respectively. The flat amplification curves include the cloacal swabs inoculated with 0 and 8 × 10-1 M. iowae. Lower panel: Standard curve generated from the above data by the real-time PCR software.
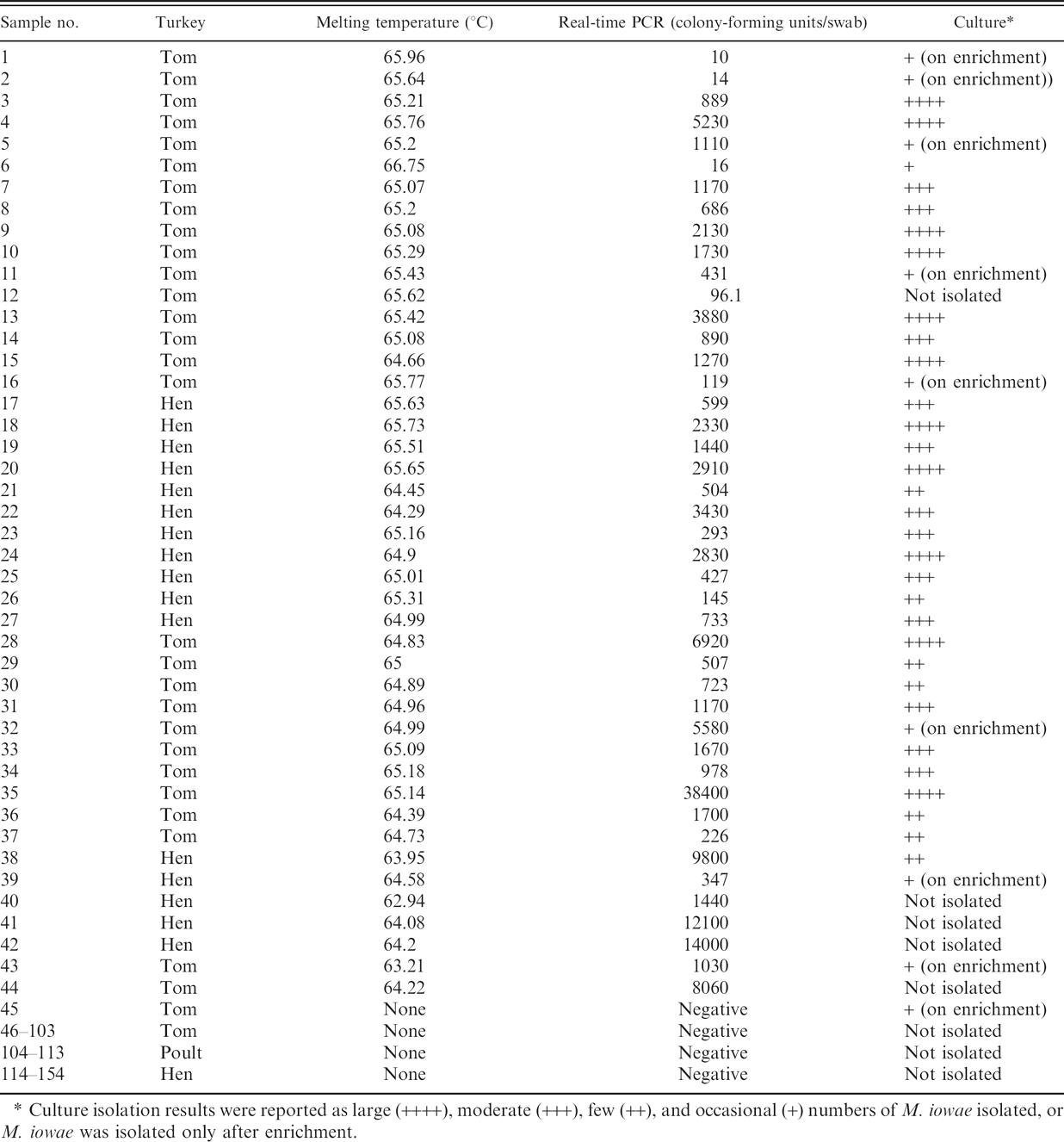
From January 2006 to July 2007, 154 field cloacal swab samples were submitted to the AHL from turkey farms suspected of M. iowae infection with no obvious clinical M. iowae disease signs. The samples were tested by M. iowae real-time PCR in parallel with Mycoplasma culture isolation with results summarized in Table 2. There were 41 positive samples and 107 negative samples as determined by both PCR and isolation. Five samples were PCR positive but negative by culture isolation. Using culture isolation as the gold standard, the diagnostic sensitivity of the PCR was 97.6% and the specificity was 95.5%. The 5 PCR-positive and culture-negative samples were overgrown with bacteria in culture, and the PCR results were confirmed by sequence analysis of the PCR products (data not shown). One sample was negative by real-time PCR but positive by culture isolation (on enrichment only). The Mycoplasma quantity in the swabs ranged from 10 to 5.6 × 105 cfu per swab as determined by the real-time PCR, and most quantitative results correlated well with culture semiquantitative results (Table 2). There was no difference between the concentration of M. iowae quantities in hen or torn swabs (P = 0.9) as determined by real-time PCR.
The real-time PCR developed in this study was rapid and cost-effective with different sample sizes. Within one day. one technician can test at least 30 samples using the manual DNA extraction method and at least 100 samples with automate DNA extraction. The assay could be further automated with a large-scale real-time PCR instrument. It takes 2 to 3 weeks to complete Mycoplasma culture isolation of 30 to 100 samples. An additional 1 to 2 days is required for Mycoplasma species identification using an FA test for 10 to 15 positive samples. Material cost was similar for both PCR and culture for negative samples, but 2 to 3 times more expensive using culture method on positive samples because of the usage of additional culture media and antisera for IFA testing. A disadvantage of real-time PCR is that it identifies M. iowae to the species level only and cannot differentiate serotypes; therefore, culture isolation is required to obtain an isolate if further characterization is necessary.
Several conventional (gel-based) PCR assays for M. iowae detection have been described previously targeting the 16S rRNA gene 1,6 or DNA fragment of unknown function. 12 These PCR assays were found to be specific and sensitive, being able to detect 0.1 pg to 100 ng of M. iowae DNA. 1,6,12 However, most published M. iowae PCR assays were not validated with field samples. 1,6 In another study, instead of using electrophoresis, a species-specific oligonucleotide probe was described that was able to detect the PCR amplification product of 80 cfu M. iowae, with some false-positive results using culture as the gold standard. 5 The real-time PCR assay developed in this study had a detection limit of 80 cfu per cloacal swab (3.2 cfu per PCR reaction), but compared with the gel-based PCR or PCR-probe-detection approach, there was no post-PCR handling in real-time PCR, which saved time and reduced the opportunity for contamination. In addition, real-time PCR using probes can increase the specificity of the assay, and the TM of the probe and amplicon hybrid can be measured for further verification of PCR specificity. In this study, the specific TM varied from 62.2 to 65.9°C. It has been described that different sample salt concentrations due to the difference in sample type and DNA extraction may cause variation in TM, which may be overcome using internal TM control and high resolution melting techniques. 10 Another advantage of real-time PCR is that it can be more accurate and easy to quantitate the target DNA concentration. In the present study, most of real-time PCR quantitation results correlated well with culture isolation. However, it should be noted that the quantitation in this study was calculated for each swab; therefore, it may not be precise because of the possible inconsistency of fecal bearing among different swabs.
Depending on the type of transport medium used for sample delivery, overgrowth of bacteria can be problematic for culture isolation, but it did not seem to have a negative impact on the real-time PCR assay. Some samples (29 of 154), including 6 PCR-positive and culture-positive samples, required filtration to remove other bacteria to enable mycoplasma culture isolation. It was found that using modified Ortmayer medium (containing antibiotics and thallium acetate) as a transport medium resulted in much less bacterial contamination and fewer samples (10 of 123) required filtration compared with using viral transport medium that contained antibiotics without thallium acetate (20 of 31 required filtration). Four PCR-positive and culture-negative samples were submitted in viral transport medium and were overgrown with bacteria even after filtration. Only one sample submitted in modified Ortmayer medium was PCR-positive but culture-negative, and the samples was overgrown with bacteria.
Field cloacal samples tested by Mycoplasma iowae real-time polymerase chain reaction (real-time PCR) and culture
Culture isolation results were reported as large (++++), moderate (+++), few (++), and occasional (+) numbers of M. iowae isolated, or M. iowae was isolated only after enrichment.
Multiplex PCR 11 or single PCR assays with species-specific oligonucleotide probe assays 5 have been developed to detect multiple avian Mycoplasma species in the same assay. Future studies can be carried out to develop similar assays using real-time PCR with multiple probes or using an array of different real-time PCR assays.
Acknowledgements. The authors thank Patricia McRaild and Rebeccah Travis for their assistance in some experiments and Karin Raymond (Roche Diagnostics) for assistance in interpreting various real-time PCR results.
Footnotes
a.
Difco, Detroit, MI.
b.
Becton Dickinson, Cockeysville, MD.
c.
Qiagen DNeasy Tissue kit, Qiagen, Mississauga, Ontario, Canada.
d.
MagNA Pure LC and MagNA Pure LC DNA Isolation Kit I, Roche Diagnostics, Montreal, Quebec, Canada.
e.
Vector NTI 10.1, Invitrogen, Burlington, Ontario, Canada.
f.
LightCycler Probe Design Software 2.0, Roche Diagnostics, Montreal, Quebec, Canada.
g.
LightCycler 2.0, Roche Diagnostics, Montreal, Quebec, Canada.
h.
Roche Diagnostics, Montreal, Quebec, Canada.
i.
TIB Molbiol, Adelphia, NJ.
j.
GraphPad InStat V3.06, GraphPad Software, Inc., San Diego, CA.
