Abstract
The suitability of nested reverse transcription polymerase chain reaction (nRT-PCR) to detect betanodavirus in blood samples from naturally infected Senegalese sole (Solea senegalensis) was evaluated in comparison with other diagnostic methods. Results indicated that histologic examination of brain lesions could be regarded as the most consistent indicator of nodavirus infection in this species. The nRT-PCR showed low to moderate levels of detection; the best values were obtained in brain samples followed by blood samples. Inoculation of SSN-1 and SAF-1 cells with fish samples did not cause cytopathic effect, although virus was detected by reverse transcription polymerase chain reaction in approximately 25% of the SSN-1 inoculated wells. The efficiency of detection of the viral genome was dramatically increased by the use of nRT-PCR, reaching 90.6% of positives in brain samples and 84.4% in blood samples. The sensitivity and the negative predictive value of nRT-PCR in blood samples were slightly lower than those obtained using brain samples. Nevertheless, it is suggested that the advantage of being able to perform diagnosis on live fish adequately counterbalances the slightly lower sensitivity of nRT-PCR on blood samples. This technique is proposed as a useful tool, not only for the selection of nodavirus-free breeders but also to check the fish status during ongrowing.
Viral encephalopathy and retinopathy, or viral nervous necrosis (VNN), is considered to be one of the most serious viral diseases affecting not only larvae but also juvenile marine fish throughout the world. 14 The causative agents of the disease are viruses belonging to the genus Betanodavirus (family Nodaviridae). Betanodaviruses are nonenveloped virions with icosahedral symmetry, about 25 nm in diameter, with a viral genome consisting of 2 single-stranded (ssRNA) segments. The main target organ of these viruses is the central nervous system, including brain, spinal cord, and retina. Clinical signs of the disease include lack of appetite, changes in pigmentation, and neurologic signs such as abnormal swimming. Both horizontal 5,15 and vertical transmission of nodavirus have been demonstrated in different fish species. 3,9,15
In general, the diagnosis of VNN relies on histopathologic identification of necrosis and vacuolization in brain and retina, combined with positive immunohistochemical assessment. However, other diagnostic procedures, such as enzyme-linked immunosorbent assay (ELISA) 4,14 and reverse transcription polymerase chain reaction (RT-PCR). have been developed in recent years. 7,14,19
The use of PCR-based technology applied to blood samples has proven to be an excellent diagnostic tool for other fish viruses 6,8,13 and has been previously used to detect betanodavirus in sea bream and sea bass. 7 The present study reports the suitability of nested RT-PCR (nRT-PCR) for detecting betanodavirus in blood samples of naturally infected Senegalese sole (Solea senegalensis) compared with RT-PCR using other tissue samples, isolation in cell culture, cell culture combined with RT-PCR, and histologic visualization of the lesions.
In November 2005, sole weighing 145.4 ± 86.0 g and reared in a 4-m2 tank supplied with flow-through water (11.8 ± 1.3°C) at the facilities of the CCMAR-Centro de Ciências do Mar (University of Algarve, Portugal) started to show mildly chronic mortality, anorexia, spiral swimming, and ulceration in the skin. In a first sampling performed in December 2005, 3 moribund fish were individually processed for viral detection by RT-PCR using specific primers for betanodavirus, Infectious pancreatic necrosis virus (IPNV), Infectious hematopoietic necrosis virus (IHNV), and Viral hemorrhagic septicemia virus (VHSV) following the procedure described as follows. All fish were positive only for betanodavirus.
In February 2006, 32 fish were individually sampled both antemortem (blood samples) and postmortem (internal organs). Blood samples were obtained by puncturing the caudal vein (200–500 μl from each fish); the samples were immediately heparinized. After immersion in ice, individual fish were necropsied, and kidney, spleen, and brain were aseptically collected from each fish. To evaluate the efficiency of the diagnostic techniques, 8 apparently healthy fish kept in a different tank were also sampled.
For histologic analysis, brain samples were fixed in Bouin's fixative, dehydrated in graded ethanol, and embedded in paraffin. Next, 4-μm-thick sections were cut, mounted on glass slides, and stained with hematoxylin and eosin. Bacterial isolation was accomplished by inoculating samples of kidney, spleen, and liver onto tryptone soy agar supplemented with 1% (wt/vol) NaCl (TSA-1) a and TCBS (thiosulfate-citrate-bile salts-sucrose) agar a (Vibrio selective agar) and incubating the samples at 25°C for 24 hr. Pure cultures of the different colony types were subjected to taxonomic analysis by standard morphologic, physiologic, and biochemical tests. Two cell lines, SSN-1 (derived from striped snakehead, Channa striatus)> and SAF-1 (derived from gilhead sea bream, Spams aurata), and 2 tissue samples from each fish (brain and a mix of kidney and spleen) were used for viral isolation. Samples were processed as previously described, 1 and inoculated (diluted up to 10-1 and 10-2) in duplicate onto SSN-1 and SAF-1 cells grown on 24–well plates. Infected monolayers were incubated at 20°C and 25°C and examined daily for the presence of cytopathic effects (CPEs). Every 15 days one of the wells inoculated with each sample was subjected to blind passages (up to 2 passages) whereas the other well was maintained up to a maximum of 45 days.
To extract RNA from blood and other fish tissues. QIAamp RNA Blood Mini Kit a and RNeasy Mini Kit b were used, respectively, following the manufacturer's instructions. Primer pairs used for RT-PCR were BA4D-BA4U for IPNV, 6 cm3a-cm3b for VHSV, 13 cm4a-cm4b for IHNV (5′AGATCGGGGCGGTGCTTAGA-3′, 5′-CCATCGCCATCGTACTTGAGGACA 3′), and F2–R3 18 for nodavirus. Complementary DNA (cDNA) synthesis and amplification rounds were carried out using SuperScript III Platinum One-Step Quantitative RT-PCR System Kit c as previously described. 13 The nRT-PCR for nodavirus detection was performed using 3 μl from the RT-PCR reaction, the specific primers F21–R31 (5′-GATTTCGTTCCATTCTCTTG-3, 5′-AGTGTCTCCAG CTTTCTTCT-3′) using the same protocol as described. The PCR products were analyzed by electrophoresis on a 2% agarose gel. All RT-PCR and nRT-PCR assays were repeated 2 or 3 (in the case of negative samples) times.
The performance of PCR-based methods for nodavirus detection was evaluated by sensitivity (probability of a positive test result in a nodavirus-infected fish) and specificity (probability of a negative result in a noninfected fish) using results from histopathologic examination as the gold standard. In addition, positive and negative predictive values for each test were determined. Positive predictive values (PPV) are defined as the probability for a fish testing positive of actually having nodavirus infection and the negative predictive values (NPV) are defined as the probability for an individual testing negative of not being infected.
The results of the histopathologic observation showed VNN-specific histologic lesions, such as necrosis with vacuolization, in all brain samples (Fig. 1). Bacterial growth was detected in 15 of 32 fish analyzed (46.87%). Bacterial colonies were identified as Vibrio fisheri (5 fish). Vibrio splendidus (3 fish), Photobacterium damselae subsp. piscicida (2 fish), Pseudomonas aeruginosa (3 fish), and Vibrio sp. (1 fish). In addition, one mixed culture of V fisheri and V splendidus was obtained from one fish. No bacteria were isolated from the control group. None of the samples produced CPE in first passage on SSN-1 or SAF-1 15 days after inoculation. After 2 blind passages, there was no evidence of CPE in both cell lines. All second blind passages in SSN-1 or SAF-1 cells were examined by RT-PCR to determine whether any of the samples harbored noncytopathic virus. Only SSN-1 cells yielded positive results by RT-PCR in 17 of the 32 monolayers inoculated with brain samples and 8 inoculated with kidney/spleen samples (Table 1). No virus was isolated and no RT-PCR product was obtained from the cells inoculated with samples from the control fish.
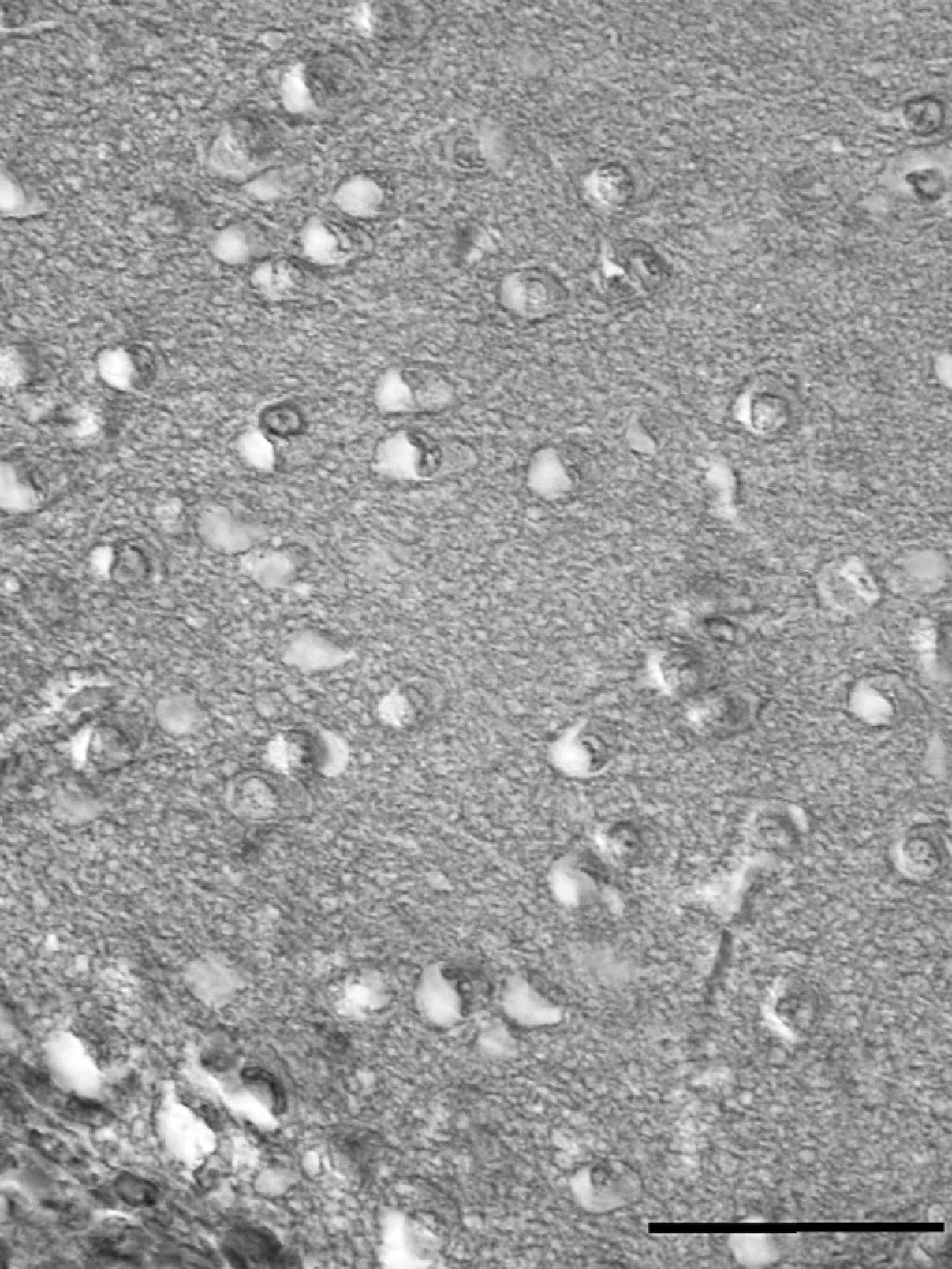
Light micrograph showing vacuolation in the brain of Solea senegalensis. Hematoxylin and eosin. Bar = 25 μm.
The results of betanodavirus detection by RT-PCR are summarized in Table 1. A specific RT-PCR amplification product of 420 base pair (bp; Fig. 2) was obtained from 18 fish, which represents 56.3% of the fish population. Brain was the most effective organ for viral detection (46.9% positive) followed by blood with 28.1% of positives, whereas only 15.6% of the kidney/spleen samples were positive. These percentages were slightly increased if RT-PCR was applied onto cell culture infected with the tissue homogenates: up to 53% from brain and 25% from kidney/spleen (blood samples were not subjected to cell culture isolation). No RT-PCR amplification was obtained from the fish used as negative control (data not shown). As expected, all the samples considered positive by RT-PCR were also found to be positive by nRT-PCR, which detected considerably more positive samples (Table 1). In fact, the specific amplification product of 156 bp (Fig. 2) was obtained from all 32 fish, although using different types of samples. The highest level of detection, as occurred in the RT-PCR analysis, was achieved in the brain (90.6%), followed by blood and kidney/spleen, which had the same detection capacity (84.4%). Sixty-nine percent of the fish were regarded as positive for nodavirus using the 3 types of samples. All fish from the control group were negative (data not shown).
Detection of betanodavirus by reverse transcription polymerase chain reaction (RT-PCR) and nested (n)RT-PCR in infected cell cultures and fish tissues.
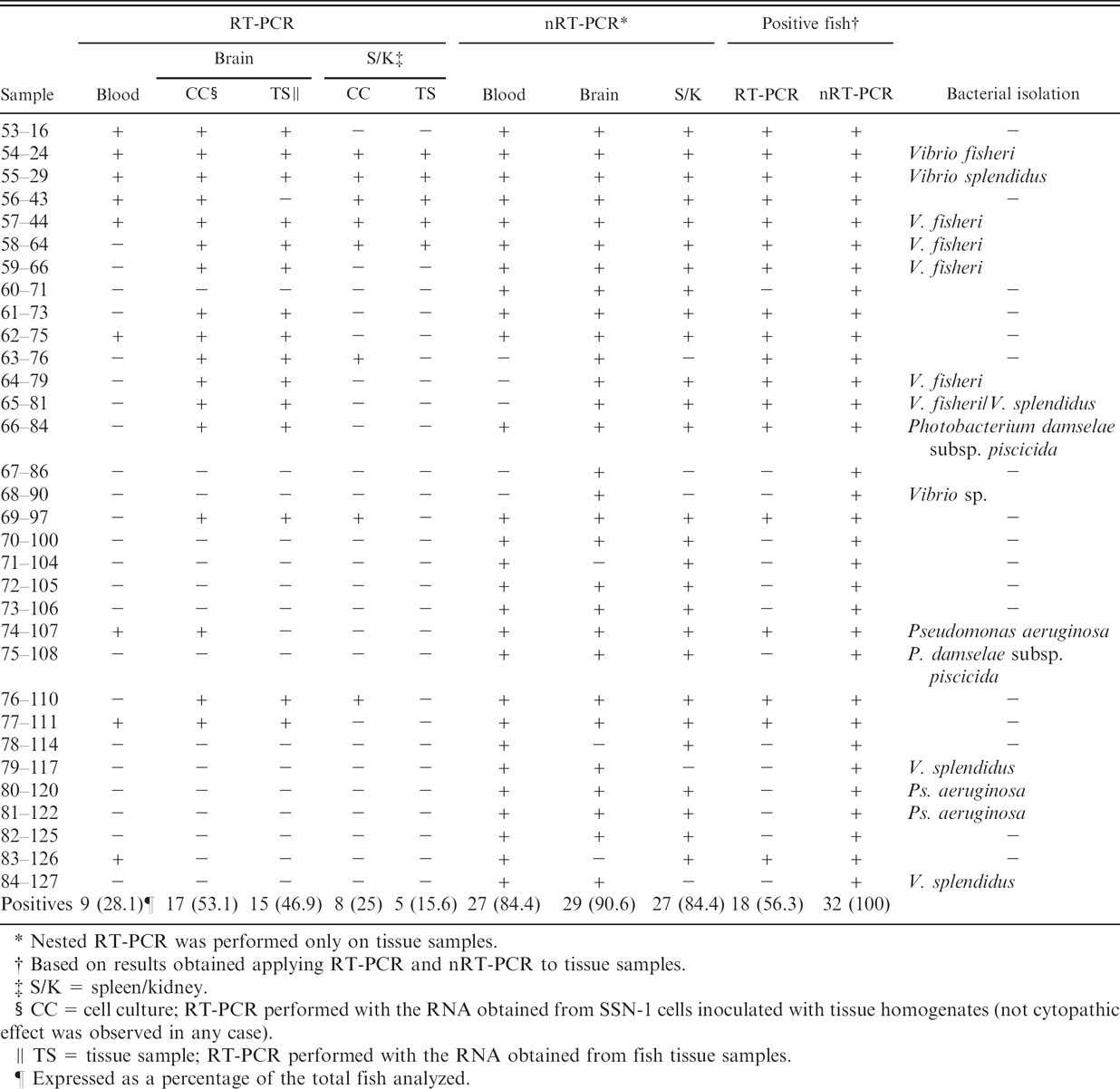
Nested RT-PCR was performed only on tissue samples.
Based on results obtained applying RT-PCR and nRT-PCR to tissue samples.
S/K = spleen/kidney.
CC = cell culture; RT-PCR performed with the RNA obtained from SSN-1 cells inoculated with tissue homogenates (not cytopathic effect was observed in any case).
TS = tissue sample; RT-PCR performed with the RNA obtained from fish tissue samples.
Expressed as a percentage of the total fish analyzed.
The results of the performance evaluation of the diagnostic tests assayed are shown in Table 2. All RT-PCR and nRT-PCR assays showed the maximum specificity values (100%), resulting from the absolute absence of false-positive results; PPV was also 100%. As expected, the highest sensitivity was obtained with nRT-PCR applied to brain samples with a value of 91%. The lowest level was yielded by RT-PCR applied to kidney/spleen samples (16%). The NPV ranged from 73% for nRT-PCR in brain tissues to 32% for RT-PCR applied to kidney/brain samples. Nested RT-PCR applied to blood and kidney/spleen samples provided identical sensitivity and PPV values (84 and 62%, respectively).
Results obtained indicated that the observation of histologic lesions in brain could be regarded as the most consistent indicator of nodavirus infection in sole (100% detection). However, this procedure is time consuming and not available in all laboratories. Use of RT-PCR showed low to moderate levels of detection, but the highest percentage of positives was obtained also using brain samples. It has already been reported that samples positive for nodavirus by immunohistochemical observation can yield a negative result in RT-PCR. 7,19 Those false-negative results have been attributed to the presence of inhibitory components for the PCR reaction in the tissue samples or to differences between the sequences of the primers and different viral strains. The efficiency nodavirus genome detection was dramatically increased by the use of nRT-PCR (90.6% of positives in brain samples, 84.4% in blood and kidney/spleen).
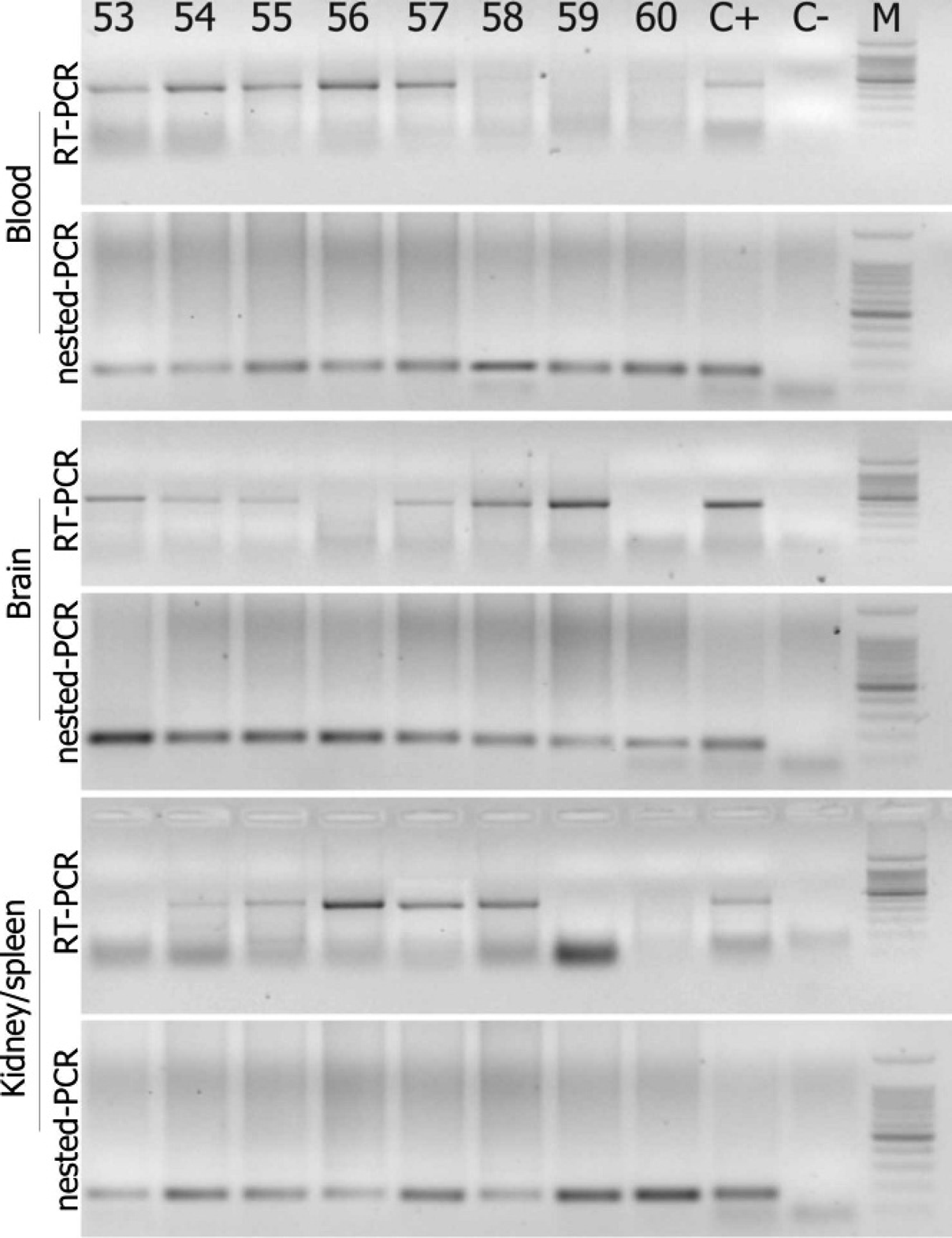
Detection of nodavirus in different tissue samples of Solea senegalensis by reverse transcription polymerase chain reaction (RT-PCR) and nested (n)RT-PCR. The results were obtained after visualization of amplification products in agarose gel under ultraviolet light. Lane numbers correspond to fish numbers: C+ = RT-PCR or nRT-PCR applied to RNA obtained from nodavirus strain 411/96/ERV (420 and 156 base pair [bp]. respectively); C- = RNase-free water; M = 100-bp DNA ladder: 11 fragments ranging from 100 to 1000 bp in 100-bp increments, with an additional band of 1,500 bp.
Inoculation of SSN-1 and SAF-1 cells with fish samples did not cause CPE. However, virus was detected by RT-PCR in some of the SSN-1–infected cells, especially those inoculated with brain samples, allowing the increase of the number of positive results obtained compared with plain RT-PCR performed directly on tissue samples. Viral replication without CPE has already been reported for VNN 10 and other fish viruses. 12 It has been suggested that the low number of nodavirus particles inoculated into cells is the cause of the absence or delay in the development of CPE, and the use of a combined procedure of cell culture and RT-PCR was proposed to detect samples with low virus levels. 11 The results of the present study seem to support the suitability of these combined procedures even though the authors tested second-blind passage cultures instead of the 72-hr cultures reported previously. 11 On the other hand, although it could be expected that coinfection bacteria/nodavirus would act as a stressful factor facilitating viral replication, no improvement was observed on the nodavirus detection by PCR and/or isolation in cell culture in the fish infected with bacteria. Only 9 of 15 fish showing bacterial growth were positive for nodavirus, and only 8 of the 17 fish positive by RT-PCR in cell culture showed bacterial infection, which suggests that no correlation exists between coinfection and viral detection.
Sensitivity, specificity, and predictive values of reverse transcription polymerase chain reaction (RT-PCR) and nested (n)RT-PCR procedures in different types of samples using the results of histopathologic examination as the gold standard (%).
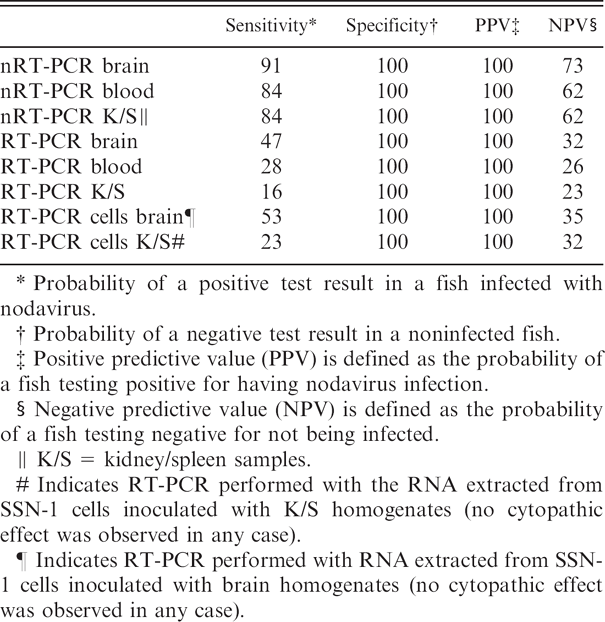
Probability of a positive test result in a fish infected with nodavirus.
Probability of a negative test result in a noninfected fish.
Positive predictive value (PPV) is defined as the probability of a fish testing positive for having nodavirus infection.
Negative predictive value (NPV) is defined as the probability of a fish testing negative for not being infected.
K/S = kidney/spleen samples.
Indicates RT-PCR performed with the RNA extracted from SSN-1 cells inoculated with K/S homogenates (no cytopathic effect was observed in any case).
Indicates RT-PCR performed with RNA extracted from SSN-1 cells inoculated with brain homogenates (no cytopathic effect was observed in any case).
In recent years, there has been an increasing emphasis on the use of antemortem sampling methods that would allow determination of the presence/absence of nodavirus in fish stock, particularly in breeders, without killing the individual fish. Different procedures have been used to analyze broodstock for nodavirus detection, including detection of serum antibodies in sea bass (Dicentrarchus labrax), 2,4 detection of the virus by RT-PCR in gonads and eggs of stripped jack (Pseudocaranx dentex), 16 or detection with both techniques in barfin flounder (Verasper moseri) 20 and sea bass, 3 with contradictory results. It is important to realize that the viruses may not reside in the reproductive organs at all times 17 ; thus, PCR procedures may not detect infection in brood fish if the gonad has not been invaded. 14 In those circumstances it has been suggested that the additional demonstration of specific antibodies in blood or body fluids can be a very useful complementary procedure. 3,14 However, the development of nRT-PCR procedures for detecting this virus in blood samples may represent a significant improvement in speed and sensitivity using minimum amounts of sample. In a previous study, 7 it was concluded that blood samples could be appropriate for selecting nodavirus-free spawners of sea bass and sea bream only when nRT-PCR was used. The results of the present study indicate that nodavirus could be detected in blood samples using plain RT-PCR, albeit at moderate levels (28%), and that the use of nRT-PCR increases the detection to 84%. Although nRT-PCR on antemortem sampling (blood) is less sensitive than on postmortem sampling (brain), it is suggested that the advantage of keeping fish alive counterbalances the slightly lower sensitivity of the antemortem versus postmortem sampling.
Acknowledgements. This work was partly supported by grants PTR1995/0985/DP from the Ministerio de Education y Ciencia (Spain), PGIDIT04RMA261014PR and PGIDIT06RMA008E from Xunta de Galicia, and REDAQUA/SP5.E27/02 programme INTERREG III A, cofunded by FEDER (European Fund for Regional Development; European Commission). The authors wish to thank all members of Aquagroup who participated in the sampling process.
Footnotes
a.
Oxoid Ltd., Basingstoke, Hampshire, UK.
b.
Qiagen Gmbh, Hilden, Germany.
c.
Invitrogen Corporation, Carlsbad, CA.
