Abstract
Viral nervous necrosis (VNN) is a worldwide disease affecting several species of cultured marine fish. In Korea, VNN has been identified in several species of cultured marine fish. In this study, the authors present data of the amplified nested polymerase chain reaction product (420 bp) of 21 nodavirus strains from different species of apparently healthy wild marine fish on the southern coast of Korea. Phylogenetic analysis based on the partial nucleotide sequence (177 bases) of the RNA2 coat protein gene of 21 strains was highly homologous (93–100%) and closely related to that of the known betanodavirus, redspotted grouper nervous necrosis virus. These results indicate that betanodaviruses occur in large populations of wild marine fish in the southern part of the Korean peninsula, suggesting the importance of these subclinically infected fish as an inoculum source of betanodavirus that is horizontally transmitted to susceptible cultured fish species.
Keywords
Introduction
For the past 2 decades, betanodavirus infections that cause viral nervous necrosis (VNN) have emerged as major constraints on the culture and sea ranching of marine fish in almost all parts of the world. More than 30 species of marine fish have been devastatingly affected during the seedling period and the culture process. 28,33 The characteristic lesions of VNN are necrosis and vacuolation of central nervous tissues and retina of the affected larvae and juvenile fishes showing abnormal swimming behavior. The virus contains 2 segments of positive-sense single-stranded RNA. The RNA1 (3.1 kb) and RNA2 (1.4 kb) of striped jack nervous necrosis virus (SJNNV) encode 100 kDa (presumably RNA-dependent RNA polymerase) and a major coat protein of 42 kDa, respectively. 26 Based on a partial nucleotide sequence of coat protein gene, betanodaviruses are divided into 4 genotypes: SJNNV type, tiger puffer nervous necrosis virus (TPNNV) type, barfin flounder nervous necrosis virus (BFNNV) type, and redspotted grouper nervous necrosis (RGNNV) type. 30
VNN was first reported in larvae and juveniles of hatchery-reared Japanese parrotfish Oplegnathus fasciatus. 42 The same viral disease, otherwise called viral encephalopathy and retinopathy, has also been observed in barramundi Lates calcarifer 12 and later in other marine fishes. 27–29,31 Recent studies have also demonstrated that a certain population of apparently healthy wild marine fish carried betanodaviruses and suggested that these wild fish can be a persistent potential source of the virus for cultured fish. 2,11,13 The spread of VNN among populations of cultured marine fish may be attributable to either vertical 1,4,17,40,41 or horizontal transmission. 5,22
In Korea, there were reported incidences of VNN that caused mortality in several cultured fish. In the present study, the authors phylogenetically analyzed 21 apparently healthy wild marine fish that tested positive for betanodavirus from nested polymerase chain reaction (nPCR) examination. This study was conducted to discover if these fish could be an inoculum source of genetically related betanodaviruses that are transmitted horizontally to cultured fishes in the Korean peninsula.
Materials and methods
Fish samples
Six hundred sixty-three samples of apparently healthy wild marine fish were collected in the coastal areas of the Korean peninsula from October 2005 to February 2006. The brains were aseptically collected and stored at −80°C until used.
RNA extraction
Total RNA was extracted from the brain samples with TRIzol reagent, a according to the manufacturer's protocol. In brief, the tissues were homogenized with 900 μl of TRIzol reagent and incubated for 5 minutes at room temperature. Two hundred microliters of chloroform were added to the suspension, incubated for 2 to 3 minutes at room temperature, and then centrifuged at 12,000 × g for 15 minutes at 4°C. The RNA in the aqueous upper phase was precipitated by adding 500 μl of isopropanol; the mixture was incubated for 10 minutes at room temperature and then centrifuged at 12,000 × g for another 10 minutes at 4°C. The pellet obtained after centrifugation was washed with 1 ml of 75% ethanol and centrifuged at 7,500 × g for 5 minutes at 4°C. The RNA pellet was air dried and dissolved in 50 μl of diethylpyrocarbonate-treated water. a
Polymerase chain reaction amplification
The PCR amplification was conducted using primers BNV-RT, BNV-UR1, and BNV-UF1 for reverse transcription polymerase chain reaction (RT-PCR), and BNV-UR2 and BNV-UF2 were used for nPCR. It was designed for the target regions 570 bp and 420 bp of the sevenband grouper nervous necrosis virus (SGNNV) RNA2, as previously described. 13 After reverse transcription using Reverse Transcriptase SuperScript II a at 45°C for 60 minutes. PCR was conducted using Ex Taq polymerase b in a thermal cycler c with 30 cycles of denaturation at 94°C for 30 seconds, annealing at 57°C for 20 seconds, and a final extension at 72°C for 60 seconds. Nested PCR was conducted using the same protocol described above. The PCR products were analyzed by 2% agarose d gel electrophoresis. The RNA from uninfected (negative control) and infected (positive control) redspotted grouper (Epinephelus akaara) larvae was used for RT-PCR and nPCR, respectively.
Virus isolation
Fish samples that tested positive in PCR assays were used for virus isolation using an E-11 cell line 14 previously cloned from the striped snakehead fry (SSN-1) cell line. 9 The E-11 cells were cultured using Leibovitz's L-15 medium a supplemented with 5% fetal bovine serum e in a 75 cm 2 tissue culture flask. f Cultured cells were grown at 25°C for 2 days, then seeded in 24-well tissue culture plates and incubated at 25°C until approximately 60% to 70% confluent. Tissue (brain) samples were weighed and homogenized in 9 volumes of Hanks balanced salt solution, a then centrifuged at 2,000 × g for 10 minutes. The supernatant was passed through a membrane filter g (0.45 μm) and added in 500-μl volumes to wells in 24-well tissue culture plates f containing E-11 cells at 60% to 70% confluency. The results were observed for 2 weeks for any cytopathic effect (CPE).
Genetic sequencing
Twenty-one of 48 positive nPCR products were electrophoresed in 2% agarose gel in 0.5 X TAE buffer (Trisacetate-ethylenediamine tetra-acetic acid). h The band representing the target size was purified from the gel using the Power Gel Extraction kit, b following the manufacturer's protocol in a microcentrifuge. Sequencing of the purified nPCR products was done at the Macrogen Genomic Division, South Korea. i,j
Sequence analysis
As the nPCR products include variable regions and homologous regions commonly observed in betanodaviruses, 15 variable region sequences were selected and used for phylogenetic analysis. The variable region of the SGNNV RGNNV-type RNA2 (177 nt from nt no. 666 to 842) 15 was aligned with the betanodavirus sequences obtained in this study as well as those retrieved from the GenBank database. Extra sequences were trimmed by comparing them with the 177 nt SGNNV RNA2 sequence. All nucleotide sequences and deduced amino acid sequences were aligned using the multiple alignment algorithms in the MegAlign package k to give a phylogenetic tree. The GenBank accession numbers of the known betanodavirus sequences used in this phylogenetic analysis are D30814 (striped jack), D38635 (barfin flounder), D38636 (redspotted grouper), and D38637 (tiger puffer).
Results
Polymerase chain reaction amplification
The specific nPCR products (420 bp) amplified from all 48 samples (21 species) are summarized in Table 1. Positive results were obtained in the brains of the following fish species: Apogon lineatus (Indian perch), Argyrosomus argentatus (white croaker), Caelorinchus multispinulosus (spearnose grenadier), Callionymus lunatus (moon dragonet), Cheilidonichthys spinosus (bluefin searobin), Cynoglossus robustus (robust tonguefish), Doederleinia berycoides (black throat seaperch), Lagocephalus lunaris (moon-tail puffer), Leiognathus nuchalis (silver ponyfish), Lophius litulon (yellow goosefish), Muraenesox cinereus (daggertooth pike conger), Okamejei kenojei (skate rayfish), Pagrus major (red seabream), Pleuronectes yokohamae (marbled sole), Pseudorhombus pentophthalmus (five spots flounder), Scomber japonicus (chub mackerel), Sillago japonica (silver whiting), Takifugu niphobles (grass puffer), Trachurus japonicus (Japanese jack mackerel), Zeus faber (John dory), and Zoarces gilli (blotched eel pout).
Virus isolation
Virus CPEs that were observed from 21 of 48 fish samples are summarized in Table 1. Virus was isolated from the following fish species: Apogon lineatus (Indian perch), Argyrosomus argentatus (white croaker), Caelorinchus multispinulosus (spearnose grenadier), Callionymus lunatus (moon dragonet), Cheilidonichthys spinosus (bluefin searobin), Lagocephalus lunaris (moontail puffer), Leiognathus nuchalis (silver pony-fish), Lophius litulon (yellow goosefish), Pleuronectes yokohamae (marbled sole), Pseudorhombus pentophthalmus (five spots flounder), Scomber japonicus (chub mackerel), and Zoarces gilli (blotched eel pout).
Betanodavirus-positive nested polymerase chain reaction (nPCR) results obtained in 21 different species of wild marine fish.
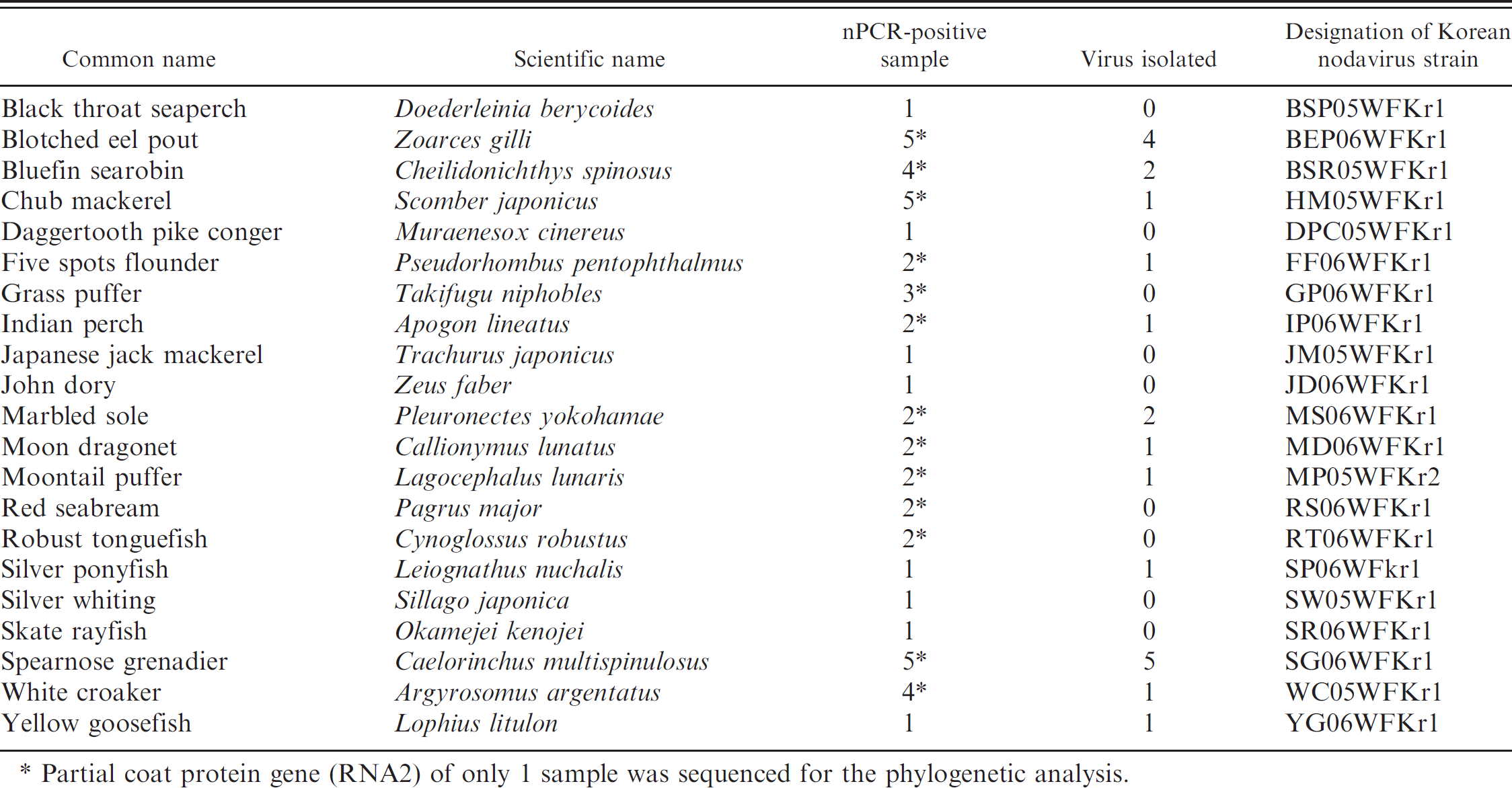
Partial coat protein gene (RNA2) of only 1 sample was sequenced for the phylogenetic analysis.
Sequence and phylogenetic tree analysis
Partial coat protein gene (RNA2) nucleotide sequences of the 21 representative strains were determined (Table 1). The 21 Korean nodavirus strains showed high nucleotide similarity to RGNNV (93.3–100%) but low nucleotide similarity to BFNNV (76.7–80%), SJNNV (63.3–68.3%), and TPNNV (68.3–71.7%; Table 2). The identities of the sequences of nucleotides 666 to 842 of the RNA2 coat protein gene between RGNNV, BEP06WFKr1, BSR05WFKr1, FF06WFKr1, HM05WFKr1, TP06WFKr1, JD06WFKr1, MD06WFKr1, MS06WFKr1, RS06WFKr1, RT06WFKr1, SP06WFKr1, SW05WFKr1, WC05WFKr1, and YG06WFKrl were 100% homologous to each other. BSP05WFKr1, DPC05WFKr1, GP06WFKr1, JM05WFKr1, MP05WFKr2, SG06WFKr1, and SR06WFKrl were related to each other (less than 11% nucleotide difference; Table 2; Fig. 2). Representative Korean nodavirus strains among homologous strains were included for a simplified table (Table 2). The identities of the deduced amino acid sequences encoded in RNA2 coat protein gene (aa 224–281) between Korean nodavirus strains and RGNNV were >93%, while the identities among the Korean nodavirus strains and the other 3 genotypes ranged from 66.3% to 80 %. The alignment of deduced amino acid sequences for the coat protein gene RNA2 showed variability in positions corresponding to aa 254 and 266 between the Korean nodavirus strains and RGNNV genotype. Amino acid at position 254 was changed from glycine (G) to arginine (R) in 5 strains, and position 266 was changed from valine (V) to aspartic acid (D) in 2 strains (Fig. 1). Phylogenetic tree analysis was constructed based on RNA2 coat protein gene. The host range of Korean nodavirus strains includes 21 species of wild marine fish; the phylogenetic analysis revealed that all Korean strains belong to the RGNNV genotype (Fig. 2).
Nucleotide (nt 666–842) percentage similarity among nodavirus strains.*
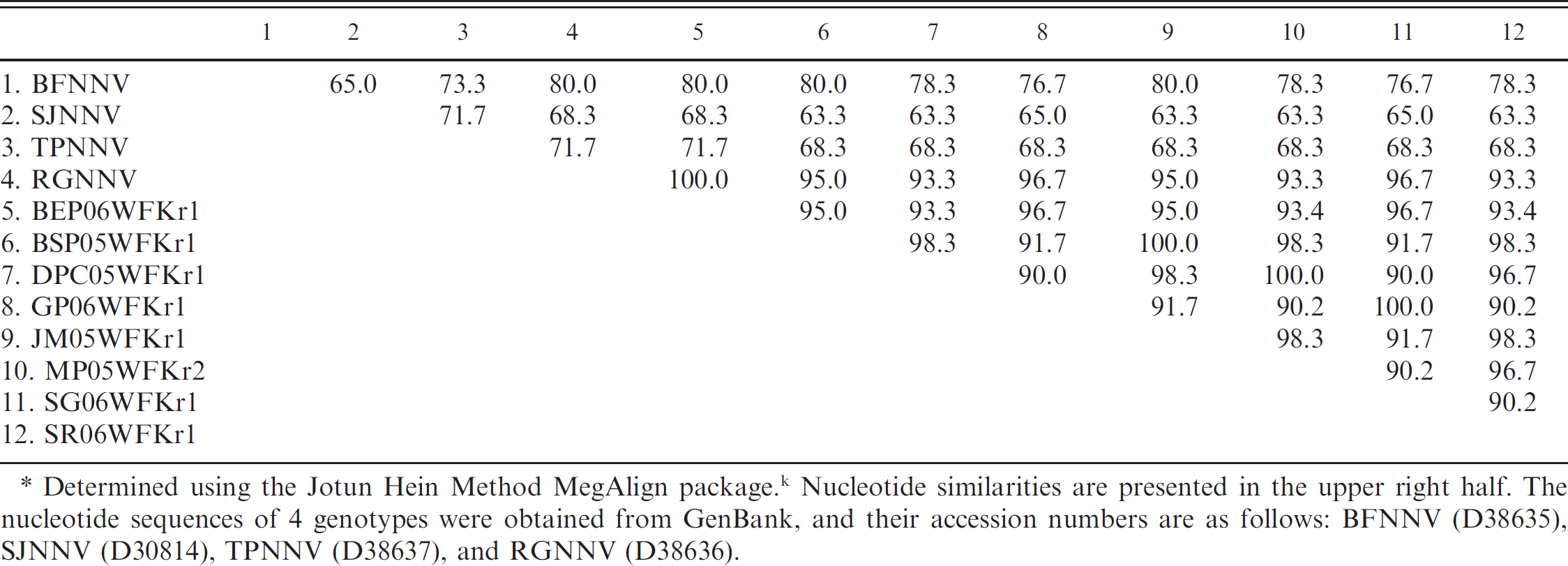
Determined using the Jotun Hein Method MegAlign packaged. k Nucleotide similarities are presented in the upper right half. The nucleotide sequences of 4 genotypes were obtained from GenBank, and their accession numbers are as follows: BFNNV (D38635). SJNNV (D30814), TPNNV (D38637), and RGNNV (D38636).

Alignment of representative amino acid sequences of RNA2 coat protein gene (aa 224–280) of betanodavirus. Outlined boxes (□) indicate changes in the amino acid position.
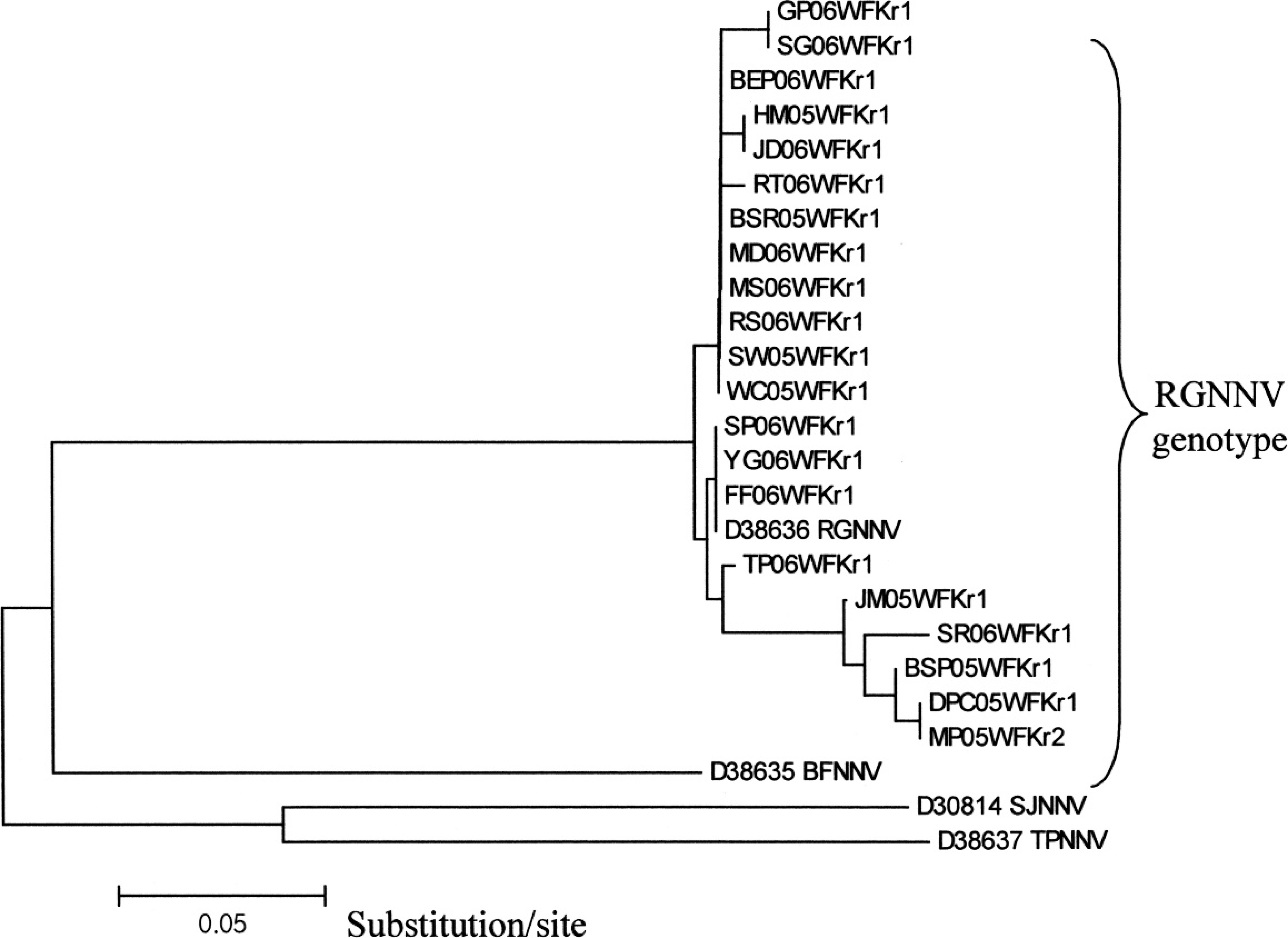
Phylogenetic tree based on the 4 genotypes and 21 Korean strain nucleotide sequences (nt 666–842) of RNA2 coat protein gene of betanodavirus. This tree was constructed by the neighbor-joining method (multiple alignment by the Jotun Hein Method in the MegAlign package k ).
Discussion
The first description of VNN was reported in cage-cultured groupers (Epinephelus septemfasciatus) on the southern coast of Korea. 38 However, in this case, the causative agent was not identified. There were also incidences of nodavirus disease outbreaks from 1999 to 2000 in 4 hatcheries culturing red drum (Sciaenops ocellatus) larvae located in the Yochon and Yosu areas of the southern coast of Korea. 34 The VNN infection might be attributable to either vertical or horizontal transmission. It was reported that mortality caused by VNN during the seed production of red drum was vertically transmitted from infected spawners. 17 However, the authors have suspected that VNN infection in Korea was caused by horizontal transmission via carrier or subclinically infected wild marine fish.
A diagnostic assay capable of detecting betanodaviruses directly in carrier fish samples would be of significant benefit to fish health management. However, isolation of betanodavirus from asymptomatic population of adult fish, in which the presence of low levels of infectious virus is usual, is more difficult than isolation from diseased fish. The usual method of nodavirus detection is culture of the virus from supernatants of brain or eye homogenates in tissue culture cells, 32 but there is evidence that this may not detect many carriers that tested positive by PCR methods. It was reported that infectious pancreatic necrosis virus may also be undetectable from kidney homogenates of carrier fish using cell cultures that are positive by RT-PCR. 39
Low levels of genetic material can be amplified by RT-PCR or nPCR assay, which is now used in a variety of viral epizootics. Although the authors did not detect samples by RT-PCR in the present study, 48 positive samples consisting of 21 species were detected by nPCR. To eliminate the presence of PCR product contamination in the present study, a confirmatory method using virus isolation was still performed in all positive nPCR samples. From 48 samples, only 21 virus strains from 12 species offish were isolated using E-11 cell lines. This may be due to the presence of low virus titer in carrier fish and a low virus detection limit in E-11 cell lines. The detection of viral pathogens by cell culture is very complex, and not all groups of viruses can be isolated on regular cell lines. A delay in the appearance of the first CPE is usual when the viral concentration is low, and some blind passages are needed to increase the yield. For this reason, PCR provides very sensitive, specific, and rapid detection of viruses in a variety of environmental samples. 16 Nested PCR amplification is also reported to ensure the specificity of detection and to increase the amplification signal, providing the method with very high sensitivity, especially for environmental samples that contain low numbers of viruses. 35
Twenty-one nodavirus strains detected were sequenced and analyzed by phylogenetic tree. The partial sequences of the RNA2 coat protein gene of these 21 strains were identical with RGNNV type, with a homology ranging from 93% to 100%. The homology with the other 3 genotypes (SJNNV, TPNNV, and BFNNV) ranged from only 63% to 80%. It was also reported previously that VNN-affected moribund or diseased maricultured fish in Korea, such as red drum and oblong rockfish (Sebastes oblongus), were also identical with grouper (RGNNV type). It has a homology ranging from 96% to 98% (RGNNV type) and <85% homology with Japanese flounder (Paralichthys olivaceus; JFNNV) and striped jack (Pseudocaranx dentex; SJNNV). 18 In the present study, another confirmatory method was done on the 2 representative viruses isolated from E-11 cell lines. Viruses were processed for nucleotide sequencing, and the results were confirmed to be of the same strain compared with the virus strain sequenced directly from nPCR product (data not shown). Detection of betanodavirus from these asymptomatic fish may indicate a horizontal transmission via water. The authors speculated that following a disease outbreak, surviving fish might become virus carriers without showing symptoms. The presence of these carrier fish in the marine environment could lead to epizootics, suggesting that some of these carriers shed infectious particles in the water and that both cultivated and wild fish in the marine environment are at risk from virus-contaminated environments. This has implications for the disease security of existing and new marine aquaculture. Since natural infections of nodavirus in marine fish occur within a wide range of temperatures from cold to warm waters, it is also important to know their survivability or stability under different water conditions. Betanodaviruses are a major cause of viral nervous necrosis; their presence in the water is an important factor to be considered in the transmission process either directly or indirectly to all healthy fish, cultured or wild, young or adult. So far, information on their stability in water is not yet available, and no study has been conducted on the survival pattern of 4 representative betanodavirus genotypes under different water conditions.
Environmental factors should also be considered as one of the sources of nodavirus transmission to marine fishes. Many environmental factors have been reported to influence virus survival in the marine environment. These include temperature, soluble chemicals such as salts and heavy metals, 36 sunlight, 3 the presence of microoorganisms 10 or their enzymes, 7 and some other heat labile factor. 24 Factors that enhance virus survival in seawater have also been discovered. The presence of sewage pollution has been claimed to both prolong virus survival 25 and enhance inactivation, 3 and the presence of nutrients 23 can lead to greater viral survival. Previous investigators have also demonstrated protection of viruses from inactivation by adsorption to solids. In the results of an experiment, IHNV adsorbed to naturally occurring substances in various aquatic environments, and it may provide a source of infection for susceptible fish inhabiting these environments. 43 It has been shown that enteric viruses will readily adsorb to sediment 19 and can survive longer in the estuarine environment when associated with sediment. 20,37
Carnivorous fish may also play an important role by feeding on nodavirus-contaminated fish or other biological organisms such as crustaceans, shellfish, aquatic plants, or other aquatic organisms present in the marine environment. In Japan, high frequencies (67%) of betanodaviruses were detected from apparently healthy cultured and wild marine fish. 13 All the nucleotide sequences of the fish species from Japan were highly homologous to each other (99–100%) and were closely related to RGNNV. It was also reported that viruses in the estuarine environment could be accumulated in filter-feeding shellfish or transported to other marine environments. 21 Therefore, it is important to detect carriers to control the spread of nodavirus.
In addition, one of the most noted methods of spreading nodavirus was by movement of infected fish. As speculated previously, 6,8 apparently uninfected pelagic and highly migratory fish such as Japanese jack mackerel may be an effective vector to transmit betanodaviruses to geographically remote areas such as Korea, considering the closer distance between the 2 countries. In this present study, species of Japanese jack mackerel that tested positive for nodavirus were 96% homologous with the RGNNV type. This might explain the distribution of betanodaviruses among wild and cultured fish populations. Furthermore, the present study's results illustrate that subclinically infected wild marine fish horizontally transmitted genetically related betanodaviruses and thus may be an inoculum source for susceptible fish species cultured in Korea.
To understand the epidemiology of betanodavirus affecting wild marine fish, further studies are needed on its survivability in various water conditions, the impact of other environmental factors, and the presence of biological organisms in the marine environment that may provide a source of infection to susceptible fish.
Acknowledgements
A Korea Research Foundation grant (KRF-2006-005-J02903) supported this study. The authors are grateful for a graduate fellowship provided by the Ministry of Education through the Brain Korea 21 program for Veterinary Science, Seoul National University. The authors are also thankful to Dr. T. Nakai, Hiroshima University. Japan, for kindly providing the E-11 cell line for virus isolation.
Footnotes
a.
Invitrogen Corporation, Carlsbad, CA.
b.
TAKARA Bio Inc., Seta 3-4-1, Otsu, Shiga, Japan.
c.
Biometra T-personal 48, Biometra GmbH Goettingen. Rudolf-Wissell, Germany.
d.
Cambrex Bio Science Rockland Inc., Rockland, ME.
e.
JRH Biosciences Inc., Lenexa, KS.
f.
Becton Dickinson and Company, Franklin Lakes, NJ.
g.
Sartorius AG 37070, Goettingen, Germany
h.
WelGENE Inc., Daegu, South Korea.
i.
Genome Sequencer (GS) 20 system sequencing kit, Roche Applied Science, Mannheim, Germany.
j.
Automatic Sequencer 3730xl DNA analyzer, Roche Applied Science, Mannheim, Germany.
k.
Windows version 3.12e, DNASTAR, Madison, WI.
