Abstract
The collection of biological material (e.g., blood) directly onto filter paper for subsequent use in laboratory assays such as polymerase chain reaction (PCR), has become a common practice. Dried cells or fluid on the paper can be readily rehydrated and retrieved into a standard volume of an appropriate elution buffer but introduces a dilution factor to the sample. The use of a common cutting instrument for excising a standard-sized piece of paper that contains the material also introduces the potential for transferring biological material from one sample to subsequent samples, causing false-positive results by PCR. In the present study, filter-paper-collected blood that contained beak and feather disease virus was used to determine if viral DNA could be transferred between samples by a hole punch used to excise sequential filter papers. It was determined that false-positive results could be obtained at least 13 times after a positive sample. Subsequently, the efficacy of 4 methods of hole punch disinfection, flaming, VirkonS, bleach, and a bleach-ethanol combination, was assessed. The only effective and practical method to destroy DNA was a method where the hole punch was agitated in commercial bleach, rinsed in water, the water was displaced with 100% ethanol and air-dried. This method was simple, cheap, and relatively rapid, and allowed for the use of a single hole punch for a series of samples, without carryover contamination and consequent false-positive results.
Keywords
Molecular diagnostic methods (e.g., polymerase chain reaction [PCR]) are widely used for the detection of etiological agents, including viruses, bacteria, and parasites, as well as for genotyping of both fauna and flora and in forensic investigations. PCR allows the detection of DNA from samples that originally contain a very few copies of the DNA of interest. 1 However, the fidelity of the PCR reaction relies on avoiding contamination of true-negative samples by carryover of target material from other samples.
Recently, it has become common practice to apply biological material to filter paper a,b for subsequent use in PCR. The material dried on filter paper is easily stored and transported, without the need for refrigeration. 3,6,11,13,15,16 Typically, such samples are eluted from the sample paper, and nucleic acid is subsequently isolated from the eluate for use in PCR. Furthermore, some filter-paper products include reagents that lyse cell membranes and stabilize nucleic acids, allowing material to be easily washed and deposited directly into the PCR reaction. 2,13 To access the biological material for PCR, the material dried on filter paper is commonly excised by using a hole punch. 3,4,6,9,12,13,16,17 This method allows for expedient processing of samples and provides a standard volume of filter paper. However, the use of a single piece of equipment for multiple samples makes the method vulnerable to carryover of nucleic acid between samples.
The aim of this experiment was to investigate the potential for contamination of true-negative samples by carryover of DNA caused by using a hole punch to excise material from filter paper. Subsequently, a method to minimize the potential of DNA transfer from a positive sample to the other samples was developed.
Ethylenediamine tetra-acetic acid (EDTA)-preserved blood originating from a long-billed corella (Cacatua tenurostris) previously diagnosed beak and feather disease virus (BFDV) positive by clinical examination and PCR by the Murdoch University veterinary diagnostic service, was spotted onto filter paper a and dried at room temperature overnight.
To determine whether the hole punch could cause carryover contamination, a new, unused hole punch c was used to excise filter paper. a In accordance with the DNA extraction kit d specifications, 3 circles were excised from each filter paper. Sequentially, “material” was excised from an unused filter paper (precut control) from a previously unopened pack, from filter paper with blood from the infected corella sample (known positive sample), and from 5 unused filter papers (contamination controls 1–5). The sequence of filter paper excisions is illustrated in Fig. 1. The excised circles were placed into 1.5-ml microcentrifuge tubes, and DNA was extracted according to the kit manufacturer's instructions.
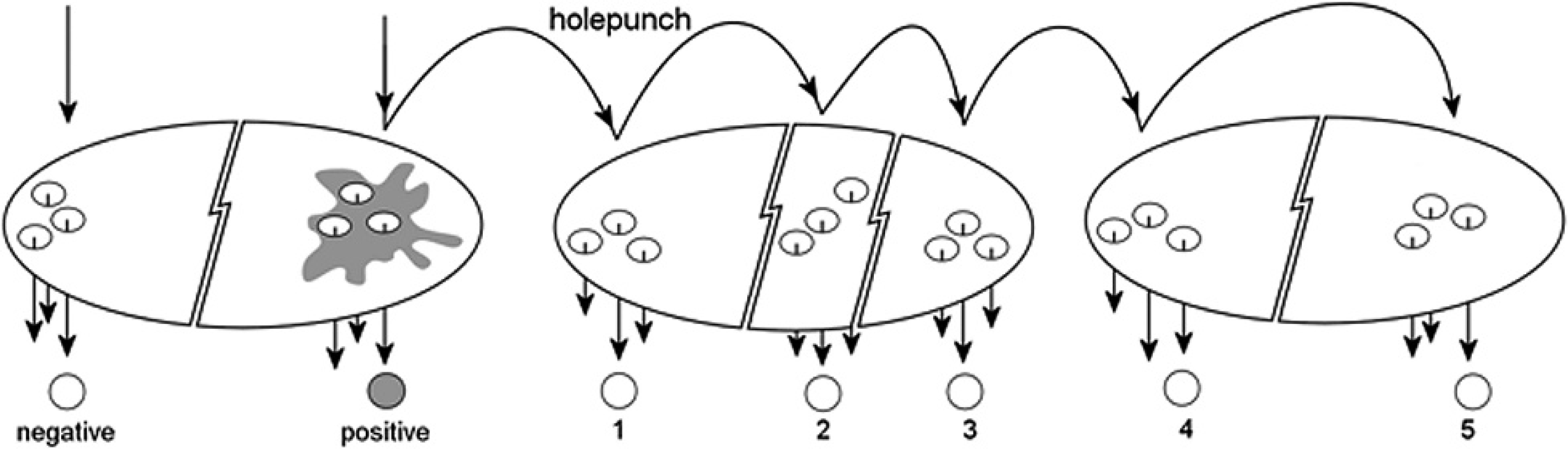
Method used to assess carryover of biological material by a hole punch used to excise a standard sized hole from filter paper.
Four methods of disinfection were assessed to determine an efficient and practical method of avoiding carryover contamination between samples in a test panel. These were 1) flaming the hole punch with a Bunsen burner; 2) soaking the hole punch in 0.5% (w/v) VirkonS e for 30 min, with subsequent washing in running tap water for 1 hr; 3) soaking the hole punch in bleach (42 g/liter sodium hypochlorite f ) for 20 min, with subsequent washing in running tap water for 20 min (after the washing step in methods 2 and 3, the hole punch was dried by removing excess water with tissue paper and drying at 50°C in a heated cabinet); and 4) agitating the hole punch in 100% bleach f (approximately 5 sec), rinsing in water (approximately 5 sec), immersing and agitating in 100% ethanol (5–10 sec) to displace the water, and finally air-drying.
The dryness of the hole punch after each method of liquid disinfection was assessed by flicking the instrument against a clean dry surface and observing for drops. In methods 2 and 3, the hole punch was dry after approximately 60 min at 50°C. In method 4 the ethanol evaporated, leaving the hole punch dry within seconds.
For each of the disinfection methods, a new, unused hole punch was used to excise filter paper. In accordance with the instructions in the DNA extraction kit, 3 circles were excised from each filter paper. Sequentially, “material” was excised from an unused filter paper (precut control) from a previously unopened pack, from filter paper with blood from the infected corella sample (known positive sample), then from an unused filter paper (carryover positive control), to positively control carryover of a nondisinfected hole punch. Further, the known positive sample was excised (without retaining circles) to contaminate the hole punch before disinfection. After disinfection, unused filter paper was excised (disinfection control). Finally, to control for significant disinfectant residue retention (and associated interference in subsequent applications), a single circle was excised from the known positive sample (postdisinfection positive control). The sequence of sample paper excisions is illustrated in Fig. 2. The excised circles were placed into 1.5-ml microcentrifuge tubes, and the DNA (if any) was extracted according to the kit manufacturer's instructions.
To detect DNA that might be present in the samples prepared above, PCR for detecting a fragment of BFDV open reading frame V1 was conducted as previously described 21 by using forward primer P2 (5′-AACCCTACAGACGGCGAG-3′) and reverse primer P4 (5′-GTCACAGTCCTCCTTGTACC-3′). Reactions consisted of 2 μl of 25 mM MgCl2, g 5 μl of 5× polymerase buffer, g 12.8 pmol of each primer, 0.12 μl of Tth polymerase, g 2 μl of DNA extract, and water-for-injection h added to a final volume of 25.12 μl. Reactions were subjected to the following conditions: 95°C for 5 min, then 32 cycles of 95°C for 30 sec, 60°C for 30 sec, 72°C for 1.5 min, and finally 72°C for 10 min. PCR products were electrophoresed in a 1% agarose gel at 90 V for 30 min, and the results were visualized by ultraviolet transillumination.
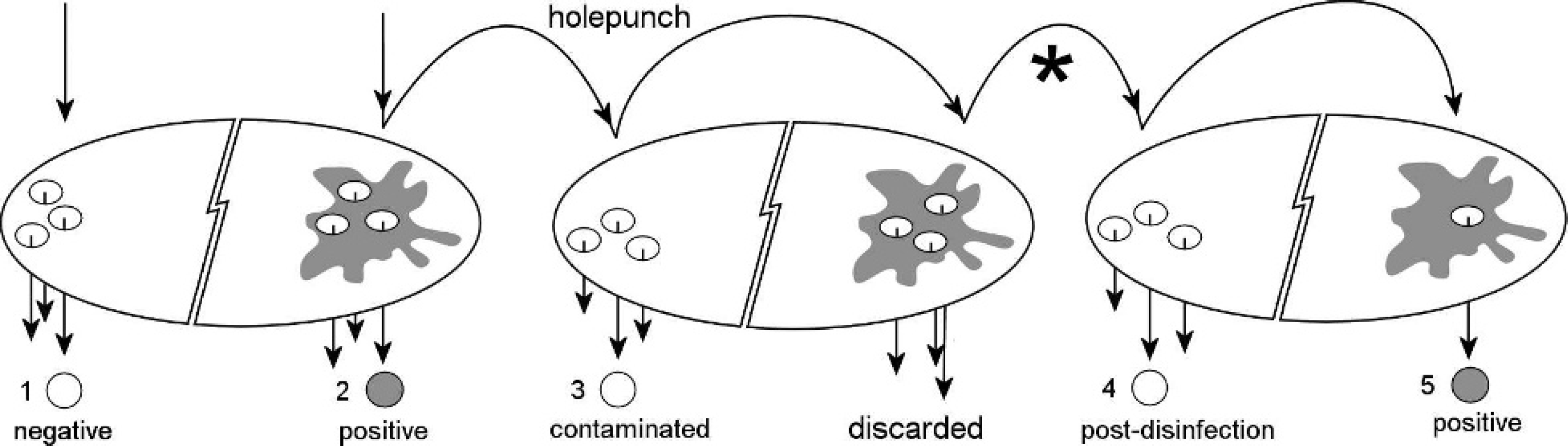
Method used to assess disinfection procedures of a hole punch. For assessing methods of decontamination, the points where the hole punch was disinfected with method 1 (flaming), method 2 (VirkonS), method 3 (bleach), or method 4 (rapid bleach and ethanol combination) are shown as an asterisk. A postdisinfection positive sample was used to assess the effects of any carryover of disinfectant that might have interfered with the assay (causing a false-negative result).
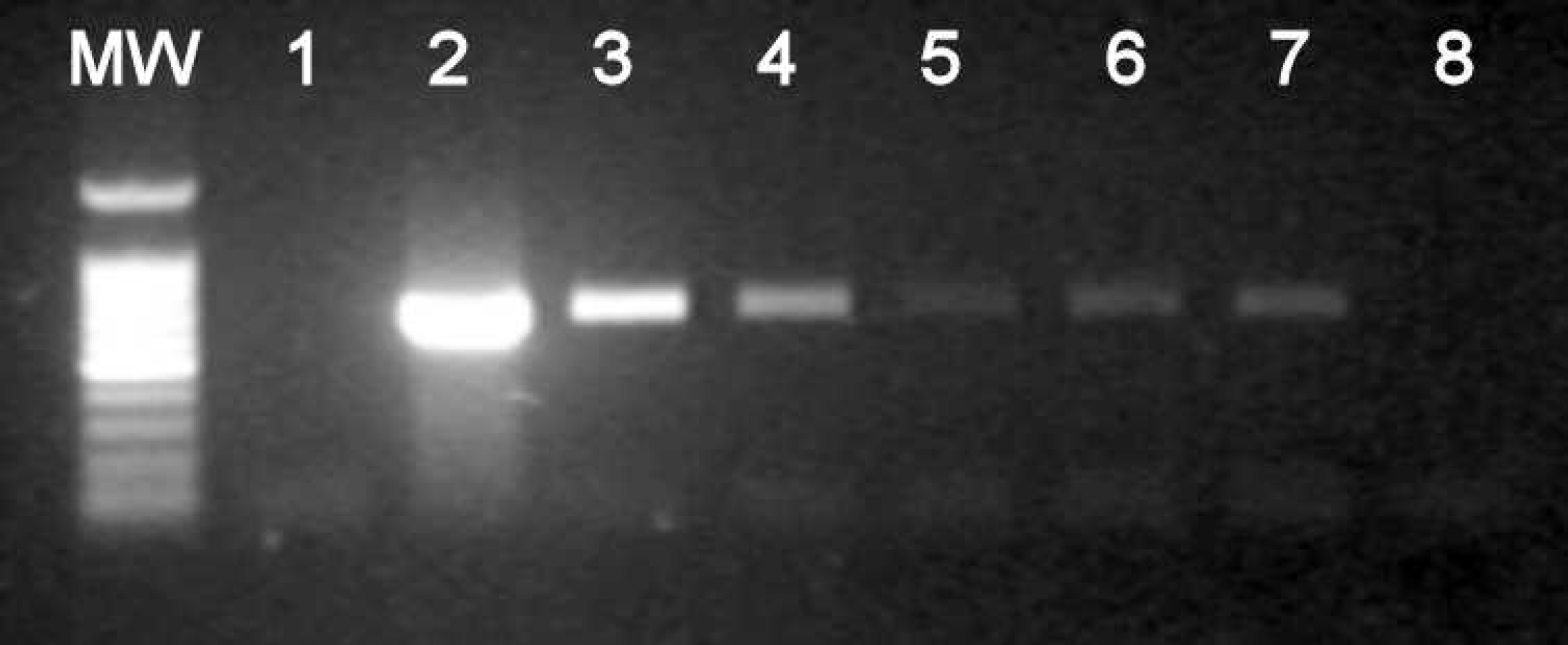
Agarose gel, illustrating the effect of using a non-disinfected hole punch to excise filter paper for DNA extraction and subsequent polymerase chain reaction (PCR) analysis. Lane MW: Promega 100-bp molecular weight marker. The sample sources for lanes 1–7 are shown diagrammatically in Fig. 1. Lane 1: Precut control. Lane 2: Known positive sample. Lanes 3–7: Contamination controls 1–5. Lane 8: PCR negative control.
In the sample panel prepared by using a hole punch that was not disinfected between excision of the various filter papers, the following results were obtained: PCR of precut control did not yield any detectable amplification (Fig. 3. lane 1). The known positive control (lane 2) yielded the expected strong signal. Contamination controls 1–5 (lanes 3–7) yielded progressively weaker signals at the same molecular weight as the positive control.
In the sample panels prepared by using a hole punch that was disinfected between excision of the various filter papers, the following results were obtained. In method 1 (Bunsen burner), PCR of the precut control did not yield any detectable amplification. The known positive sample yielded a strong product at the expected molecular weight. The carryover positive control yielded a weaker product at the expected molecular weight for this fragment. PCR of the disinfection control sample yielded a product at the molecular weight of the known positive sample but was significantly weaker than the known positive. Finally, the postdisinfection positive control yielded the expected product (data not shown). In method 2 (VirkonS and tap water) and method 3 (bleach and tap water), the precut control did not yield any detectable amplification from PCR. The expected positive result was obtained with the known positive sample. PCR of the carryover positive control resulted in a positive signal, weaker than that of the known positive. The disinfection control did not yield detectable amplification. Finally, the postdisinfection positive control yielded a strong signal at the same molecular weight of the known positive sample. Methods 2 and 3 both caused corrosion of the hole punch (data not shown). In method 4 (bleach, tap water, ethanol), no detectable amplification was obtained with PCR of the precut control (Fig. 4, lane 1). The known positive sample yielded the expected positive result (lane 2). PCR of the carryover positive control obtained a product at the molecular weight of the known positive sample, however, significantly weaker than the known positive (lane 3). The disinfection control sample did not yield a detectable signal (lane 4). Finally, the postdisinfection positive control resulted in a strong positive signal (lane 5). This method did not cause visible corrosion of the hole punch.
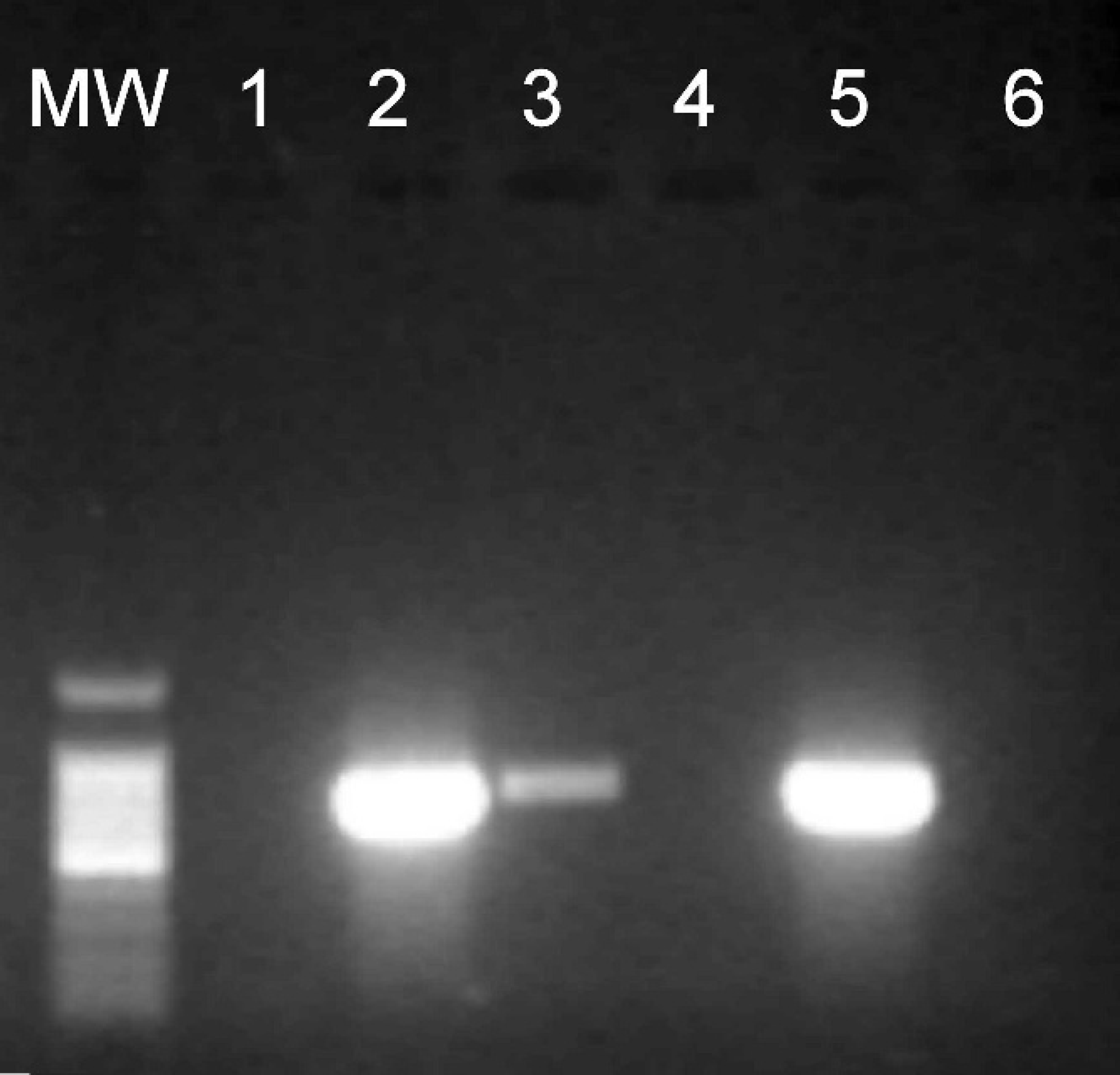
Agarose gel of polymerase chain reaction (PCR) products yielded from filter paper cut with a disinfected hole punch. Lane MW: Promega 100-bp molecular weight marker. The sample sources for lanes 1–5 are shown diagrammatically in Fig. 2. Lane 1: Precut control. Lane 2: Known positive sample. Lane 3: Carryover positive control. Lane 4: Postdisinfection result (method 4). Lane 5: Postdisinfection positive control. Lane 6: PCR negative control.
Many studies used hole punches to excise material dried on filter paper 2–4,7,9,12,14,16,18,20 . however; only a few reports mention any protocols for detecting possible carryover or detection of false positives in filter-paper-based PCR methods. 9,11 Furthermore, many published studies overlooked any consideration for the possibility of such contamination. 2,3,7,12,14,18
Quality assurance methods to avoid such contamination were reported by some investigators. 4,9,13,16,17,20 Some laboratories report punching holes in blank paper between samples to remove possible contaminants. 17 One study reported ethanol flaming the hole punch before making an excision, 16 and other studies reported wiping or pressing the hole punch against ethanol-wetted paper towel, 4,13 or a combination of wiping the hole punch with ethanol and excising blank papers. 9
The present study shows that there may be significant carryover at least 13 excisions after the first excision of a positive sample. It was concluded that sequential cutting of fresh blank paper alone is inadequate for cleaning or removing contaminating material from the hole punch. Furthermore, the data obtained in the present experiment show that exposing a contaminated hole punch to a high-temperature Bunsen flame may not result in elimination of all residual DNA.
Studies on the effects of ethanol for disinfection showed that, although various concentrations of ethanol may be, in some cases, effective at reducing infectivity, it does not destroy nucleic acid effectively. 5,8 Thus, it can be concluded that a hole-punch disinfection procedure that depends on alcohol will not be sufficient to avoid nucleic-acid carryover contamination.
The likely possibility of a hole punch carrying contamination to consecutive samples and associated false-positive results may have been overlooked by many laboratories, and some may be using methods that are insufficient to ensure decontamination. In the present study, a virgin hole punch was used to determine whether hardware used for processing dried blood samples on filter paper could serve as a vessel for carryover. The results obtained in this experiment give support for this hypothesis, because clean filter paper processed subsequently to the known positive control resulted in progressively weaker positive results for as many as 13 punches afterward. This result is not surprising in light of the ability of the PCR to detect few copies of the target DNA.
The present study demonstrated that the simple, cheap, and relatively rapid disinfection technique involving bleach, tap water, and ethanol was the most successful. This method allowed for the use of a single hole punch for a series of samples, without carryover and consequent false-positive results.
Forensic scientists and scientists who study ancient DNA commonly use bleach to eliminate contaminating DNA on bone samples. 10 Furthermore, studies have determined the effectiveness of bleach for eliminating DNA in PCR reactions, 19 but the present study is the first to demonstrate its use to decontaminate hole punches used for filter-paper excision.
The results presented in the present study provide evidence for the potential of a hole punch to transfer material between filter-paper dried samples. The possibility of carryover of material between samples when using a piece of hardware should be considered by all researchers who use PCR analysis of biological material and appropriate methods used to minimize its occurrence.
Footnotes
a.
Whatman no. 3 filter paper, Whatman International Ltd. Maidstone, England.
b.
FTA filter paper, Whatman International Ltd, Maidstone. England.
c.
One hole punch, Officeworks Superstores Pty Ltd, Box Hill. Vic, Australia.
d.
QIAamp DNA blood mini kit, Qiagen Pty Ltd, Doncaster, Vic, Australia.
e.
ANTEC VirkonS, Janos-Hoey Pty Ltd, Forbes, NSW, Australia.
f.
White King, SaraLee Household and Body Care, Pymble. NSW, Australia.
g.
PCR reagents, Fisher Biotech, Wembley, WA, Australia.
h.
Water-for-injection BP, AstraZeneca Pty Ltd, North Ryde. NSW, Australia.
