Abstract
Aves polyomavirus 1, psittacine beak and feather disease virus, and psittacid herpesvirus 1 are important pathogens of psittacine birds with the potential to cause substantial morbidity and mortality. Using publically available nucleotide sequences, we developed and validated a triplex real-time PCR (rtPCR) assay to rapidly detect these 3 viruses. The assay had high analytical sensitivity, detecting <6 copies of viral DNA per reaction, and 100% analytical specificity, showing no cross-reactivity with 59 other animal pathogens. Archived formalin-fixed, paraffin-embedded tissues from psittacine birds diagnosed at postmortem as infected with each of the viruses as well as virus-negative birds were used to validate the utility of the assay. Birds were selected for the positive cohort if they showed histologic evidence of infection (i.e., characteristic inclusion bodies in tissues); birds in the negative cohort had final diagnoses unrelated to the pathogens of interest. The triplex rtPCR assay confirmed 98% of histopathology-positive cases, and also identified subclinical infections that were not observed by histologic examination, including coinfections. Birds that tested positive only by rtPCR had significantly higher cycle threshold values compared to those with histologic evidence of infection. Positive, negative, and overall percentage agreements as well as the kappa statistic between the results of the assay and histopathology were high, demonstrating the usefulness of the assay as a tool to confirm disease diagnoses, and to improve detection of subclinical infections.
Introduction
Psittacine birds (order Psittaciformes) include many species that are commonly kept as pets in North America and other countries. Given that these species are popular domestic companions, efficient laboratory methods are necessary to prevent and manage diseases. Rapid diagnoses are especially important for infectious diseases to reduce bird-to-bird transmission. Three DNA viruses are known to cause serious morbidity and mortality in psittacine birds: aves polyomavirus 1 (APV1; agent of budgerigar fledgling disease); beak and feather disease virus (BFDV; agent of psittacine beak-and-feather disease); and psittacid herpesvirus 1 (PsHV1; agent of Pacheco’s disease and internal papillomatosis).8,9
APV1 (family Polyomaviridae, genus Gammapolyomavirus) 18 causes severe disease in young psittacine birds, characterized by systemic virus dissemination, necrosis, and the presence of large intranuclear inclusion bodies in multiple parenchymatous tissues. 14 BFDV (family Circoviridae, genus Circovirus) 2 replicates in lymphocytes and macrophages, causing necrosis and atrophy of lymphoid organs, leading to severe immunosuppression and often results in coinfection with other viruses or pathogens.23,26 Histologically, the virus produces intracytoplasmic and intranuclear botryoid, basophilic inclusions, mainly in the thymus and cloacal bursa. 26 Chronic infections can produce feather dystrophy as a result of replication of the virus in feather follicles. 20 PsHV1 (family Herpesviridae, genus Iltovirus, species Psittacid alphaherpesvirus 1) 17 can become latent with reactivation at times of stress, causing tissue necrosis and high mortality. 8 Histologically, the virus produces eosinophilic, intranuclear inclusions bodies (Cowdry type A), primarily in the liver. 11
Although multiplex PCR tests are increasingly utilized in veterinary laboratories, to date the only multiplex assay that has been reported in the literature for psittacine birds is a duplex conventional PCR assay for APV1 and BFDV. 19 Real-time PCR (rtPCR) assays using SYBR green or SYTO9 (intercalating fluorescent dyes) have been independently developed for BFDV, APV1, and PsHV1,13,24 and validations of TaqMan-based rtPCR assays have also been published for BFDV in 2 reports.3,10 Psittacine birds can be infected with multiple viruses simultaneously, as documented in numerous reports of coinfection with APV1, BFDV, and/or PsHV1.6,15,16,25 A multiplex rtPCR assay that can test for multiple pathogens in a single reaction would reduce the cost of processing, providing additional data on presence or absence of pathogens that would otherwise not be assessed with individual tests.
Our objectives were: 1) to develop a triplex rtPCR assay to detect APV1, BFDV, and PsHV1 and validate it using formalin-fixed, paraffin embedded (FFPE) tissues from captive psittacine birds; 2) to assess the agreement between histopathology and the rtPCR assay; 3) to determine the extent of subclinical infection and coinfection with these pathogens using postmortem samples; and 4) to assess which tissues are most appropriate for pathogen detection.
Materials and methods
Assay development and protocols
Viral sequences available in GenBank (www.ncbi.nlm.nih.gov/genbank) as of March 28, 2018 were used to design primers and probes against conserved regions of the genomes from the 3 test viruses and the enterovirus control (Table 1). Genome regions targeted were conserved regions in the origin of replication (ORI), the replicase gene (rep), and the UL16 gene (tegument protein) for APV1, BFDV, and PsHV1, respectively. Primer specificity was tested using a panel of 59 veterinary agents (Supplementary Table 3) for which testing is routinely conducted at the Animal Health Laboratory (AHL; University of Guelph, Ontario, Canada), which is an American Association of Veterinary Laboratory Diagnosticians–accredited diagnostic facility. Analytical sensitivity of each reaction was assessed separately for each virus using 10-fold dilutions of synthetic double-stranded target gene fragments (gBlocks; Integrated DNA Technologies, Coralville, IA).
Primers and probes, genes targeted, and concentrations used in the reactions of a multiplex rtPCR assay developed for the detection of APV1, BFDV, and PsHV1 in psittacine birds.
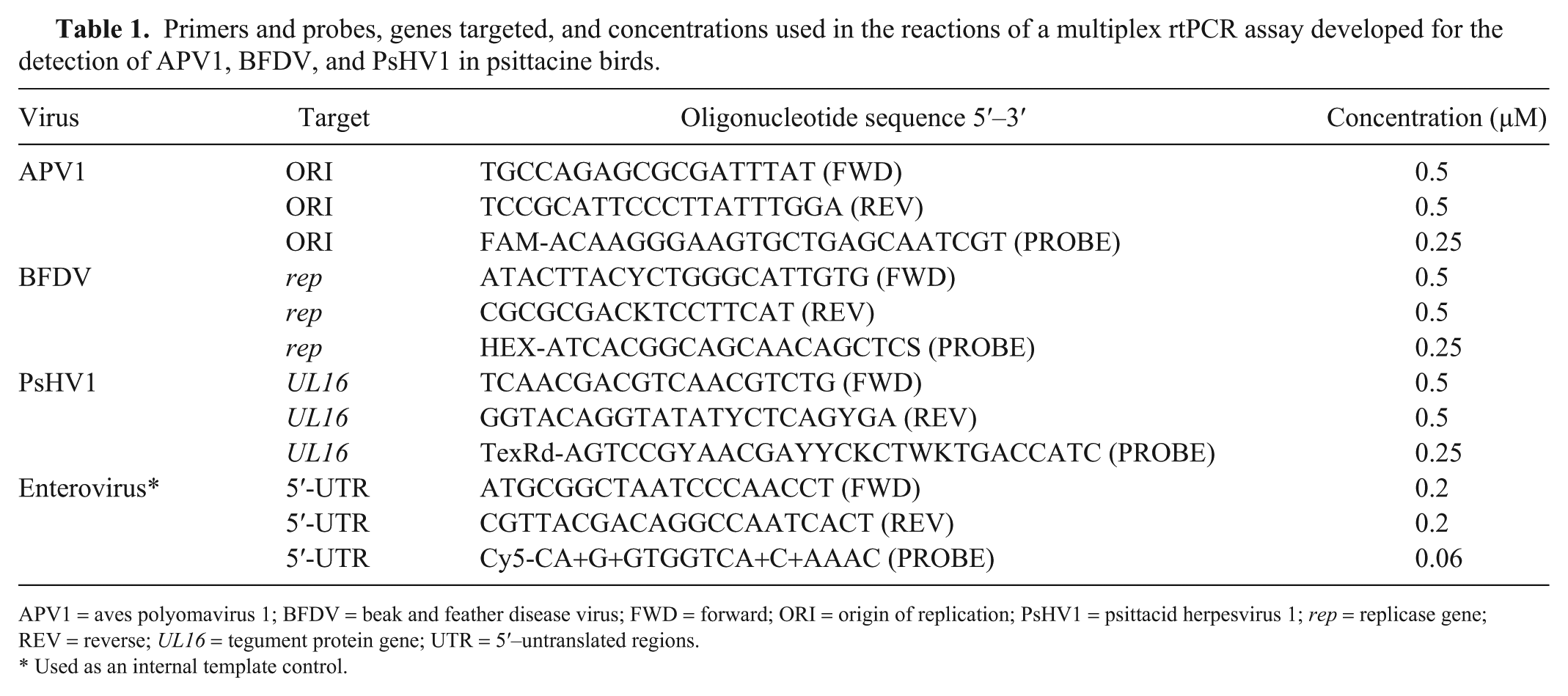

APV1 = aves polyomavirus 1; BFDV = beak and feather disease virus; FWD = forward; ORI = origin of replication; PsHV1 = psittacid herpesvirus 1; rep = replicase gene; REV = reverse; UL16 = tegument protein gene; UTR = 5′–untranslated regions.
Used as an internal template control.
Selection and processing of diagnostic cohorts for test validation
To validate the utility of the triplex rtPCR assay for viral detection in real tissues, FFPE tissues from psittacine birds, submitted to the University of Guelph for postmortem examination, were used to compile positive and negative diagnostic cohorts. Positive birds (i.e., histopathology-positive) were selected based on the final diagnosis in the pathology report, as well as histologic identification of viral inclusion bodies (Supplementary Figs. 1–3) in at least one of the following tissues: liver, kidney, spleen, or cloacal bursa. Negative birds (i.e., histopathology-negative) were selected from birds that did not show evidence of viral infection (i.e., absence of extensive necrosis and viral inclusions). For all birds, demographic data and cause of death were recorded. Age was delimited into 4 categories: chick, juvenile, adult, and senior, which were defined by cutoffs of 3% of average lifespan,7,29 age at sexual maturity, 30 and overall average lifespan for each genus. 30 Whenever possible, birds in the negative cohort were selected to be conspecific or congeneric and age-matched with those in the positive cohort. Based on the inclusion criteria above, we retrieved 40 blocks from 21 birds diagnosed with APV1, 43 blocks from 17 birds with BFDV, and 30 blocks from 16 birds with PsHV1. These included 2 birds diagnosed by histopathology with coinfections, 1 with APV1 and 1 with PsHV1, each with concurrent BFDV infections. The cohort of histopathology-negative cases included 31 birds, resulting in 184 blocks tested with the assay.
Using a standard microtome set at 20 µm, scrolls (shavings of wax and tissue) were taken from paraffin blocks that contained liver, kidney, and spleen for each bird. Cloacal bursa was included for histopathology-positive BFDV cases, and 9 histopathology-negative birds. Therefore, each bird could have a maximum of 4 scrolls (in the case that each tissue was in a separate block). New microtome blades were used for scroll sampling of each paraffin block, to prevent cross-contamination. Other tissues (besides liver, kidney, spleen, and cloacal bursa) were inconsistently included in blocks and were not included in downstream analysis with the exception of thymus, which was considered lymphoid tissue along with cloacal bursa.
Nucleic acid extraction and testing
Real-time PCR and extraction of total nucleic acids from FFPE tissues were conducted at the AHL. Tissue scrolls from paraffin blocks were transferred to a safe-lock, 1.5-mL tube, and 200 μL of the ATL buffer (catalog 939011; Qiagen, Hilden, Germany) was added. Twenty μL of proteinase K (>600 mAU/mL, catalog 19131; Qiagen) and 160 μL of deparaffinization solution (catalog 19093; Qiagen) was then added and mixed by vortexing. Tubes were incubated in a shaking heat block, first at 60°C for 45 min, 300 rpm, and then at 80°C for 30 min, 300 rpm. Once samples cooled down to room temperature, 50 μL of the bottom phase was collected and used for extraction (MagMax viral RNA isolation kit, catalog AM1836; Thermo Fisher Scientific, Waltham, MA) in a magnetic particle processor (MagMAX Express-96; Thermo Fisher). Armored RNA enterovirus-like particles (catalog 42050; Asuragen, Austin, TX) were added to the extraction kit lysis solution as an internal control. Eight µL of total nucleic acid was tested by rtPCR in 25-µL reactions (Light Cycler 480; Roche, Basel, Switzerland), using AgPath-ID one-Step RT-PCR kit (Thermo Fisher) under conditions described in Table 2. Samples with cycle threshold (Ct) values <37 were considered rtPCR-positive, and ≥37 were considered inconclusive. Samples that returned no Ct value were considered negative.
Cycling parameters used in a multiplex rtPCR assay for the detection of aves polyomavirus 1, beak and feather disease virus, and psittacid herpesvirus 1.
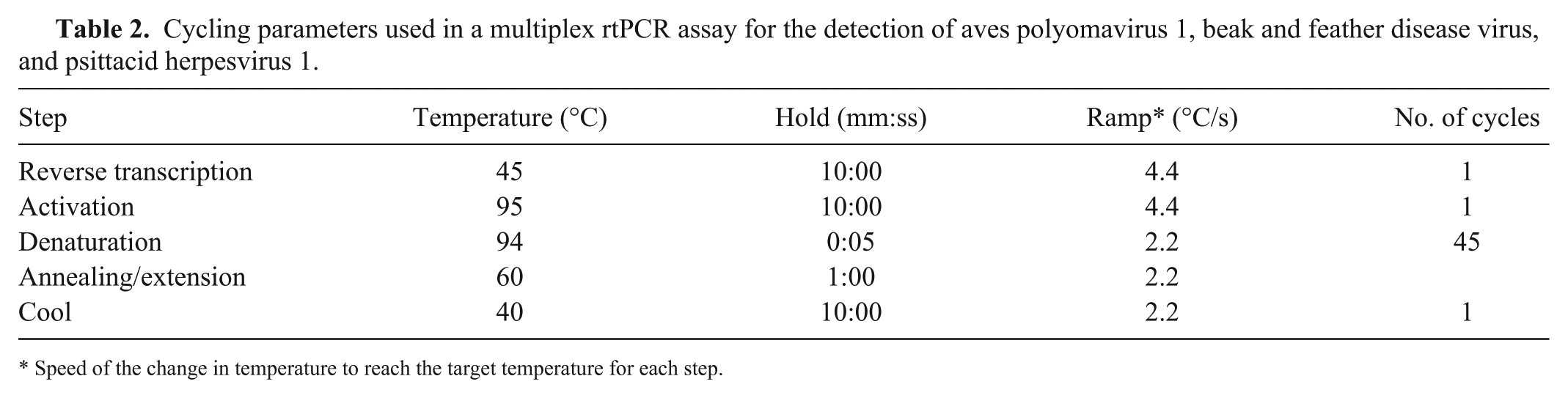
Speed of the change in temperature to reach the target temperature for each step.
Analyses
For each bird, the scroll with the lowest Ct value for each virus was used to define the infection status, resulting in a maximum of one Ct value per virus per bird. The numbers of histopathology-positive cases that were also identified by rtPCR for each virus were tallied. No true gold standard test could be defined in this situation, given that disease or infection status could not be absolutely confirmed in all diagnostic cases, and therefore terminology such as sensitivity and specificity were avoided; percentage agreements were calculated in lieu of these. Percentage agreements between both test methods were calculated using rtPCR as the gold standard to represent infection. 4 Specifically, positive percentage agreement (PPA) is the total number of histopathology-positive birds of the total rtPCR-positive birds, and similarly, negative percentage agreement (NPA) is the number of histopathology-negative birds of the total rtPCR-negative birds. Overall percentage agreement (OPA) is the sum of PPA and NPA, which gives an overall measure of the performance of the test. Percentage agreements were also calculated using histopathology as the gold standard, and these numbers were used in overall characterization of agreement. Further, the Cohen kappa statistic was calculated for each virus to measure the statistical agreement between the 2 test methods. 5 Average Ct values from individual histopathology-positive birds and birds without histologic lesions for each virus were compared using 2-tailed t-tests for all rtPCR-positive cases. Given that many infections were clinically silent, the proportion of birds coinfected with multiple viruses was also calculated. Lastly, considering all birds that were positive by rtPCR for each virus, the Ct values yielded by each scroll containing liver, kidney, spleen, or cloacal bursa and/or thymus separately were recorded. This analysis was carried out to evaluate whether specific tissues in the paraffin block provided any diagnostic advantage.
Results
Analytical sensitivity and specificity
The triplex rtPCR assay detected <6 copies per reaction of synthetic target oligonucleotide for each of the 3 viruses evaluated (Ct range: 34.9–36.7). The triplex assay did not cross-react with any of the panel of 59 other agents, indicating 100% specificity of the primers/probe sets (data not shown).
Positive and negative diagnostic cohorts
Psittacine species sampled represented 16 genera (Supplementary Table 1), of all age classes, including chicks (34%), juveniles (28%), adults (13%), seniors (7%), and unspecified age (18%). All birds in the histopathology-positive cohort for APV1 and PsHV1 died of the associated viral disease except for a macaw (Ara sp.) chick diagnosed with APV1 infection, whose proximate cause of death was aspiration. Histopathology of birds diagnosed with APV1 and PsHV1 infection was characterized by multifocal necrosis in multiple organs and typical viral inclusion bodies (Supplementary Figs. 1, 2). For BFDV-positive birds, bacterial or fungal (n = 4) and psittacine bornavirus (n = 1) infections were the proximate cause of death, and BFDV for the remaining 12 birds. Lesions caused by BFDV consisted primarily of lymphoid depletion in cloacal bursa and thymus with typical botryoid inclusion bodies (Supplementary Fig. 3). Causes of death in the histopathology-negative cohort included: neoplasia (n = 10), inflammation with no identified cause (n = 4), bacterial or fungal infection (n = 4), aspiration or asphyxiation (n = 3), idiopathic hemorrhage (n = 2), and one case each of congenital disease, squamous metaplasia, egg binding, exocrine pancreatic insufficiency, myodegeneration, and trauma. The cause of death was undetermined in 2 cases.
Diagnostic agreement between the triplex assay and histopathology
The majority of cases (98%) in the histopathology-positive cohort were also positive by the triplex rtPCR assay. Inclusion bodies consistent with APV1 and PsHV1 were histologically detected separately in 2 birds that were rtPCR-negative for the suspected virus; however, both birds were rtPCR-positive for BFDV (Supplementary Table 1). Ct values in the histopathology-positive cohorts were on average 20.7, 15.3, and 24.7 for APV1, BFDV, and PsHV1, respectively. All Ct values in the negative cohort were >22, with an average of 33.6 (Supplementary Table 1). Twenty-four (65%) birds in the histopathology-negative cohort were confirmed as virus-negative or inconclusive by triplex rtPCR, and 7 (23%) birds tested positive for at least 1 virus.
Agreement between tests (Table 3) showed that histologic assessment was able to identify 71%, 41%, and 83% of birds infected with APV1, BFDV, and PsHV1 as determined by triplex rtPCR, respectively (PPA). NPAs were ≥98% for all 3 viruses, indicating that almost all cases that were negative by multiplex rtPCR were also negative by histopathology. The OPA indicated that 89%, 68%, and 95% of all results for APV1, BFDV, and PsHV1, respectively, were in agreement, as confirmed by the kappa statistics (Table 3). Percentage agreements using histopathology as a gold standard were also calculated (Supplementary Table 2), highlighting the ability of rtPCR assay to confirm infection in 98% of histopathology-positive cases.
Positive, negative, and overall percentage agreements between pathology diagnoses and real-time PCR (rtPCR) results defining rtPCR as the gold standard for identification of infection with APV1, BFDV, and PsHV1 in psittacine birds.
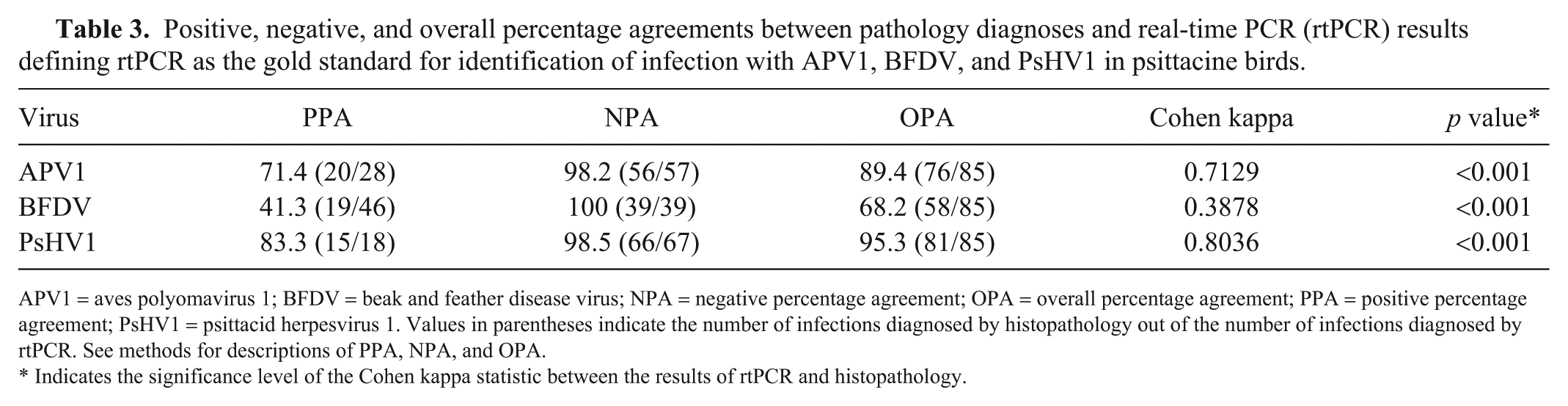
APV1 = aves polyomavirus 1; BFDV = beak and feather disease virus; NPA = negative percentage agreement; OPA = overall percentage agreement; PPA = positive percentage agreement; PsHV1 = psittacid herpesvirus 1. Values in parentheses indicate the number of infections diagnosed by histopathology out of the number of infections diagnosed by rtPCR. See methods for descriptions of PPA, NPA, and OPA.
Indicates the significance level of the Cohen kappa statistic between the results of rtPCR and histopathology.
Coinfections were common, especially between BFDV and the other 2 viruses (Fig. 1). Overall, 57%, 24%, and 69% of birds that were histopathology-positive for APV1, BFDV, or PsHV1, respectively, also tested positive for at least one other virus by rtPCR. Of the 2 birds originally diagnosed with coinfections by histopathology, the BFDV:APV1 coinfection was confirmed by rtPCR, but the BFDV:PsHV1 coinfection was only positive for PsHV1. In total, the multiplex rtPCR identified coinfections with 2 or more viruses in 29 birds (Fig. 1). Two of these 29 birds were positive by rtPCR for all 3 viruses; the histopathology diagnosis identified only APV1 infections in these 2 cases.
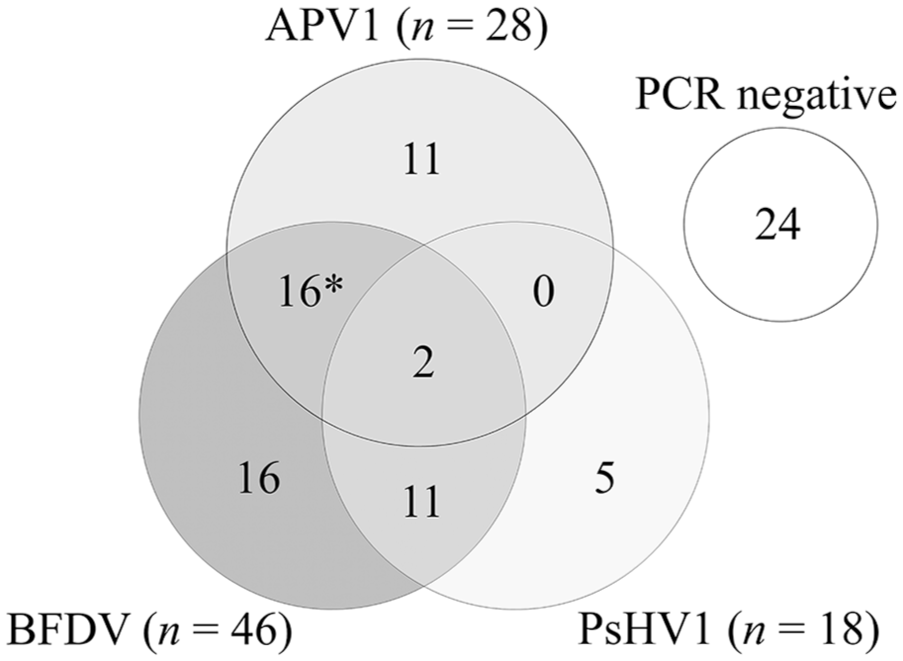
Non-proportional Venn diagram showing the numbers of birds infected with aves polyomavirus 1 (APV1), beak and feather disease virus (BFDV), and/or psittacid herpesvirus 1 (PsHV1) as determined by a multiplex real-time PCR in a cohort of 85 psittacine birds. Asterisk indicates one bird that was diagnosed with an APV1:BFDV coinfection by histopathology.
Given the presence of many silent infections detected by rtPCR, the Ct values of birds diagnosed by histopathology and rtPCR were compared with those identified by rtPCR alone. For each virus, the average Ct values were significantly higher for birds that did not have inclusion bodies evident microscopically (i.e., histopathology-negative) compared to those that did (Fig. 2).
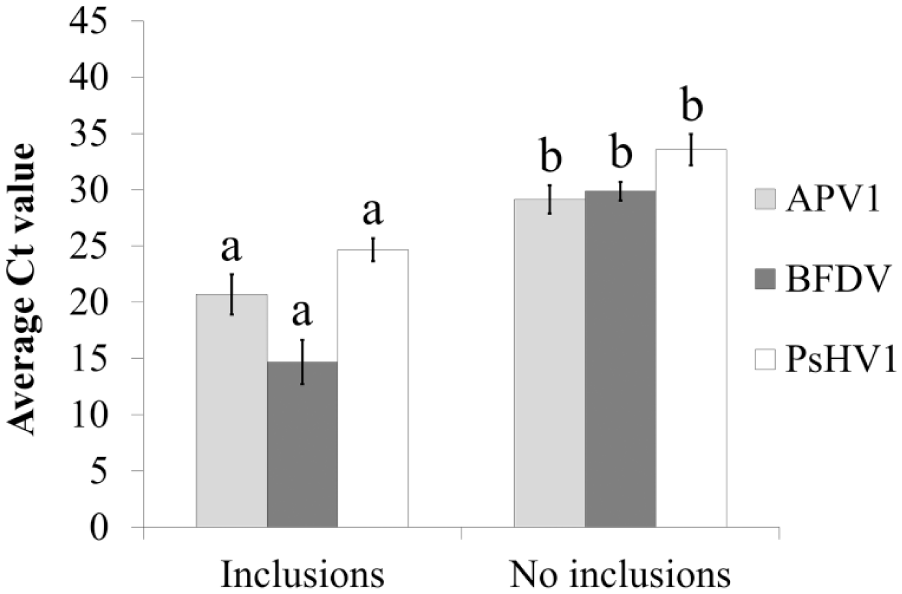
Average Ct values of samples from birds with and without histologic lesions (viral inclusion bodies) in 61 psittacine bird infections detected by real-time PCR with aves polyomavirus 1 (APV1), beak and feather disease virus (BFDV), and psittacid herpesvirus 1 (PsHV1). For each virus, different letters indicate significant differences using 2-tailed t-tests.
Utility of tissues for rtPCR testing
For the 61 birds that were diagnosed by rtPCR with 1 or more infections, 28, 22, 25, and 5 blocks that separately contained kidney, liver, spleen, and cloacal bursa or thymus, respectively. Evaluation of the Ct value yielded by these blocks showed that spleen, cloacal bursa or thymus, and kidney and liver were consistently (100%) useful for detection of APV1, BFVD, and PsHV1, respectively (Table 4). Spleen was positive in only 73% of birds, and was hence the least useful tissue for identification of BFDV infection.
Percent of positive samples (paraffin block scrolls) that contained specific tissues from psittacine birds that tested positive by real-time PCR for each virus.
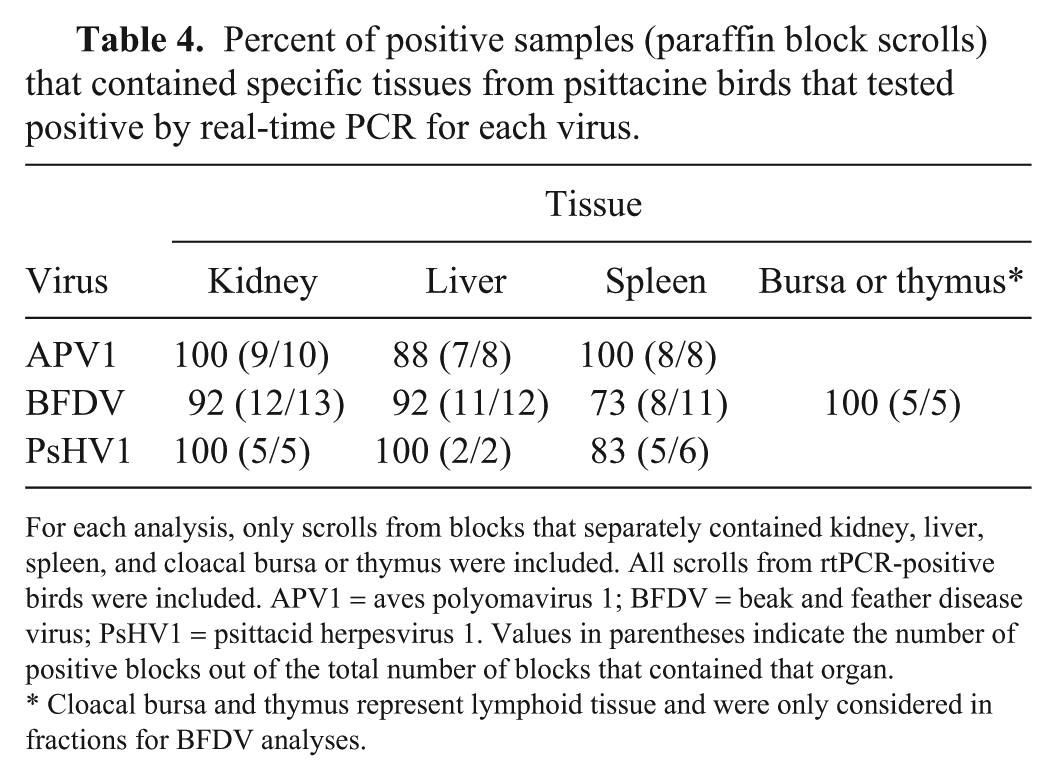
For each analysis, only scrolls from blocks that separately contained kidney, liver, spleen, and cloacal bursa or thymus were included. All scrolls from rtPCR-positive birds were included. APV1 = aves polyomavirus 1; BFDV = beak and feather disease virus; PsHV1 = psittacid herpesvirus 1. Values in parentheses indicate the number of positive blocks out of the total number of blocks that contained that organ.
Cloacal bursa and thymus represent lymphoid tissue and were only considered in fractions for BFDV analyses.
Discussion
Our multiplex assay demonstrated high analytical sensitivity and specificity for 3 important avian viral pathogens. Less than 6 copies of DNA per reaction were detected by rtPCR, indicating that viral infections may be detected in samples with very low viral loads, and the assay did not cross-react with a variety of veterinary viral and bacterial agents, suggesting high target specificity. One major advantage of our assay is the ability to simultaneously detect multiple pathogens in a single sample, allowing for more efficient testing in addition to gaining information on subclinical coinfections in birds representing natural or spontaneous disease in a given population.
We tested the utility of this assay on tissues from a cohort of psittacine birds submitted for postmortem examination at the AHL. Results showed that the overall diagnostic agreement was high between histopathology-based diagnoses of viral disease and virus detection by rtPCR. However, although most of the birds that were negative by rtPCR were also negative by histopathology, many birds that had no histologic lesions of viral infection were positive by rtPCR. This was particularly true for BFDV, which was detected by histopathology in only 41% of infected birds, as indicated by rtPCR. These data highlight the lower diagnostic sensitivity of detecting infection by histopathology compared to molecular methods, and reflect the substantial difference between disease diagnosis and infection status. Similarly, the average Ct values from birds with histologic evidence of infection was significantly lower than those without, suggesting that the viral load is higher in tissues that have evidence of virus infection.
The identification of rtPCR-positive birds without histologic evidence of infection may be attributable to latent or low-level chronic infections, which have been shown to occur with APV1, 6 BFDV, 1 and PsHV1. 28 Alternatively, acute infections might also result in the lack of histologic lesions, especially for birds that died of other (e.g., non-viral) causes. In cases of low-level infection, birds may represent sources of virus with the potential to lead to outbreaks, representing a threat to cohabiting birds or overall populations. For example, BFDV subclinical infections in adult birds used for breeding can be transmitted to offspring or other young birds in the flock, resulting in significant mortality. 22 Most coinfections in our cohort were identified by the multiplex rtPCR assay, with only one confirmed coinfection identified by light microscopy. This suggests that a bird can have non-clinical infections with more than one pathogen, and further highlights the importance of adequate screening protocols when introducing new birds to a population. Therefore, use of molecular testing would improve detection of viruses to help define the infection status of an aviary or breeding colony, and to complement quarantine protocols.
Microscopic findings (e.g., viral inclusion bodies) can be used to confirm the etiology of lesions and thus, can be helpful in disease diagnosis caused by some viruses. However, considering that inclusions can be rare or similar in appearance (i.e., nonspecific) among some viruses, this ambiguity presents a challenge in making definitive diagnoses by histopathology alone. Additionally, non-viral inclusions or pseudoinclusions (e.g., invaginations of nuclear membranes) can occur and are often impossible to differentiate from viral inclusions by light microscopy. 12 Therefore, detection of pathogen genetic material through molecular testing can be used to confirm a diagnostic suspicion or to identify more accurately the etiology of ambiguous lesions.
Based on our results, the tissues that most often yielded positive results for viral testing were the spleen for APV1, cloacal bursa or thymus for BFVD, and kidney or liver for PsHV1. However, in most cases, combinations of these tissues (i.e., liver, kidney, spleen, and thymus or cloacal bursa) were all useful for consistently detecting infections with all 3 viruses. Because APV1 and PsHV1 cause lesions in multiple parenchymatous organs, liver and/or spleen are often selected for testing to confirm these viral infections.21,28 For BFDV, primary lymphoid organs (cloacal bursa or thymus) are often suggested for testing given that BFDV replicates primarily in lymphocytes; however, specific tissues (e.g., liver) have been identified that contained sufficient levels of BFDV DNA for molecular detection. 22 Fecal swabs from herpesvirus-positive birds with latent infections can test negative, and viral shedding can resume subsequently, making some clinical samples inconsistent for detection. 28 However, testing parenchymatous tissue (e.g., biopsy samples) from these same birds accurately detected infection in non-shedding birds. 28
In addition to FFPE tissues, clinical samples, including blood, and oral or fecal swabs, are often submitted for viral testing in psittacine birds. 13 We have not validated our rtPCR assay on fresh tissue or other clinically relevant samples, such as blood or feces. The psittacine species included in the diagnostic cohorts were restricted to cases submitted for postmortem examination, and therefore our multiplex assay was not validated against the full range of potential species in the target population. Divergent genotypes are known for several of the target viruses in our study (e.g., psittacine herpesviruses, 27 for which we included all 4 genotypes in the primer design to maximize the range of PsHV1 variants that our assay will detect). Because our inclusion criteria for selecting cases for the diagnostic cohorts required the microscopic detection of inclusion bodies in affected tissue, selection bias may have resulted in psittacine species or viral genotypes with more severe disease, thereby potentially biasing the validation step of the assay; the assay may have performed differently on samples from less severe disease. Results of the assay are also semiquantitative, which allows for the inference of viral load; however, true quantitative methods are required to accurately compare Ct values with viral genetic material or virus titer.
Supplemental Material
DS1_JVDI_10.1177_1040638719870218 – Supplemental material for Development and use of a triplex real-time PCR assay for detection of three DNA viruses in psittacine birds
Supplemental material, DS1_JVDI_10.1177_1040638719870218 for Development and use of a triplex real-time PCR assay for detection of three DNA viruses in psittacine birds by Daniel J. Gibson, Nicole M. Nemeth, Hugues Beaufrère, Csaba Varga, Davor Ojkic, Anna Marom and Leonardo Susta in Journal of Veterinary Diagnostic Investigation
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This project was funded by the Ontario Veterinary College Pet Trust program (grant 052654). Mr. Gibson’s stipend was in part supported by an Ontario Graduate Scholarship.
Supplementary material
Supplementary material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
