Abstract
A survey was conducted to investigate viral infection in 253 wild marine fishes harvested in the southern coastal area of Korea from 2010 to 2012. The fish that were captured by local anglers were randomly bought and sampled for virus examination. The samples were tested for presence of virus by virus isolation with FHM, FSP, and BF-2 cells and molecular methods (polymerase chain reaction and sequencing). Of the 253 fish sampled, 9 fish were infected with virus. Aquabirnaviruses (ABVs), Viral hemorrhagic septicemia virus (VHSV), and Red seabream iridovirus (RSIV) were detected in 7, 1, and 1 fish, respectively. Molecular phylogenies demonstrated the detected viruses (ABV, VHSV, and RSIV) were more closely related to viruses reported of the same type from Korea and Japan than from other countries, suggesting these viruses may be indigenous to Korean and Japanese coastal waters.
With the development of culturing techniques in Korea, marine fish culture, including olive flounder (Paralichthys olivaceus) and Schlegel’s black rockfish (Sebastes schlegelii), has grown rapidly since the 1980s. However, because of high culture densities and environmental pollution, infectious diseases have been implicated in high mortality.1,7,9,22 In particular, viral diseases are a threat to Korean aquaculture because of the lack of available treatments. Several viral diseases have been identified from fishes in Korea, including but not limited to red seabream iridovirus disease, viral hemorrhagic septicemia, viral nervous necrosis, and lymphocystis disease.7,9,12,15
Fresh minced wild marine fish such as pacific herring (Clupea pallasii), chub mackerel (Scomber japonicus), and horse mackerel (Trachurus japonicus) have been commonly used as feed for marine cultured fish in Korea. However, several pathogenic fish viruses have been isolated or detected from wild marine fish in coastal areas of Korea.4,10,11 Thus, feeding virus-infected wild marine fish to domestic stock is considered to be a potential route of viral infection.23,25
A survey of fish viruses from wild marine fishes in coastal areas can provide significant information. Knowledge of pathogen presence in wild populations can help alert managers to potential problems in cultured fish. The origin of a virus can be understood through the phylogenetic relationship between a native virus from wild fish and a putative virus introduced from outside the country. New viruses with high pathogenicity can be found in cultured fish, and a change in the infection pattern of a pathogen or the introduction of a foreign pathogen due to an environmental change such as climate change may be recognized by a continuous survey.2,8,18,28
In the present study, a survey was conducted to investigate viral infections in wild marine fishes in the coastal area of the southern part of Korea. When viruses were detected or isolated, polymerase chain reaction (PCR) amplification and sequencing of informative gene sequences for each respective virus was performed, and phylogenetic relationships to isolates from different geographical regions were determined.
Wild fish (n = 253) were collected from the coastal areas of Yeosu, Wando, Tongyeong, and Jeju in the southern part of Korea during 2010–2012. The live fish that were auctioned at the landing centers were randomly selected and procured for virus examination. The local anglers catch fishes by trap netting or angling and sell to the middlemen who auction at the landing center. Fish samples were transported on ice and immediately subjected to virological examination. Kidney and spleen tissues were homogenized using a stomacher, a diluted 1:10 in Hanks balanced salt solution (HBSS), b and centrifuged at 3,000 × g for 20 min. Supernatants were then passed through a 450-nm membrane filter, and 100 µl was inoculated onto 3 fish cell lines in 24-well tissue culture plates, c including a fathead minnow (FHM) cell line, an olive flounder spleen (FSP) cell line, and a bluegill fry (BF-2) cell line. All cells were maintained in Dulbecco minimum essential medium d with 10% fetal bovine serum e and antibiotics (100 IU of penicillin/ml and 100 µg of streptomycin/ml). Cell cultures were incubated at 20°C for 14 days and examined for development of cytopathic effects (CPEs). If CPE was not developed after the primary incubation, 100 µl of supernatant from the primary culture were reinoculated into homologous cells according to the above-mentioned method. In order to identity the isolated viruses, the cell culture supernatants from cells showing CPEs were submitted to a neutralization test using antiserum against aquabirnavirus (ABV, isolate NC1). 6 If the neutralization test was negative, PCR for pathogenic fish viruses (Viral hemorrhagic septicemia virus [VHSV], Nervous necrosis virus [NNV], and Red seabream iridovirus [RSIV]) previously reported in Korea was performed (Table 1). Briefly, 10-fold serially diluted viral suspensions of each isolate were mixed with an equal volume of antiserum diluted 10−2 or with HBSS (control) and incubated for 1 hr at 20°C. Then, 50 µl of each mixture was inoculated onto FSP or BF-2 cells cultured in 96-well microtiter plates (4 wells for each dilution). Inoculated cells were incubated at 20°C for 14 days, and the samples showing a reduced viral titer (>103.0 TCID50) were determined to be ABV. DNA or RNA was extracted from the cell culture supernatants of cells showing CPEs using a DNA extraction kit f or an RNA extraction kit g following the manufacturer’s instructions. Polymerase chain reaction or reverse transcription (RT)-PCR was performed according to methods as previously described (Table 1)13,19,21: VGsense and VGanti primers for VHSV, F2 and R3 primers for NNV, 1F and 1R primers for RSIV. Reverse transcription PCR of ABV isolates identified by the neutralization test was also performed using the P1 and P2 primers according to methods as previously described (Table 1). 20 In addition, kidney and spleen tissues were examined for the presence of Infectious spleen and kidney necrosis virus (ISKNV) by PCR as previously described 31 because of the lack of an ISKNV-sensitive cell line in the authors’ laboratory. Briefly, DNA was extracted from the kidney and spleen tissues using a DNA extraction kit. f The PCR amplification was conducted in PCR premix h containing 20 pmol of each primer (mF1 and mR1), 40 mM KCl, 10 mM Tris–HCl (pH 9.0), 1.5 mM MgCl2, 250 µM each deoxyribonucleotide triphosphate, 1 U of Taq DNA polymerase, and 100 ng/µl of template DNA using a thermal cycler i with 35 cycles (94°C for 1 min, 50°C for 1 min, and 72°C for 1 min; Table 1).
Detailed information on the primes used for polymerase chain reaction (PCR) and reverse transcription PCR.*
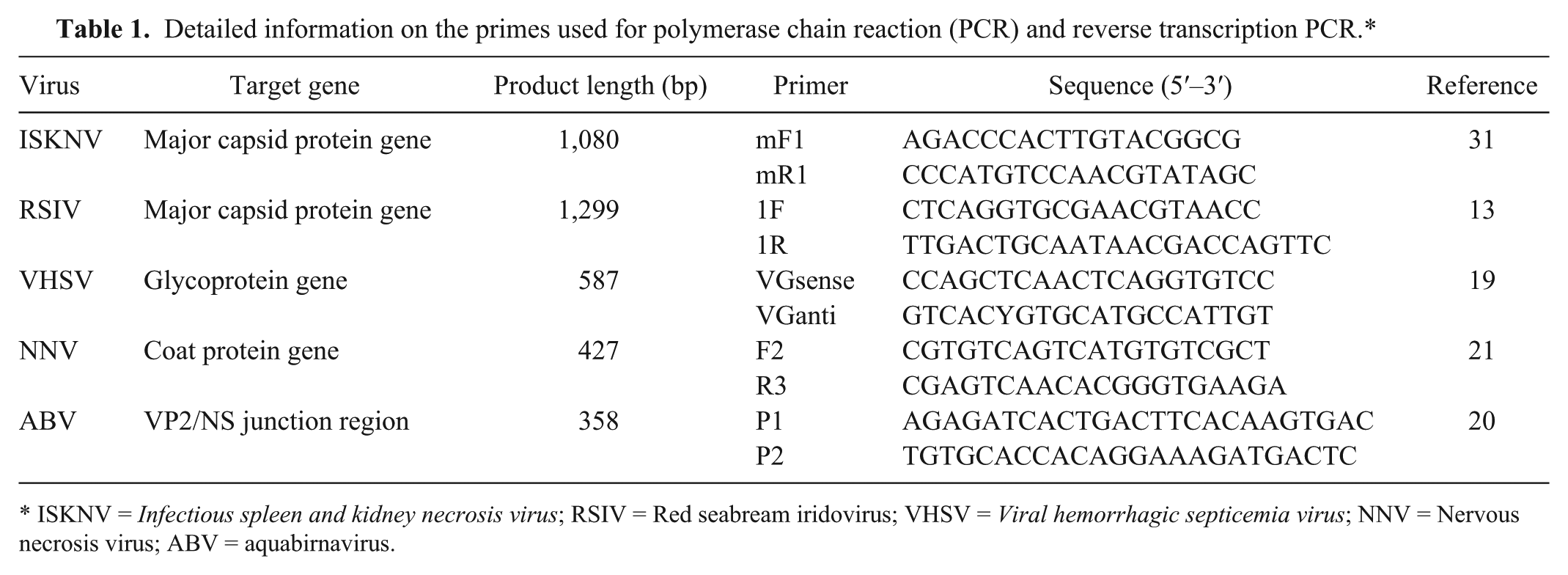
ISKNV = Infectious spleen and kidney necrosis virus; RSIV = Red seabream iridovirus; VHSV = Viral hemorrhagic septicemia virus; NNV = Nervous necrosis virus; ABV = aquabirnavirus.
The PCR products were analyzed by 1.5% agarose gel electrophoresis and purified using a gel extraction kit. j The amplified products were subjected to nucleotide sequence analysis using a terminator cycle sequencing kit k with the above-mentioned PCR primers. The resulting sequences were assembled with commercial software. l The sequences, excluding the PCR primer sequences, were submitted to GenBank (accession nos. KC693768–KC693774) and compared with those of previously reported viruses.3,5,6,11,12,19,20 A multiple alignment of the ABV sequences was constructed using ClustalX 30 to infer genetic relationships among each sequence with the neighbor-joining algorithm. The final phylogenetic tree was drawn with the MEGA4 program. 29
The results of virus detection from 253 wild marine fishes are shown in Table 2. Samples comprised fishes from 9 orders, 30 families, and 50 species. No clinical signs were observed among the fish tested. Of the 253 fish sampled, 9 fish were infected with a virus. Aquabirnaviruses, VHSV, and RSIV were detected in 7, 1, and 1 fish, respectively. No virus was detected in 95 fish from the coastal area of Yeosu in 2010. In 2011, 1 fish was ISKNV PCR positive and 7 fish were ABV positive among 126 fish from the coastal areas of Yeosu, Wando, Tongyeong, and Jeju. Viral hemorrhagic septicemia virus was only detected in 1 of 32 fish from the coastal area of Wando in 2012. From these results, it was confirmed that 5 wild fish species (yellowtail [Seriola quinqueradiata], Temminck’s surfperch [Ditrema temminckii], chicken grunt [Parapristipoma trilineatum], stone flounder [Kareius bicoloratus], and olive flounder) from the southern part of Korea were hosts for ABV and VHSV. In addition, 1 fish (striped knifejaw [Oplegnathus fasciatus], also known as rock bream) was positive for ISKNV by PCR, although this was later identified by sequencing as RSIV.
Detection of fish viruses from wild marine fishes from the coastal waters of southern Korea.*
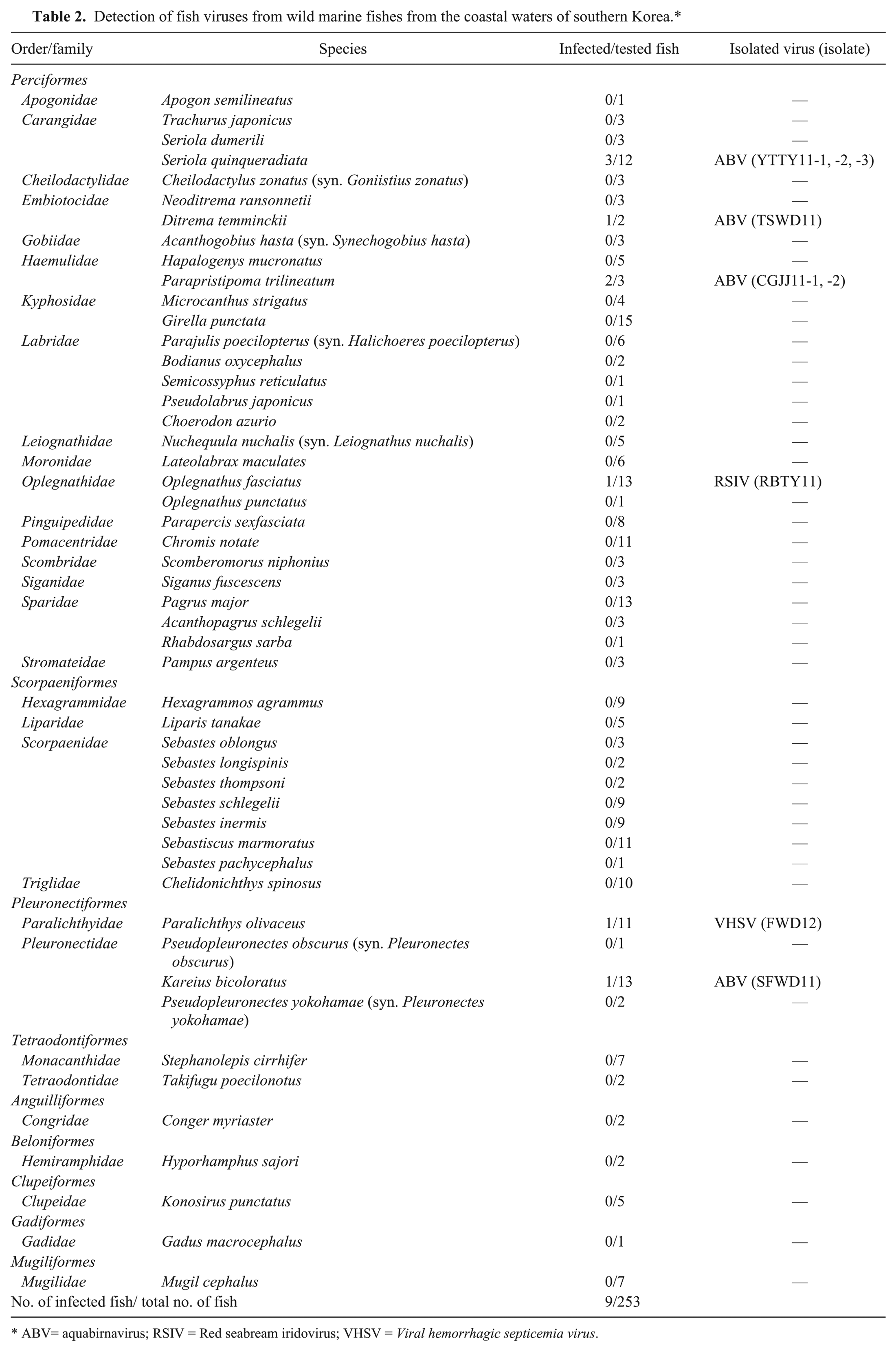
ABV= aquabirnavirus; RSIV = Red seabream iridovirus; VHSV = Viral hemorrhagic septicemia virus.
The PCR products for ABV, ISKNV, and VHSV were subjected to nucleotide sequence analysis. The nucleotide sequences were compared with those of previously reported viruses belonging to the same species. The nucleotide sequence of the VP2/NS junction gene of the 7 ABV isolates (YTTY11-1, YTTY11-2, YTTY11-3, CGJJ11-1, CGJJ11-2, SFWD11, and TSWD11) showed 94.2–100% identity to each other and more than 72.5% identity to the sequences of 97 worldwide ABV isolates (data not shown). A phylogenetic tree based on the VP2/NS gene sequences revealed 7 genogroups described in previous studies (Fig. 1).6,20 Genogroup I included strains from Canada, the United States, and Japan; genogroup II included isolates from Denmark, Taiwan, and Japan; genogroup III included isolates from France, Norway, Thailand, England, and Denmark; genogroup IV included isolates from Canada and England; genogroup V included Canadian isolates; genogroup VI included German isolates; and genogroup VII was composed of Korean isolates (FJ, FK, FWA, NC1, FM2, and DS) from cultured olive flounder and Japanese isolates from marine fish and molluscan shellfish. Seven isolates in the current study were classified into genogroup VII and were closely related to isolates from Korean and Japanese coastal areas. These results suggest that the 7 isolates could be indigenous viruses in Korean and Japanese coastal areas. Moreover, the Korean isolates from farmed olive flounder may have originated from coastal areas of Korea, but likely have not been introduced from outside the country.
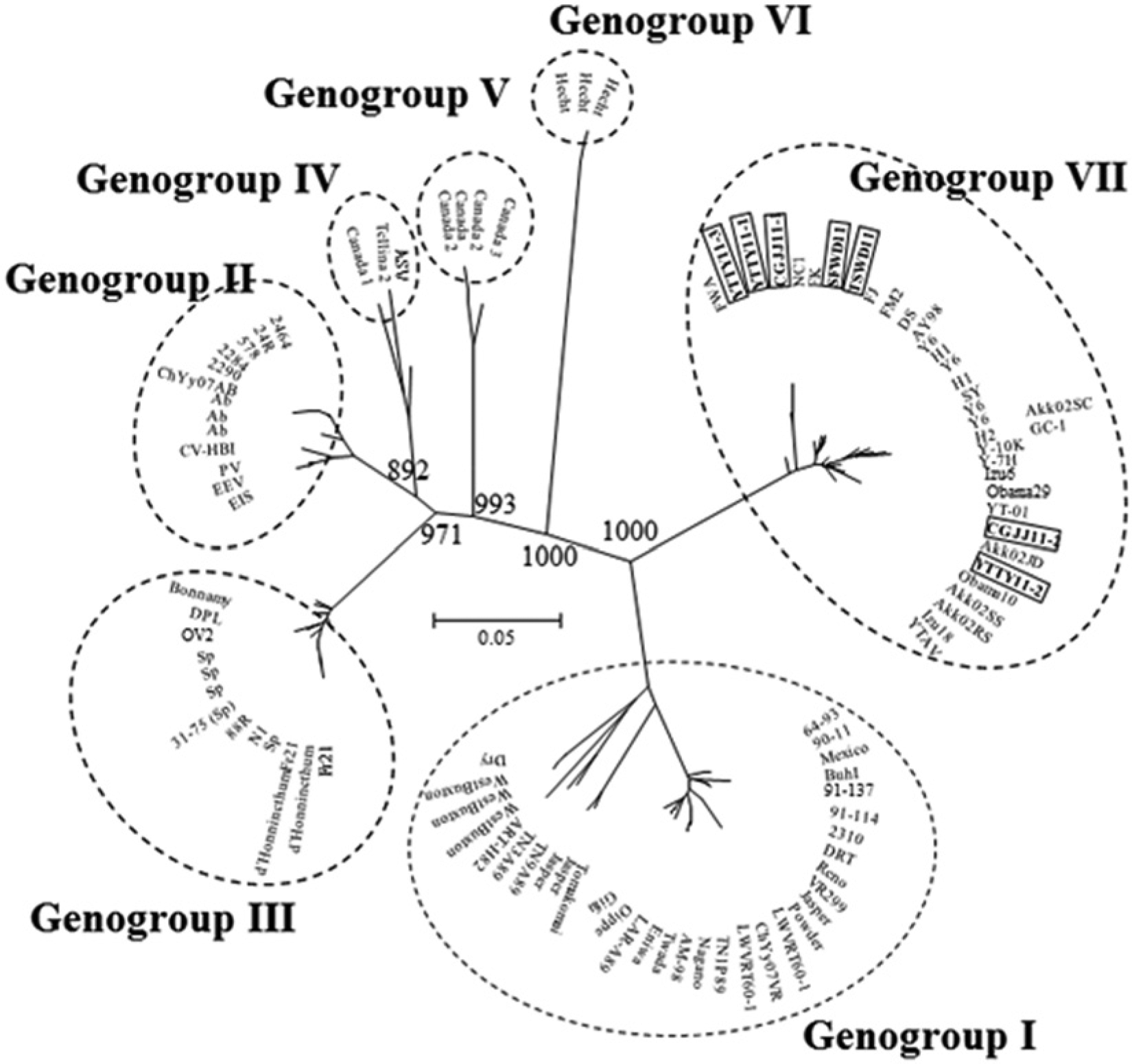
Molecular phylogenetic tree of the genetic relationship among 104 aquabirnaviruses based on VP2/NS gene nucleotide sequences. The tree was constructed by the neighbor-joining method and drawn with the MEGA4 program. Bootstrap values for 1,000 replicates are shown at major nodes in the tree. Distance marker refers to the expected number of substitutions per site. Bold character in quadrilateral indicates isolates found in the current study.
The MCP gene nucleotide sequence of the ISKNV PCR-positive virus showed 99.4% identity with RSIV (RBIV-KOR-TY1 isolate, GenBank no. AY532606) from striped knifejaw in Korea, and 94.9% identity with ISKNV (AF371960; data not shown). A phylogenetic tree based on the MCP gene sequences revealed that the isolate (RBTY11) was closely related to RSIV isolates from Korean and Japanese coastal areas (data not shown). Thus, the virus was identified from striped knifejaw as RSIV, not ISKNV. Red seabream iridovirus causes serious damage in cultured marine fishes in Korea and has been detected in various wild marine fishes in the southern and western coastal areas of Korea, but has not included wild striped knifejaw.3,7,16 Infectious spleen and kidney necrosis virus is a known virus that causes significant economic losses of several aquacultured fishes and has been isolated not only from freshwater fish species in Southeast Asian countries, including China, Indonesia, and Malaysia, but also from marine fishes caught in the South China Sea.26,27,31 Infectious spleen and kidney necrosis virus has never been detected in fishes from interior or coastal areas of Korea, even though ISKNV-like virus has been detected in imported ornamental fishes from Southeast Asia to Korea.14,17 The temperature of Korean seas is gradually increasing because of global climate change, 24 suggesting the possibility of the introduction of ISKNV from the South China Sea. Thus, although ISKNV was not detected from the samples analyzed in the current study, surveys should continue to recognize the possibility of introduction of ISKNV.
The nucleotide sequence of the VHSV partial G gene from olive flounder showed 100% identity with several Korean VHSV isolates11,12 (FJeju05, FWando05, FJeju05, BF05-1, LH03, and YBS05) from cultured olive flounder and wild marine fishes in coastal areas of Korea (data not shown). Phylogenetic analysis based on the G gene sequences of the isolate FWD12 formed 1 minor cluster including Korean and Japanese isolates, in genogroup IVa, while the North American isolates formed a different minor cluster in genogroup IVa (data not shown). The result concurs with previous studies.11,19 Viral hemorrhagic septicemia virus causes serious damage on Korean olive flounder farms and has been detected in various wild marine fishes such as chub mackerel, striped mullet (Mugil cephalus), silver pomfret (Pampus argenteus), Atlantic cutlassfish (Trichiurus lepturus, also known as largehead hairtail), hireguro (Glyptocephalus stelleri, also known as Korean flounder), and yellowback seabream (Dentex tumifrons, syn. Evynnis tumifrons).10–12 However, it has not previously been detected in wild olive flounder in Korea. Thus, wild olive flounder in Korean coastal waters could serve as a source of infection to farmed flounder in Korea.
In conclusion, the present survey revealed that 3 viruses (ABV, RSIV, and VHSV) were found in 9 of 253 wild marine fishes, and testing identified the viruses as indigenous in Korean and Japan coastal areas. In addition, the results can be helpful in providing evidence to support policies that prohibit feeding wild fish and at detecting the proximity of infected fish to their cultured counterparts.
Footnotes
a.
IUL S.A., Barcelona, Spain.
b.
Gibco, Life Technologies, Grand Island, NY.
c.
Nalge Nunc International Corp., Rochester, NY.
d.
Gibco, Life Technologies, Grand Island, NY.
e.
Gibco, Life Technologies, Grand Island, NY.
f.
Bioneer, Daejeon, South Korea.
g.
Bioneer, Daejeon, South Korea.
h.
Bioneer, Daejeon, South Korea.
i.
Bioneer, Daejeon, South Korea.
j.
Qiagen Inc., Valencia, CA.
k.
BigDye, Applied Biosystems, Foster City, CA.
l.
Genetyx Windows version 5.1, Software Development Co., Tokyo, Japan.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The current work was supported by the Technology Development Program for Fisheries, Ministry for Food, Agriculture, Forestry and Fisheries, Republic of Korea.
