Abstract
Hepatoblastomas exhibit the lowest mutational burden among pediatric tumors. We previously showed that epigenetic disruption is crucial for hepatoblastoma carcinogenesis. Our data revealed hypermethylation of nicotinamide N-methyltransferase, a highly expressed gene in adipocytes and hepatocytes. The expression pattern and the role of nicotinamide N-methyltransferase in pediatric liver tumors have not yet been explored, and this study aimed to evaluate the effect of nicotinamide N-methyltransferase hypermethylation in hepatoblastomas. We evaluated 45 hepatoblastomas and 26 non-tumoral liver samples. We examined in hepatoblastomas if the observed nicotinamide N-methyltransferase promoter hypermethylation could lead to dysregulation of expression by measuring mRNA and protein levels by real-time quantitative polymerase chain reaction, immunohistochemistry, and Western blot assays. The potential impact of nicotinamide N-methyltransferase changes was evaluated on the metabolic profile by high-resolution magic angle spinning nuclear magnetic resonance spectroscopy. Significant nicotinamide N-methyltransferase downregulation was revealed in hepatoblastomas, with two orders of magnitude lower nicotinamide N-methyltransferase expression in tumor samples and hepatoblastoma cell lines than in hepatocellular carcinoma cell lines. A specific TSS1500 CpG site (cg02094283) of nicotinamide N-methyltransferase was hypermethylated in tumors, with an inverse correlation between its methylation level and nicotinamide N-methyltransferase expression. A marked global reduction of the nicotinamide N-methyltransferase protein was validated in tumors, with strong correlation between gene and protein expression. Of note, higher nicotinamide N-methyltransferase expression was statistically associated with late hepatoblastoma diagnosis, a known clinical variable of worse prognosis. In addition, untargeted metabolomics analysis detected aberrant lipid metabolism in hepatoblastomas. Data presented here showed the first evidence that nicotinamide N-methyltransferase reduction occurs in hepatoblastomas, providing further support that the nicotinamide N-methyltransferase downregulation is a wide phenomenon in liver cancer. Furthermore, this study unraveled the role of DNA methylation in the regulation of nicotinamide N-methyltransferase expression in hepatoblastomas, in addition to evaluate the potential effect of nicotinamide N-methyltransferase reduction in the metabolism of these tumors. These preliminary findings also suggested that nicotinamide N-methyltransferase level may be a potential prognostic biomarker for hepatoblastoma.
Keywords
Introduction
Hepatoblastoma (HBL) is the most common primary liver tumor in children, accounting for ∼1% of pediatric cancers. 1 However, it is a very rare disease with an incidence of 2.6 cases per 1 million children aged 18 years and younger. 2 HBL presents a relatively normal genomic background with the lowest mutational burden reported among pediatric solid tumors. 3 These embryonal hepatic tumors usually carry cytogenetic alterations, mostly aneuploidies involving gains of chromosomes 2, 8, and 20.4–6 A few somatic mutations have already been recognized as drivers of HBL tumorigenesis, mainly as activators of the WNT pathway, with recurrent mutations in CTNNB1.7–10
This relative paucity of molecular biomarkers poses a challenge to proper risk stratification, and gene expression signatures have been reported in recent years, providing clues about specific HBL subtypes.7,11,12 A 16-gene signature discriminated two HBL subgroups that resemble early and late phases of liver development. 13 This signature stratified tumors into one group presenting fairly well-differentiated histology and favorable prognosis, and another group with a poorly differentiated histology and worse prognosis. Another model that considered differential activation of hepatic progenitor cell markers and metabolic pathways stratified HBLs in three risk groups. 7 Recently, a new approach adding epigenomics to genomic/transcriptomic data resulted in an HBL risk stratification model composed of three subgroups, based on the degree of hypomethylation, as well as the expression pattern of genes located at the 14q32 locus. 14
In a previous work, we explored the role of epigenetic mechanisms in HBL by analyzing changes in DNA methylation (DNAm) in comparison to control embryonic and differentiated liver samples; 15 a widespread and non-stochastic pattern of global low-level hypomethylation was disclosed in tumors, with enrichments at intergenic CpG sites. Loss of DNAm in HBL was also reported by Cui et al. 16 Furthermore, aberrant DNAm in specific loci has been described in HBL samples, suggesting that epigenetic alterations are an important mechanism associated with their development.17,18
Our previous work evidenced that most of the detected hypermethylated sites were mapped to CpG islands, 15 and a specific gene has drawn our attention due to its role in liver metabolism. The promoter region of NNMT (nicotinamide N-methyltransferase), a highly expressed gene in adipocytes and hepatocytes, was found to be hypermethylated in HBL. A previous study had shown that NNMT methylation detected in the fetal liver is lost in the differentiated liver and inversely correlated with gene expression. 19 This cytosolic enzyme was initially related to the N-methylation of nicotinamide, purines, and other structural analogues 20 using S-adenosyl methionine (SAM) as a methyl group donor, and the best known function of NNMT is associated with the biotransformation of drugs and xenobiotic compounds. SAM is the universal methyl donor for DNA, histones, non-histone proteins, lipids, and other metabolites, and the transfers of a reactive methyl group by NNMT to nicotinamide generate S-adenosyl homocysteine and the metabolic product 1-methylnicotinamide (1MNA). 21 This activity generates a methyl sink of 1MNA, which leads to the depletion of SAM and reduces the global methylation potential of the cell. 22
To examine whether NNMT expression in HBL could be controlled by promoter hypermethylation, we determined the methylation pattern of CpG sites mapped at the promoter region and assessed the mRNA and protein NNMT expression level in tumors. Furthermore, we explored the potential impact of NNMT changes on the metabolic profile of HBL in relation to non-tumoral control liver (NTCL) samples.
Patients and methods
Samples
The samples included in this study were HBLs surgically removed between 2016 and 2019 from patients of cancer hospitals and were selected from the biobanks for presenting at least >80% of tumoral cellularity. Thirty HBL (23 fresh-frozen and 15 paraffin block—for some samples, we had material available in both conditions) samples and 11 NTCL samples were recovered from patients enrolled in three Brazilian cancer institutions: A.C. Camargo Cancer Center, GRAACC, and ITACI (São Paulo, SP, Brazil); clinical data are summarized in Table 1. The validation cohort consisted of 15 fresh-frozen HBL samples and 15 NTCL samples from the Cincinnati Children’s Hospital Medical Center in Cincinnati (USA); clinical data of this group are summarized in Table 2. Supplementary Table 1S indicates which sample was used for each experimental procedure.
Clinical features of the 30 Brazilian HBL cases.

AFP: alphafeto protein; F: female; m: months; y: years; M: male; NA: not available; HB: hepatoblastoma; HCC: hepatocellular carcinoma.
Clinical Features of the 15 CCHMC HBL cases.

CCHMC: Cincinnati Children’s Hospital Medical Center; AFP: alphafeto protein; F: female; y: years; M: male; NA: not available; HB: hepatoblastoma; HCC: hepatocellular carcinoma.
The Research Ethics Committee of the respective institutions approved this research and the use of biological samples, and all samples were collected after informed signed consents from parents or children legal guardians.
Human induced pluripotent stem cell (iPSC) lines derived from the peripheral blood from two healthy individuals were obtained according to Okita et al. 23 Using iPSCs, hepatic differentiation was performed as described by Hay et al. 24 to obtain definitive endoderm, hepatoblasts, and precursor hepatocyte (hepatocyte-like) cells. The generation of iPSC lines derived from healthy individuals 25 was approved by the Ethics Committee of the Institute of Biosciences, University of São Paulo, Brazil (Protocol Number 1.294.118). The liver cancer cell lines, SNU-387 (ATCC® CRL-2237™), SNU-423 (ATCC® CRL-2238™), SNU-449 (ATCC® CRL-2234™), and SNU-475 (ATCC® CRL-2236™), were acquired from ATCC (USA). Following the recommended protocol, the cell lines were grown in Roswell Park Memorial Institute Medium (RPMI 1640; Thermo Fisher, USA) supplemented with 10% Fetal Bovine Serum (FBS; ThermoFisher), in conditions at 37°C and 5% CO2. The HBL cell lines HEPG2 (ATCC® HB-8065™) and C3A (ATCC® CRL-10741™) were grown in Minimum Essential Medium (MEM; ThermoFisher) supplemented with 10% FBS (ThermoFisher) and maintained at the same conditions.
Gene expression
Total RNA was isolated from HBL and NTCL samples using the RNeasy Mini Kit (QIAGEN, Germany) according to the manufacturer’s recommendations. Microfluidics-based electrophoresis (Bioanalyzer, Agilent Technologies, USA) was performed to verify the quality, and only RNA samples with RNA Integrity Number (RIN) >7.0 were used. Complementary DNA (cDNA) was synthesized with the High Capacity RNA-to-cDNA Kit (Applied Biosystems, USA) according to the standard procedures. NNMT expression was evaluated by reverse transcription-quantitative polymerase chain reaction (RT-qPCR) in 35 HBL samples and 24 NTCL samples using the TaqMan® System (Applied Biosystems), and the data were normalized to the expression level of the housekeeping gene 18S ribosomal RNA (18S rRNA). All reactions were performed in triplicate in a total volume of 10 µL containing 5 µL Master Mix, 1.5 µL water, 50 ng/µL cDNA template, and 0.5 µL of the gene-specific TaqMan Assay Probe Mixture. The delta–delta Ct (ΔΔCt) method was used for data analysis, 26 and the Kruskal–Wallis test and the post hoc Dunn test with Bonferroni correction were used for statistical analyses using GraphPad Prism 7 software.
Protein analysis
Qualitative protein analysis was performed in 14 available HBL samples from the Brazilian cohort by immunohistochemistry using the antibody anti-NNMT antibody (OTI3D8; Abcam, UK); reactions were automated in the BenchMark Ultra-VENTANA equipment. Images were obtained from Aperio Digital Pathology Slide Scanners - AT2 (Leica Biosystems, USA). HBL samples were classified as either positive or negative for NNMT protein expression.
Western blot assays were conducted using samples from the USA cohort. Protein extracts were isolated from available 10 HBL and 5 NTCL samples as previously described.27,28 This assay was performed in duplicate, where proteins (50 µg) were loaded onto a 4% to 20% gradient gel (BioRad, USA) and transferred to a nitrocellulose membrane (BioRad). The membranes were probed with anti-NNMT (OTI3D8; Abcam) and anti-beta-actin (ab8227; Abcam) antibodies. The results of Western blotting are also presented as ratios of the protein to the loading control, which were obtained by using ImageJ software (NIH, USA). 29 The t-test was used in the data analysis.
Metabolomics by nuclear magnetic resonance
Ten Brazilian HBL samples and eight NTCL samples were available for analysis by high-resolution magic angle spinning nuclear magnetic resonance (HR-MAS NMR) spectroscopy. 1H NMR spectra were acquired using a Bruker Avance spectrometer (Bruker BioSpin, Germany) operating at 400 MHz and equipped with the double nuclei 4 mm probe for HR-MAS. One-dimensional water-suppressed 1H NMR spectra were recorded with the nuclear overhauser effect spectroscopy (NOESY) pulse sequence and 128 repeats, and the T2-edited spectra were recorded using the CPMG (Carr–Purcell–Melboom–Gill) pulse sequence with 128 repetitions. All spectra were recorded at a magic angle spinning frequency of 3.5 kHz and at 293 K. Chemometrics analysis was performed using MetaboAnalyst (www.metaboanalyst.ca). Details for NMR spectra processing, data pre-processing for chemometrics, and metabolites’ assignments were previously described.30,31
Statistical analysis
To evaluate overall survival rates, Kaplan–Meier method was applied by means of the survfit function from R statistical package and the p-value from log-rank test was reported. Nonparametric Mann–Whitney–Wilcoxon rank-sum test was applied to investigate differences in NNMT gene expression levels (RT-qPCR) between pairs of groups. The Pearson correlation coefficient (ρ) between RT-qPCR relative quantification (RQ) values and metabolite variable importance in projection (VIP) scores (1HR NMR HR-MAS) per metabolite was estimated applying the Spearman nonparametric test.
Results
The DNAm values of HBL and NTCL samples were recovered from our previous work. 15 These data show that non-tumoral liver samples present a homogeneous methylation pattern, and HBL samples present a tendency of hypermethylation, although with heterogeneity. The HM450K platform contains 11 CpG sites located at the NNMT sequence, from which 10 reached all quality control parameters (Supplementary Table 2; Figure 1(a)). In Figure 1(a), boxplots of each NNMT CpG site are depicted showing the level of DNAm of HBL and NTCL samples. These boxplots and the heatmap (Figure 1(c)) showed that the CpG sites located at the 5′of the NNMT are more methylated in HBL samples than in NTCL samples. Three out of these 10 CpG sites are located at TSS1500, but only one (cg02094283, in green) was considered differentially methylated for both paired and unpaired analyses.

DNA methylation (DNAm) pattern of NNMT CpG sites in hepatoblastomas (HBLs) and non-tumoral control liver (NTCL) samples. The HM450K platform contains 11 CpG sites located at the NNMT sequence, from which 10 reached all quality control parameters; DNAm beta values (ranging from 0% to 100%) of the 10 CpG sites mapped at the NNMT gene sequence were retrieved from the Infinium Illumina 450K BeadChip arrays from previous study 15 in 19 HBL and 10 NTCL samples. Beta values were transformed into M values before performing comparison between groups, employing an empirical Bayesian framework linear model from limma. (a) The regions of the NNMT (TSS1500, TSS200, 5′ UTR, gene body, and 3′ UTR) are shown with the associated CpG sites depicted as numbered lollypops, according to their genomic coordinates; in green, the TSS1500 cg02094283 was differentially methylated for both paired and unpaired analyses.(b) Boxplots of each CpG site are presented showing the level of DNAm (%) of HBL and NTCL samples (median values indicated by lines); the asterisks (*) indicate the CpGs with significant differentially methylation (p < 0.03) between unpaired samples, and one of them (cg02094283) presents significant differentially methylation (p < 0.04) in both paired and unpaired analyses (**). (c) Heatmap showing the level of DNAm of the 10 NNMT CpG sites (the identification of each CpG site is given above in the columns) in group of tumors and NTCL; above is indicated the CpG site and below a DNAm level scale, in which blue and red correspond to lower and higher methylation content, respectively. The red asterisks marked the six differentially methylated CpG sites; 3 out of 10 CpG sites are located at TSS1500, and only one (cg02094283) was considered differentially methylated in both paired and unpaired analyses (data generated based on the results from Maschietto et al. 2016 15 ).
Significant downregulation of NNMT expression was observed in the group of 20 HBL samples compared to 9 NTCL samples (p < 0.001; Figure 2(a)). Only four HBL samples exhibited expression levels equivalent to NTCL samples, and two of them were derived from HBL patients diagnosed older than 12 years. The remaining 16 HBL samples presented NNMT expression levels similar to hepatoblasts. The analysis in iPSC samples showed a progressive increase in NNMT expression during in vitro hepatocyte differentiation; as expected, NNMT expression was very low in stem cells and derived definitive endoderm cells, while hepatoblasts exhibited a modest increase in expression, with hepatocyte-like cells presenting NNMT expression levels similar to NTCL samples. To confirm the reduction in the NNMT expression, we used a validation cohort of paired 15 HBL and NTCL samples, and a decrease in the expression of NNMT was also detected in tumor samples (p < 0.001; Figure 2(a)). Moreover, we found that NNMT expression was two orders of magnitude lower in the HBL cell lines HEPG2 and C3A than in four hepatocellular carcinoma (HCC) cell lines. The TSS1500 CpG (cg02094283) was used for the correlation analysis of DNAm (M values) versus NNMT expression (logged relative expression values), which is showed in Figure 2(b); in tumors, there is an inverse correlation between the methylation level of the TSS1500 cg02094283 and NNMT expression (R = –0.8271; p < 0.0001). In addition, using data from a published study in HBLs, 7 which evaluated 50 tumor samples, we could observe that NNMT was also more expressed in the NTCL samples, corroborating our expression findings (Supplementary Figure 1S).

Downregulation of NNMT expression in hepatoblastomas (HBLs): (a) NNMT expression analyzed by reverse transcription-quantitative polymerase chain reaction (RT-qPCR). In black dots, iPSC: induced pluripotent stem cells (iPSCs) derived from healthy individuals; DE: definitive endoderm cells; HB: hepatoblast cells; HT: hepatocyte-like cells. In blue, non-tumoral control liver (NTCL) samples from Brazilian (BR) and American (USA) cohorts. In red, tumors: HBL samples from Brazilian (BR) and American (USA) cohorts; liver cancer cell lines: in purple, HBL cell lines (HEPG2 and C3A); in gray, hepatocellular carcinoma cell lines (SNU-387, SNU-423, SNU-449, and SNU-475). For iPSC, DE, HB, and HT showing two samples for each one, corresponding to two different patients. (b) TSS1500 CpG (cg02094283) was used for the correlation analysis of DNA methylation (M values) versus NNMT expression (logged relative expression values); in tumors, there is an inverse correlation between the methylation level of the TSS1500 cg02094283 and NNMT expression (R = –0.8271; p < 0.0001).
Qualitative NNMT analysis by immunohistochemistry was performed in 15 Brazilian HBL. In hepatocytes of NTCL samples, NNMT protein labeling was observed in the membrane, cytoplasm, and nucleus. A decrease in the level of NNMT protein was validated in the majority of the HBL (Figure 3), which was detected in several cellular components, although the loss of protein labeling was heterogeneous in some tumors (Supplementary Table 3S); one sample (HB50) did not exhibit change regarding NNMT protein expression.
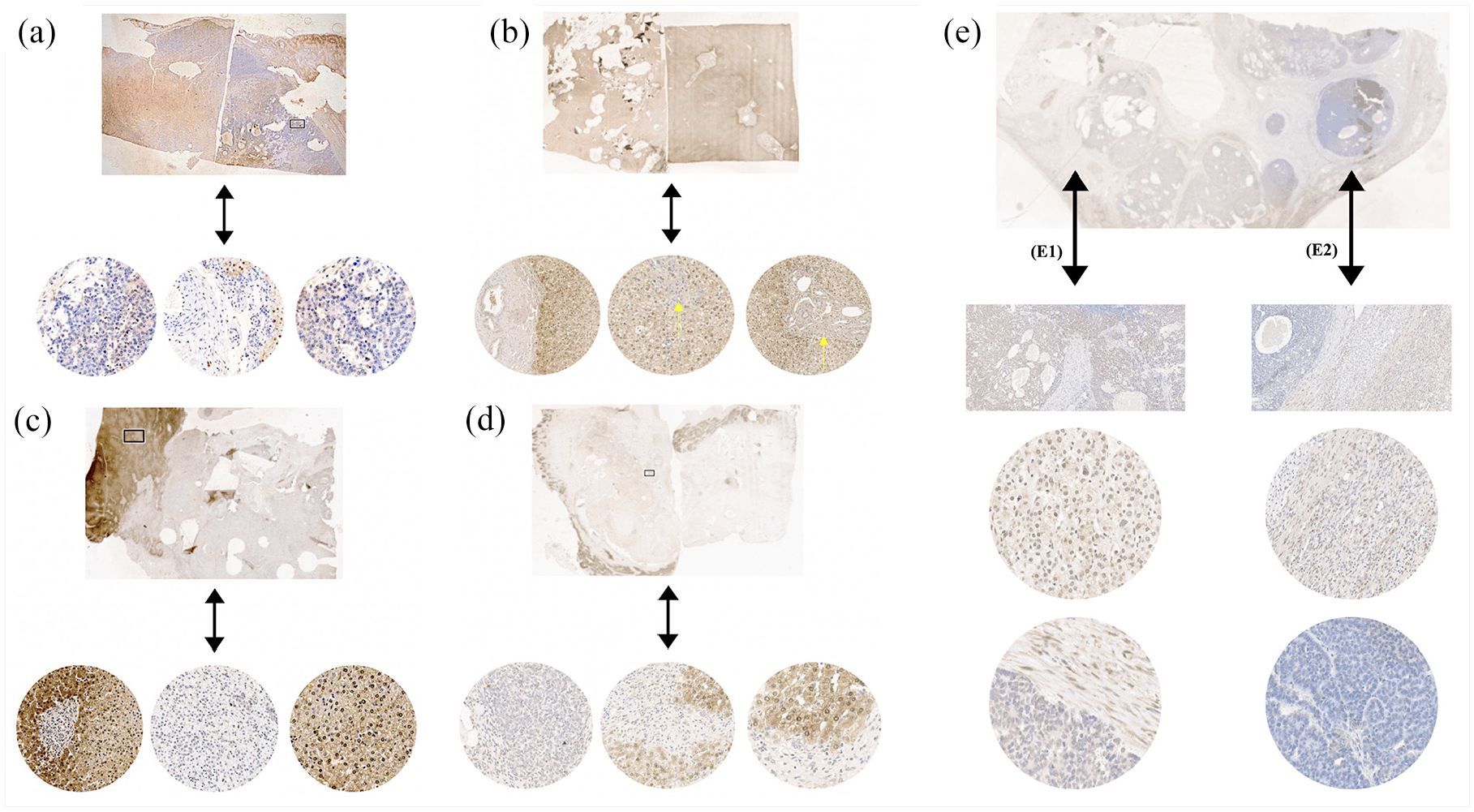
Reduction in the expression of the NNMT protein evaluated in HBLs by immunohistochemistry. In hepatocytes, the detection of the NNMT protein in the cytoplasm and nucleus is expected (brown labeling). Panels A to E represent five HBL cases, with images obtained from Aperio Digital Pathology Slide Scanners with increases of 500, 300, and 200 µm. (a) HB31 tumor showing loss of NNMT expression in some regions. (b) HB18 exhibiting loss of expression in some regions, with complete nuclear NNMT absence in specific regions, indicated by the yellow arrow. (c) HB15 and (d) HB32 exhibiting the loss of expression in some regions. (e) HB33, a congenital tumor presenting variable NNMT expression in the epithelial-fetal region (E1), and a total absence of the NNMT labeling in the epithelial-embryonal component (E2).
A Western blot assay was used in the validation cohort as another approach to verify the expression levels of the NNMT protein (Figure 4(a)). As already revealed by NNMT immunohistochemistry, the NNMT protein presented a wide reduction in HBL samples compared to NTCL samples (p < 0.001; Figure 4(b)), although its decrease was not homogeneous. In addition, the Pearson analysis showed a strong correlation of the gene and protein NNMT expression in tumor samples (R = 0.93; p = 9.2e–05; Figure 4(c)).
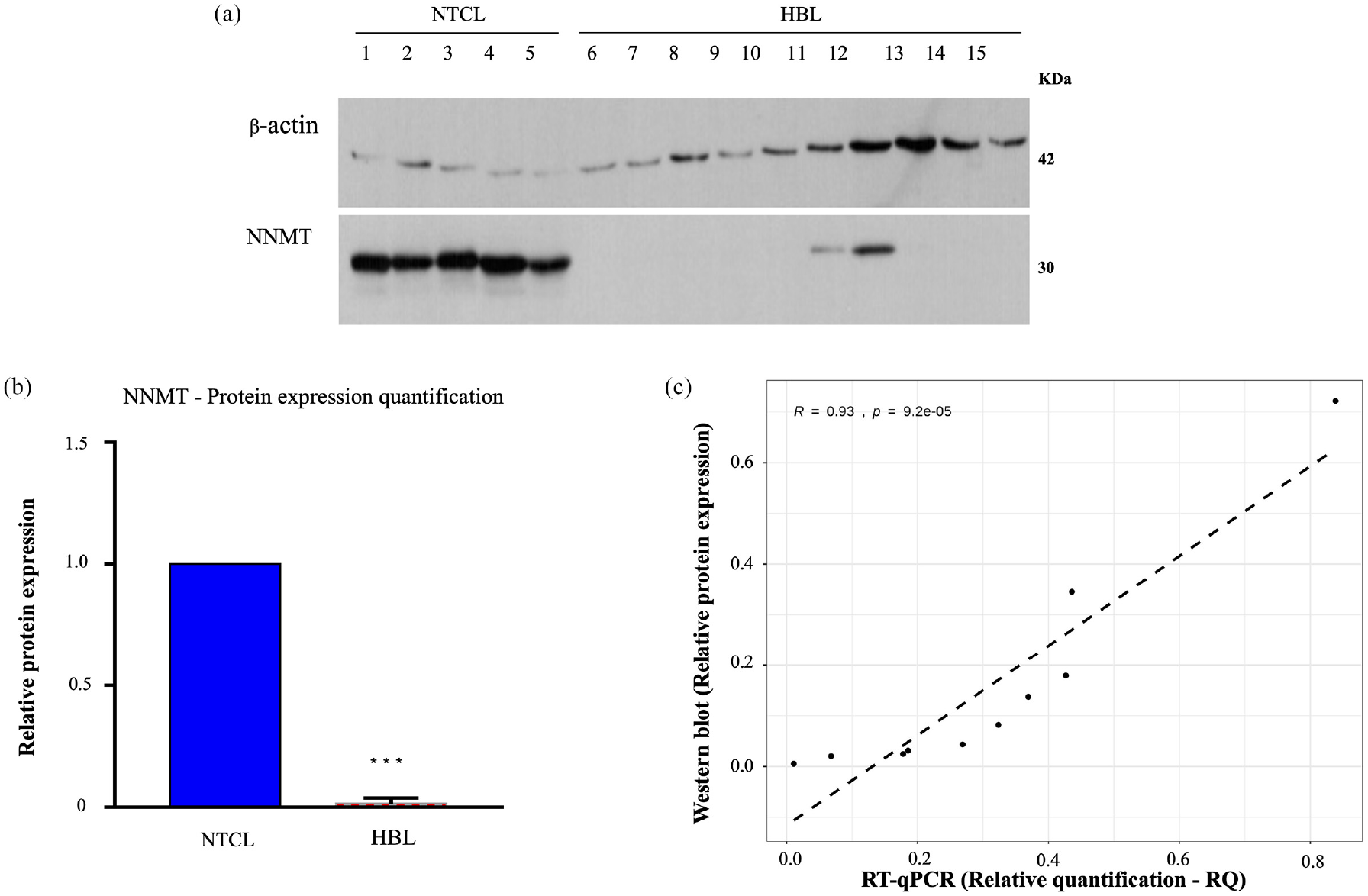
NNMT protein expression is decreased in HBLs in comparison to non-tumoral control liver (NTCL) samples by Western blot assay: (a) Immunoblotting analysis in NTCL (1–5) and HBL (6–15) samples showed a decrease in the NNMT protein in HBL. (b) Bar graph showing the NNMT protein level as a ratio to β-actin (***p-value < 0.001). (c) Correlation analysis between relative protein expression and RT-qPCR (reverse transcription-quantitative polymerase chain reaction) relative quantification (RQ) values per sample; the Pearson correlation coefficient and corresponding p-value are written in the top-left part of the graph. The dashed line represents the linear regression line.
Statistically significant differences on NNMT gene expression depending on HBL histological subtypes were not detected (Supplementary Figure 2S). Using CHIC (Children’s Hepatic tumors International Collaboration) parameters,32,33 the HBL cases were stratified according to their risk (Table 1). To evaluating the overall survival rates, Kaplan–Meier method was applied considering the level of NNMT expression (Supplementary Figure 3S), considering the first 36 months after diagnosis and the level of NNMT expression; the mean RQ value of the NNMT expression of all HBL samples was obtained (mean RQ = 0.15) and then HBL was classified as Group A: high expression level (above 0.15; n = 15 tumors) or Group B: low expression level (below 0.15; n = 20 tumors). No statistical difference was observed in HBL exhibiting higher or lower NNMT expression level than the mean of the group. Following, NNMT expression level was compared between pairs of groups according to the clinical variables associated with HBL prognosis parameters (risk stratification, age at diagnosis, alphafeto protein (AFP) dosage level in the range of 101–1000 ng/mL, PRETEXT IV, vascular invasion, metastasis, transplantation and relapse; Supplementary Figure 4Sa). It was detected a statistically significant difference related to the age at diagnosis (p < 0.015; Supplementary Figure 4Sb); patients diagnosed with more than 8 years old presented significantly higher NNMT expression. In addition, although not significant, the groups of patients with AFP levels 101 to 1000 ng/mL and high risk also appeared to be associated with higher NNMT expression.
We also investigated 10 Brazilian HBL samples and 8 NTCL samples using metabolomics by 1HR NMR HR-MAS, aiming to identify changes in metabolites that could be related to the reduction in the NNMT protein levels. HBL showed to be more heterogeneous than NTCL samples, as could be seen in the Partial least squares-discriminant analysis (PLS-DA) results (Figure 5(a)). The highest loadings and VIP values were observed for lipids, aromatic amino acids, and other metabolites that could be linked to NNMT lower activity in HBL (Figure 5(b) and (c)). In summary, 15 metabolites were detected with different concentrations between tumors and NTCL samples (boxplots in Figure 5(c)), 10 of them exhibiting increase in HBLs, and 5 with reduction. The five metabolites detected with reduced levels in tumors compared to NTCL samples were the amino acid tryptophan and four peaks of the triglyceride lipid class, used as an energy reserve source, both saturated and unsaturated fatty acids (–CH3 (C18) cholesterol; –CH3; –CH– glyceryl; –CH=CH). Among the 10 metabolites detected with increased concentrations, there are formate, some amino acids (tyrosine, alanine, phenylalanine), and several peaks of structural lipids, such as phospholipids, which are part of the composition of cell membranes. Typical 1H NMR HR-MAS spectra of HBL and NTCL samples are illustrated in Figure 5(d), where the most important metabolites (peaks 1–16) are marked.
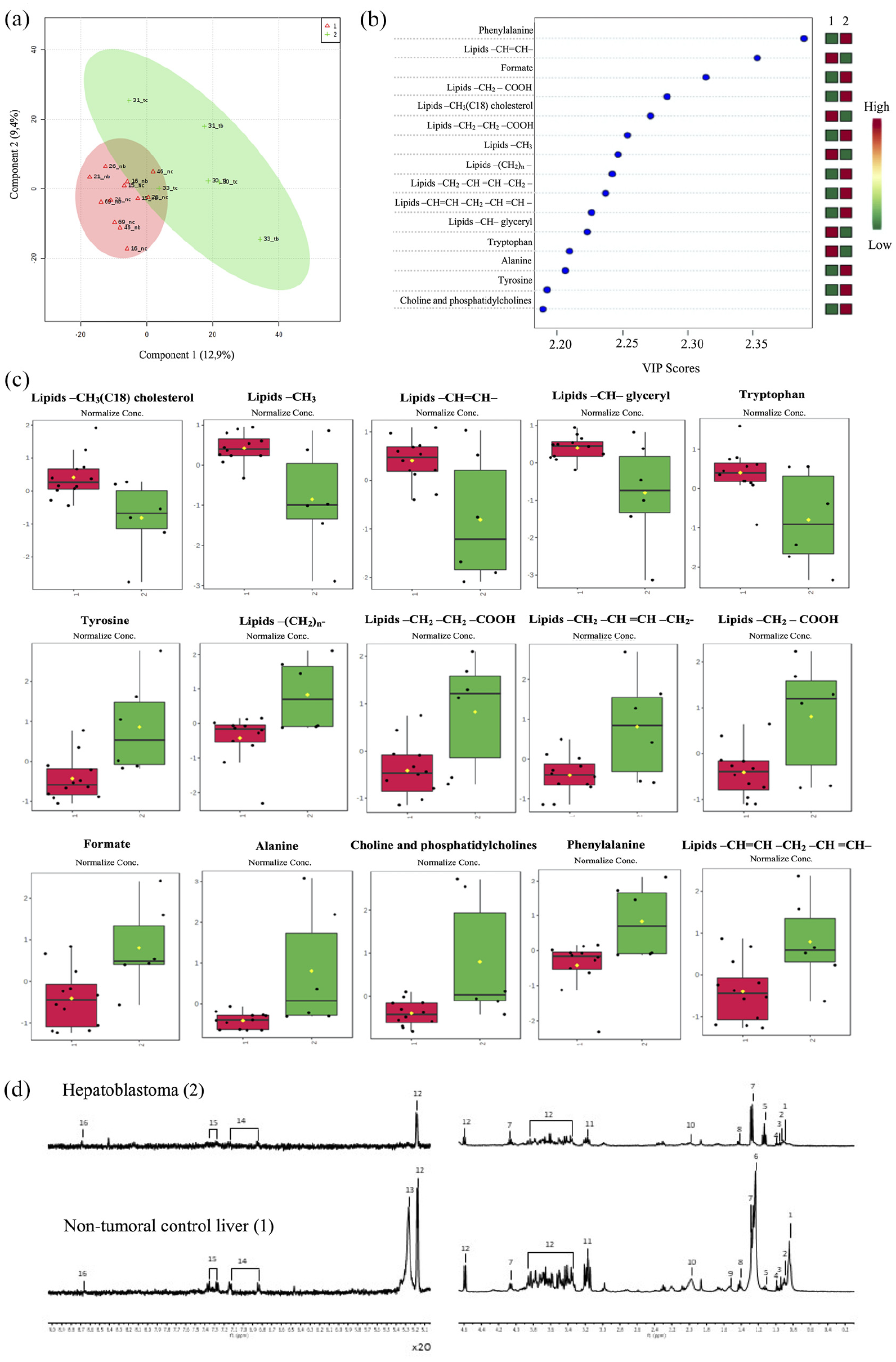
Metabolomics data obtained from HBL and non-tumoral control liver (NTCL) samples: (a) Scores of the PLS-DA model obtained using T2-edited 1H NMR (CMPG) HR-MAS spectral data, constructed from data of NTCL samples (red triangles, and indicated as Group 1) and tumor (green plus sign, and indicated as Group 2). (b) The VIP values are indicated, where the colored boxes in the right show the relative concentrations of the corresponding metabolite in each group under study. (c) The boxplots show the HBL and NTCL samples levels of the top-15 altered metabolites (Student’s t-test, p < 0.05); black dots represent the concentrations of each sample, and the yellow diamond the average values of the group (HBL illustrated in green and NTCL samples in red). (d) 1H NMR HR-MAS spectrum of HBL (upper) and NTCL samples (lower); two regions are presented between 0.20 and 4.60 ppm and amplified 20 times in the regions between 5.10 and 9.00 ppm.
Six HBLs and three NTCL samples were investigated using both methods 1HR NMR HR-MAS and RT-qPCR. Considering all studied samples, the Pearson correlation coefficients (ρ) between NNMT RT-qPCR RQ values and metabolite VIP scores (1HR NMR HR-MAS) showed strong positive correlation (ρ ≥ 0.697) in all five metabolites which are reduced in HBLs (Supplementary Table 4S and Supplementary Figure 5S), one of them statistically significant (the lipid class –CH3; p < 0.05). In addition, all 10 metabolites with increased levels in HBLs exhibited a negative correlation with NNMT expression, although not significant.
Discussion
NNMT was originally identified as the enzyme responsible for the methylation of nicotinamide (NAM), producing 1MNA, which is a precursor for NAD+, a cofactor known for donating electrons to the mitochondrial complex I and to multiple oxidoreductases. 34 Recent studies have expanded the role of NNMT, which has been increasingly associated with the regulation of multiple metabolic pathways in adipose and liver tissues through the consumption of methyl donor groups and the generation of active metabolites.35–39
NNMT overexpression has been reported in breast cancer, glioblastoma, and papillary thyroid cancer, among others.20,40,41 Ulanovskaya et al. 42 previously revealed that NNMT overexpression leads to an increase in 1MNA in various aggressive cancer cell lines; the authors proposed that the accumulation of 1MNA caused by NNMT overexpression would sequester methyl groups in cancer cells, leading to diminished methylation with changes in the epigenetic profiles.
However, it has been shown that NNMT is reduced in HCC; Kim et al. 39 analyzed 120 patients with HCC and demonstrated that NNMT expression was reduced in the majority of the examined samples and correlated with poor prognosis. Recently, these findings were corroborated in HCC samples, with the detection of significant NNMT downregulation; 43 in addition, NNMT expression was found to be heterogeneous, and tumors exhibiting high NNMT protein levels presented unfavorable prognostic features, such as vascular invasion and distant metastasis. Although the precise mechanisms by which the reduction in NNMT could contribute to liver cancer are not known, Shin et al. 44 recently revealed that NNMT depletion enhances autophagy and contributes to liver cancer cell survival and tumor growth, thus providing new insights into the mechanisms of liver cancer in adult patients. Nevertheless, very little is known about the expression pattern and role of NNMT in pediatric liver cancer.
Previous work from our group identified hypermethylation at the promoter region of NNMT in HBL. 15 As DNAm is a widely recognized epigenetic mechanism for the regulation of gene expression, 45 we speculated that NNMT dysregulation by epigenetic changes might be a plausible factor contributing to HBL tumorigenesis, deserving further investigation. Accordingly, we examined whether DNAm alterations in HBL were correlated with NNMT expression. Significant NNMT downregulation was observed in these embryonal liver tumors compared to NTCL tissues in two different HBL cohorts. Furthermore, a specific TSS1500 CpG site (cg02094283) of NNMT was hypermethylated in HBL in both paired and unpaired analyses, with an inverse correlation between its methylation level and NNMT expression, with other neighbor CpG sites following a similar methylation pattern.
In addition, regarding the reduction in NNMT observed by Shin et al. 44 in liver cancer cells, we found two orders of magnitude lower NNMT expression in HBL cell lines compared to HCC cell lines. In general, although the level of the NNMT protein was heterogeneous among HBL, remarkable reductions in NNMT mRNA and protein levels were observed using RT-qPCR, immunohistochemistry, and Western blot assays. We found a strong correlation between gene and protein expression of NNMT in tumor samples, indicating that the decrease in mRNA directly regulates the protein level in HBL.
In the HBL samples with available data, NNMT expression did not significantly impact the 36-month survival, probably because of the small sample size. However, higher NNMT expression was associated with known clinical variables of HBL prognosis. HBL from patients diagnosed with more than 8 years old exhibited higher NNMT level (p < 0.015), maybe because these tumors arose from more differentiated hepatocytes. It is also worthy to highlight that tumors from patients with AFP levels, 101 to 1000 ng/mL, and tumors from patients classified as high risk also appeared to be associated with higher NNMT expression.
Cancer cells have the ability to generate energy in a nutrient-deficient environment, and aberrant metabolism has become a characteristic hallmark of malignancies,46–48 including shifts in the metabolism of lipids.47–49 Using untargeted metabolomics by NMR, variation in many metabolites was detected in HBLs, and our data clearly showed changes in the lipid content, with an increase of the structural class of lipids, commonly detected in cancer cells because they are used for the synthesis of membranes (cytoplasmic and organelles).50,51 However, lipids used as an alternative source of energy were found to be reduced in HBLs with correlation with NNMT expression, suggesting that these tumors are using this class of molecules for their own energy demand for tumor progression.46–49,51 Structural changes in membranes, cell signaling, gene expression, protein distribution, and disruption of energy homeostasis are caused by lipid changes, and these changes may have consequences in autophagy, necrosis, apoptosis, proliferation, differentiation, growth, and chemotherapy resistance.52–54
Hepatocytes present the highest levels of 1MNA, 21 evidencing a role for NNMT in the liver. Functionally, 1MNA increases SIRT1, 55 leading to the suppression of fatty acid and cholesterol synthesis in normal hepatocytes and resulting in decreased triglyceride and cholesterol contents as well as liver inflammation. 35 Therefore, it is expected that NNMT downregulation would impact the SIRT1 function; nevertheless, the relation between NNMT and SIRT1 is complex and modulated by several factors. Adaptations of these normal conditions were reported in cancer cells, with abnormal feedback for increased or decreased NNMT content, as well as changes in other pathways of lipid metabolism, as a source of energy.34–37,56 Recent studies36,52,53 have shown that NNMT activity consumes SAM, which decreases the methylation of selected epigenetic marks, with cell-specific effects such as fat accumulation in adipocytes. Furthermore, similar to our findings, a decrease in the lipid content in liver cells, especially affecting the polyunsaturated fatty acids, was previously reported following NNMT downregulation, 57 providing an indirect evidence that supports the hypothesis that NNMT downregulation could diminish the lipid content in HBLs. Even though this hypothesis is appealing, the functional impact of NNMT downregulation should be evaluated in hepatocytes and their precursors to be causally associated with the detected lower lipid content in HBL.
This study provides further evidence that the reduction of NNMT in liver cancer, previously observed by others, is a wide phenomenon, now also validated in the embryonal tumor HBL. Furthermore, we have shown for the first time that HBL presented reduction in the level of specific classes of lipids. Taken together, our data highlight the role of DNAm in the regulation of NNMT expression in HBL and suggest a possible effect of NNMT depletion on the lipid metabolism.
The main findings of this study are summarized in Figure 6.

Main findings of this study showing effects of NNMT depletion in hepatoblastomas modulated by DNA methylation.
Supplemental Material
sj-png-1-tub-10.1177_1010428320977124 – Supplemental material for Hepatoblastomas exhibit marked NNMT downregulation driven by promoter DNA hypermethylation
Supplemental material, sj-png-1-tub-10.1177_1010428320977124 for Hepatoblastomas exhibit marked NNMT downregulation driven by promoter DNA hypermethylation by Maria Prates Rivas, Talita Ferreira Marques Aguiar, Mariana Maschietto, Renan B Lemes, Luiz Carlos Caires-Júnior, Ernesto Goulart, Kayque Alves Telles-Silva, Estela Novak, Lilian Maria Cristofani, Vicente Odone, Monica Cypriano, Silvia Regina Caminada de Toledo, Dirce Maria Carraro, Melissa Quintero Escobar, Hana Lee, Michael Johnston, Cecilia Maria Lima da Costa, Isabela Werneck da Cunha, Ljubica Tasic, Peter L Pearson, Carla Rosenberg, Nikolai Timchenko and Ana Cristina Victorino Krepischi in Tumor Biology
Supplemental Material
sj-xlsx-2-tub-10.1177_1010428320977124 – Supplemental material for Hepatoblastomas exhibit marked NNMT downregulation driven by promoter DNA hypermethylation
Supplemental material, sj-xlsx-2-tub-10.1177_1010428320977124 for Hepatoblastomas exhibit marked NNMT downregulation driven by promoter DNA hypermethylation by Maria Prates Rivas, Talita Ferreira Marques Aguiar, Mariana Maschietto, Renan B Lemes, Luiz Carlos Caires-Júnior, Ernesto Goulart, Kayque Alves Telles-Silva, Estela Novak, Lilian Maria Cristofani, Vicente Odone, Monica Cypriano, Silvia Regina Caminada de Toledo, Dirce Maria Carraro, Melissa Quintero Escobar, Hana Lee, Michael Johnston, Cecilia Maria Lima da Costa, Isabela Werneck da Cunha, Ljubica Tasic, Peter L Pearson, Carla Rosenberg, Nikolai Timchenko and Ana Cristina Victorino Krepischi in Tumor Biology
Supplemental Material
sj-pdf-3-tub-10.1177_1010428320977124 – Supplemental material for Hepatoblastomas exhibit marked NNMT downregulation driven by promoter DNA hypermethylation
Supplemental material, sj-pdf-3-tub-10.1177_1010428320977124 for Hepatoblastomas exhibit marked NNMT downregulation driven by promoter DNA hypermethylation by Maria Prates Rivas, Talita Ferreira Marques Aguiar, Mariana Maschietto, Renan B Lemes, Luiz Carlos Caires-Júnior, Ernesto Goulart, Kayque Alves Telles-Silva, Estela Novak, Lilian Maria Cristofani, Vicente Odone, Monica Cypriano, Silvia Regina Caminada de Toledo, Dirce Maria Carraro, Melissa Quintero Escobar, Hana Lee, Michael Johnston, Cecilia Maria Lima da Costa, Isabela Werneck da Cunha, Ljubica Tasic, Peter L Pearson, Carla Rosenberg, Nikolai Timchenko and Ana Cristina Victorino Krepischi in Tumor Biology
Footnotes
Acknowledgements
The authors thank the patients and their families for participating in the study.
Author contributions
M.P.R., T.F.M.A., M.M., and A.C.V.K. conceived the study and participated in its design. M.P.R., T.F.M.A., M.M., L.C.C-.J., E.G., K.A.T-.S., R.B.L., E.N., L.M.C., V.O., H.L., M.J., L.T., N.T., and A.C.V.K. performed the collection and assembly of data. M.P.R., T.F.M.A., M.C., R.B.L., S.R.C.d.T., D.M.C., C.M.L.d.C., I.W.d.C., M.Q.E., L.T., P.L.P., C.R., N.T., and A.C.V.K. realized the data analysis and interpretation. M.P.R., T.F.M.A., M.M., L.T., C.R., P.L.P., and A.C.V.K. wrote the manuscript. All authors have read and approved the final version of the manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The present study was supported by grants from FAPESP (CEPID—Human Genome and Stem Cell Research Center 2013/08028-1; 2018/21047-9; 2018/06510-4; fellowships 2015/06281-7, 2016/04785-0, 2016/23462-8) and CNPq (141625/2016-3). The funders had no roles in the study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Ethical approval
Samples were recovered from patients enrolled in three Brazilian cancer institutions: A.C. Camargo Cancer Center, GRAACC and ITACI (São Paulo, Brazil), and Cincinnati Children’s Medical Hospital Center in Cincinnati (USA). The Research Ethics Committee of the respective institutions approved the research for the used biological samples, and all samples were collected after the informed signed consents were obtained from parents or legal guardians. For induced pluripotent stem cells (iPSCs), the Ethics Committee of the Instituto de Biociências at Universidade de São Paulo, Brazil (Protocol Number 1.294.118), approved the present study, including experimental procedures involving samples from human subjects.
Guarantor
M.P.R., T.F.M.A., and A.C.V.K. are listed as guarantors of the paper.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
