Abstract
The recent investigation on PARK-2, a putative tumor suppressor gene, has found that it has been altered in multiple human malignancies. However, the clinical impact of PARK-2 alteration in uterine cervix carcinoma has not yet been studied. Therefore, we aimed to examine mutations, promoter hypermethylation, and protein expression of PARK-2 among the North Indian patients and their association with clinical parameters to evaluate the implication of PARK-2 in the genesis of cervical cancer. A total of 168 patient samples were processed for mutational analysis by single-strand conformation polymorphism, sequencing, and further in silico analysis of the identified mutations. Promoter hypermethylation by methylation-specific polymerase chain reaction and expression of PARK-2 were performed using immunohistochemistry. Statistical correlation between molecular findings and the clinicopathological parameters was taken to figure out the meaningful outcome. As per our findings, 3.5% (6/168) tumors showed novel missense mutations in exon 11 of PARK-2. In silico analysis showed high structural deviations manifested by mutations, A398D and Y391N, in both mutant proteins as compared to wild type. Promoter hypermethylation was observed in total of 29% of (48/168) tumor samples. Furthermore, 46.43% tumors (78/168) exhibited loss of PARK-2 expression in cervical carcinoma. The loss of expression of PARK-2 when correlated with clinical parameters resulted in significant association with tumor stage (p = 0.002) and with histological grade (p = 0.025). However, only clinical stage remained significant after Bonferroni correction (p < 0.007). A trend was observed between PARK-2 promoter hypermethylation and its protein expression. Our study provided sufficient information and insight for investigation of PARK-2 and highlighted its role as a tumor suppressor gene in cervical cancer in North Indian population.
Introduction
Globally, cervical cancer is the fourth most prevalent malignancy among females with an estimated incidence of 7.9% (5.3 lakh) of overall incidence and 7.5% (2.6 lakh) of overall deaths due to cancer among females in 2012. 1 It is the third major cause of mortality in underdeveloped countries where almost 9 in 10 deaths (87%) occur due to cervical cancer.1,2 In India, around 1.3 lakh new cases and 7.4 lakh deaths on account of cervical cancer occur annually, which is about one third of total worldwide deaths due to cervical cancer. 3
Infection of human papilloma virus (HPV), predominantly HPV types 16 and 18, is frequently reported during the process of cervical carcinoma.4,5 However, HPV infectivity is generally considered as rudimentary factor to transform cervical epithelium into malignant phenotype.6,7 Scientific evidences suggested the involvement of genetic and epigenetic alterations in the progression of cervical cancer pathology.8,9
PARK-2 gene is a 1.5 MB long putative tumor suppressor gene, primarily reported to be deleted and/or mutated at FRA6E locus of chromosome 6 (q25.2-27) in autosomal recessive juvenile parkinsonism (ARJP).10–13 PARK-2 contains 12 exons encoding 465 amino acids with a 52 KDa molecular weight multifaceted protein. 14 It is usually expressed in a wide variety of tissues (primarily in the muscle and brain) and functionally implicated in posttranslational mechanism regulating protein stability and activity.11,15,16 Both monoubiquitination and polyubiquitination are carried out by parkin’s E3 ligase activity involving proteosomal proteolysis, protein localization, and intracellular trafficking.17–20 The loss of PARK-2 E3 ubiquitin ligase activity leads to Lewy bodies formation in Parkinson’s disease. 21 Function of PARK-2 as tumor suppressor has been illustrated in different scientific studies and linked to tumorogenesis through its role in cell cycle regulation,22,23 mitochondrial biogenesis and hemostasis,24,25 binding and stabilization of microtubules, 20 repair of mitochondrial DNA, and identification/autophagy of damaged mitochondria.26,27 Although the mechanism adopted by PARK-2 is still not very clear but majorly it is related to different potential targets of PARK-2 such as CYC E, the accumulation of this cell cycle protein due to functional loss of PARK-2 leads to delayed proteolysis providing stimuli to cancer cell growth and mitotic instability. 22 In recent findings, the tumor suppressor role of PARK-2 is demonstrated in ubiquitination of Eps15 which plays an important role in epidermal growth factor receptor (EGFR) endocytosis and trafficking, thus, implicated in Akt signaling pathway. 18 Besides, PARK-2 is also found to be a mediator of P53 in energy metabolism and antioxidant defense thereby contributing to P53 signaling pathway. 28
In recent past, remarkable work and potential scientific investigations have substantiated the role played by PARK-2 in wide-range of tumors, including cervical cancer. 29 Tumor suppressor function of PARK-2 has been implicated in human malignancies, such as glioblastoma, 30 ovarian cancer,30,31 colorectal cancer, 32 breast cancer, 33 hepatocellular carcinoma, 34 and lung cancer.32,35 Multiple findings have demonstrated downregulation of PARK-2 primarily due to loss of heterozygosity (LOH) in breast, ovarian, and lung carcinoma,30,31 while subsequent studies described the presence of somatic mutations in PARK-2 gene affecting tumor suppressor activity in cancer such as glioblastoma, lung, and colon cancer. 32
The plethora of scientific studies clearly indicates the involvement of epigenetic factors in addition to genetic factors in tumor development and progression. 36 After PARK-2 promoter characterization 37 with the evidence of altered promoter hypermethylation of PARK-2 in leukemia and colon cancer,22,38 epigenetic deregulation of PARK-2 in carcinogenesis seems unquestionable now. The above noteworthy facts relating the role of PARK-2 in cancer and burden of cervical cancer on human population in India 39 have prompted us to further decipher the provided insight into PARK-2 molecular variations in cervical carcinoma among North Indian women.
Methods
Ethical statement
The protocol was approved by the Institutional Ethical Committee of Maulana Azad Medical College and Jamia Millia Islamia (JMI), New Delhi. Informed written consent was signed by each patient.
Tissue specimens
A total of 168 cervical cancer female patients undergoing biopsies at Maulana Azad Medical College and associated Lok Nayak Hospital, New Delhi were employed for this study. Tissue biopsies and their matched control samples (blood/precancerous lesions) were taken from each concerned subject. Tissue biopsies were collected in phosphate-buffered saline and 10% buffered formalin for immunohistochemistry (IHC). The patients were clinically staged in accordance with the International Federation of Gynaecology and Obstetrics (FIGO) standards, and histopathological features of specimens were classified according to the criteria of World Health Organization (WHO). Clinical descriptions were obtained from the medical records of hospital, including age at the time of diagnosis, tumor size, lymph node status, clinical stage, histological grade, and menopausal status, as listed in Table 1. Only cases with invasive squamous cell carcinoma were included for this study. The exclusion criteria included metastasized cases from other organs, reported prior history of any cancer, and previous exposure to radiotherapy or chemotherapy.
Clinicopathological characteristics of cervical cancer patients included in this study (N = 168).

WD: well differentiated; MD: moderately differentiated; PD: poorly differentiated.
DNA extraction
DNA was extracted as per the standardized protocol in the laboratory using standard phenol–chloroform method. Furthermore, DNA concentration and purity were analyzed by NanoDrop ND-1000 Spectrophotometer and electrophoresis on 0.8% agarose gel using standard ethidium bromide stain (Wilmington, DE, USA).
Polymerase chain reaction–single-strand conformation polymorphism
All 12 exons were considered to reveal the mutations of PARK-2 by assessing the mobility shift through single-strand conformation polymorphism (SSCP) of the amplified polymerase chain reaction (PCR) products. The primer sequence for PCR, primer size, and annealing temperature for each exon are summarized in Table 2. PCR amplification was carried out in a final volume of 25 µL, containing 100 ng of genomic DNA, 1× PCR buffer, 1.5 mM of MgCl2, 0.5 µM of forward and reverse primers, 200 µM of each deoxynucleotide triphosphates (dNTPs; deoxyadenosine triphosphate (dATP), deoxycytidine triphosphate (dCTP), deoxyguanosine triphosphate (dGTP), and deoxythymidine triphosphate (dTTP)), and 1 U of Taq DNA polymerase. The amplification was carried out under the following conditions: an initial denaturation at 95°C for 15 min, followed by 35 cycles with denaturation at 95°C for 30 s, annealing temperature (58°C–64°C) specific for each primer for 30 s, and extension at 72°C for 1 min, followed by single cycle of final extension at 72°C for 7 min.
Representative table of primer sequence for PCR and their respective temperature and amplicon size.

PCR: polymerase chain reaction; F: forward; R: reverse.
The SSCP analysis of PARK-2 exons was conducted systematically on the PCR products. PCR aliquots and loading buffer (95% formamide, 20 mM ethylenediaminetetraacetic acid (EDTA), 0.05% bromphenol blue, and 0.05% xylene cyanol) in a proportion of 1:1 were mixed and centrifuged for 15 s. Then, it was heat denatured at 95°C for 7 min and immediately incubated on ice for 10 min to maintain the denaturation. Mobility shift was assayed on 8% nondenaturing polyacrylamide gel (29:1 acrylamide to bisacrylamide) containing 50 mM of Tris–borate (pH 7.5) and 2.5 mM of EDTA prepared with 1× Tris–borate–EDTA (TBE) buffer and electrophoresed for 16–20 h at constant temperature. Silver staining method was performed as illustrated earlier. 40 DNA from normal tissue was used to differentiate the shifts obtained in tumor samples. Genomic DNA from positive PCR-SSCP samples was amplified again in 40 mL of reaction system for DNA sequencing carried out at Xcelris Labs. Ltd., India. These sequences were correlated with the wild type PARK-2 sequence (accession number: NG_008289.1) and results were compared with “GenBank” database National Centre of Biotechnology Information (NCBI). Results were confirmed by repeating the sequencing.
Structure preparation and molecular dynamics simulations
PARK-2 structure (5C1Z) was retrieved from the protein data bank (PDB) and missing residues were modeled using MODELLER V9.17 (1989–2017 Andrej Sali, University of California San Francisco). Both the mutants (Y391N and A398D) were constructed using WHAT IF server (http://swift.cmbi.ru.nl/servers/html/index.html). These structures were further subjected for energy minimization. In order to check the quality of the modeled structure Ramachandran plot was generated, which confirmed that residues were well fitted in the allowed region.
Molecular dynamics simulations
The calculations were performed with GROMACS, using the GROMOS 96 force field (http://www.gromacs.org/). The box dimensions ensured that any protein atom was at least 1 nm away from the wall of the box with periodic boundary conditions and solvated by simple point charge (spc) water molecules. NaCl counter ions were added to satisfy the electro-neutrality condition. Energy minimization was carried out using the steepest descent method. Berendsen temperature coupling and Parrinello–Rahman pressure coupling were used to keep the system in a stable environment (300 K, 1 bar), and the coupling constants were set to 0.1 and 2.0 ps for temperature and pressure, respectively. The particle mesh Ewald (PME) algorithm was employed for electrostatic and Van der Waals (VdW) interactions; cut-off distance for the short-range VdW (rvdw) was set to 1.4 nm, where Coulomb cut-off (r coulomb) and neighbor list (rlist) were fixed at 0.9 nm. All the bond lengths were constrained using the LINear Constraint Solver (LINCS) algorithm, and the time step was set to 0.002 ps. The complexes in medium were equilibrated for 1000 ps in constant Number of particles, Volume, and Temperature (NVT) and constant Number of particles, Pressure, and Temperature (NPT) ensembles, respectively. Finally, a 10 ns molecular dynamics simulation was carried out for both wild and mutant proteins. All trajectories were stored at every 2 ps for further analysis.
Analysis of molecular dynamics simulations
Structural properties of the wild and mutant (Y391N and A398D) PARK-2 were calculated from the trajectory files with the built-in functions of GROMACS. Structural analysis, such as root mean square deviation (RMSD) and root mean square fluctuation (RMSF), and radius of gyration were analyzed through the use of g_rmsd, g_rmsf, and g_gyrate, respectively, with the built-in functions of GROMACS. The number of hydrogen bonds formed within the protein during the simulation was calculated using g_h bond utility. Numbers of hydrogen bond were determined on the basis of donor–acceptor distance smaller than 3.6 Å and of donor–hydrogen–acceptor angle larger than 90°. Solvent-accessible surface area (SASA) of PARK-2 protein was analyzed using g_sas tool in GROMACS. To generate the plot for three-dimensional backbone RMSD, RMSF of carbon-alpha (Cα), gyration of backbone, and SASA analysis, we used Graphing, Advanced Computation and Exploration (GRACE) program (http://plasma-gate.weizmann.ac.il/Grace/).
Methylation-specific polymerase chain reaction
Extracted genomic DNA was subjected to bisulfite modifications using EZ DNA Methylation-Gold™ Kit (Zymo Research, Orange, CA, USA) as per the manufacturer’s instructions. Bisulfite converted genomic DNA from invasive tumor tissues and control samples was assessed for qualitative analysis of PARK-2 promoter region using methylation-specific polymerase (MSP) chain reaction. 41 Primers specific for unmethylated and methylated PARK-2 promoter regions were used as previously described. 22 Unmethylated PARK-2 promoter was amplified using primers 5′-AAGTGATTGGTTAATATGGTGGGTG-3′ (forward) and 5′-ACACAACCCAAAACCTACTAAAAATCAT-3′ (reverse) yielding 160 bp Amplicon, while methylated sequences was amplified using primers 5′-AGGTAAGTTTTTCGGTTGTTAAGCG-3′ (forward) and 5′-GCGACCCAAAACCTACTAAAAATCG-3′ (reverse) yielding 178 bp amplicon. Positive controls for methylated and unmethylated alleles were considered using commercially available totally methylated and unmethylated bisulfite converted human genomic DNA (Zymo Research). In each PCR reaction double-distilled water (ddH2O) was used as negative control.
PCR amplification was carried out in a final volume of 25 µL, containing 100 ng of modified DNA, 1× PCR buffer, 1.5 mM MgCl2, 0.5 µM of forward and reverse primer, 200 µM of each dNTPs (dATP, dCTP, dGTP, and dTTP), and 1 U of Hot Start Taq DNA polymerase (Qiagen, Valencia, CA, USA). The amplification was carried out by an initial denaturation at 95°C for 15 min, followed by 35 cycles with denaturation at 95°C for 30 s, annealing at 58°C for 30 s, and extension at 72°C for 1 min. The cycle was terminated with a final extension at 72°C for 7 min. Amplified PCR aliquots were further electrophoresed on 2% agarose gels with 100 bp DNA ladder as a standard reference and analyzed on Gel Doc (Bio-Rad laboratories, CA, USA) under ultraviolet illumination. Each experiment was repeated twice to avoid any discrepancy in the result.
Immunohistochemical analysis
Immunohistochemical staining was performed on formalin-fixed paraffin-embedded tissue blocks for each case as demonstrated by Ali et al.
42
The 4–5 µm thin tissue sections obtained on poly-
Staining interpretation
Immunostained slides were reviewed by expert histopathologist under light microscope and photographed at 400× magnification. The staining graded as low expression (upto 5%) were considered as negative and moderate expression (5%–25%) or more than 25% staining were measured as positive.
Statistical analysis
All Statistical analyses were done using Statistical Package for the Social Sciences, version 17 (SPSS Inc., Chicago, IL, USA); Fisher’s exact test was used to assess the statistical significance with two-tailed p values less than 0.05, and the confidence intervals were quoted at 95% level. In addition, each p value was statistically adjusted by applying Bonferroni correction for multiple comparisons.
Results
PARK-2 mutation analysis in cervical cancer patients
Overall, band shift was observed in 6 of 168 (3.5%) studied cervical carcinoma samples in exon 11 of PARK-2, while paired control samples did not result in any band shifts. We are hereby reporting novel missense mutations in exon 11 of PARK-2 as shown in Table 3. Sequencing analysis illustrated that all mutations were of transversion and missense type. Four samples demonstrated C to A transversion at nucleotide position 1294 leading to conversion of alanine to aspartate at codon 398, while other two mutations resulted into transversion of T to A at nucleotide position 1272 leading to conversion of tyrosine to asparagine at codon 391. Result of SSCP and sequencing histogram for positive samples are depicted in Figure 1.
Summary of mutations obtained along with codons and nucleotide alterations in tumor samples.

MD: moderately differentiated; WD: well differentiated; PD: poorly differentiated.
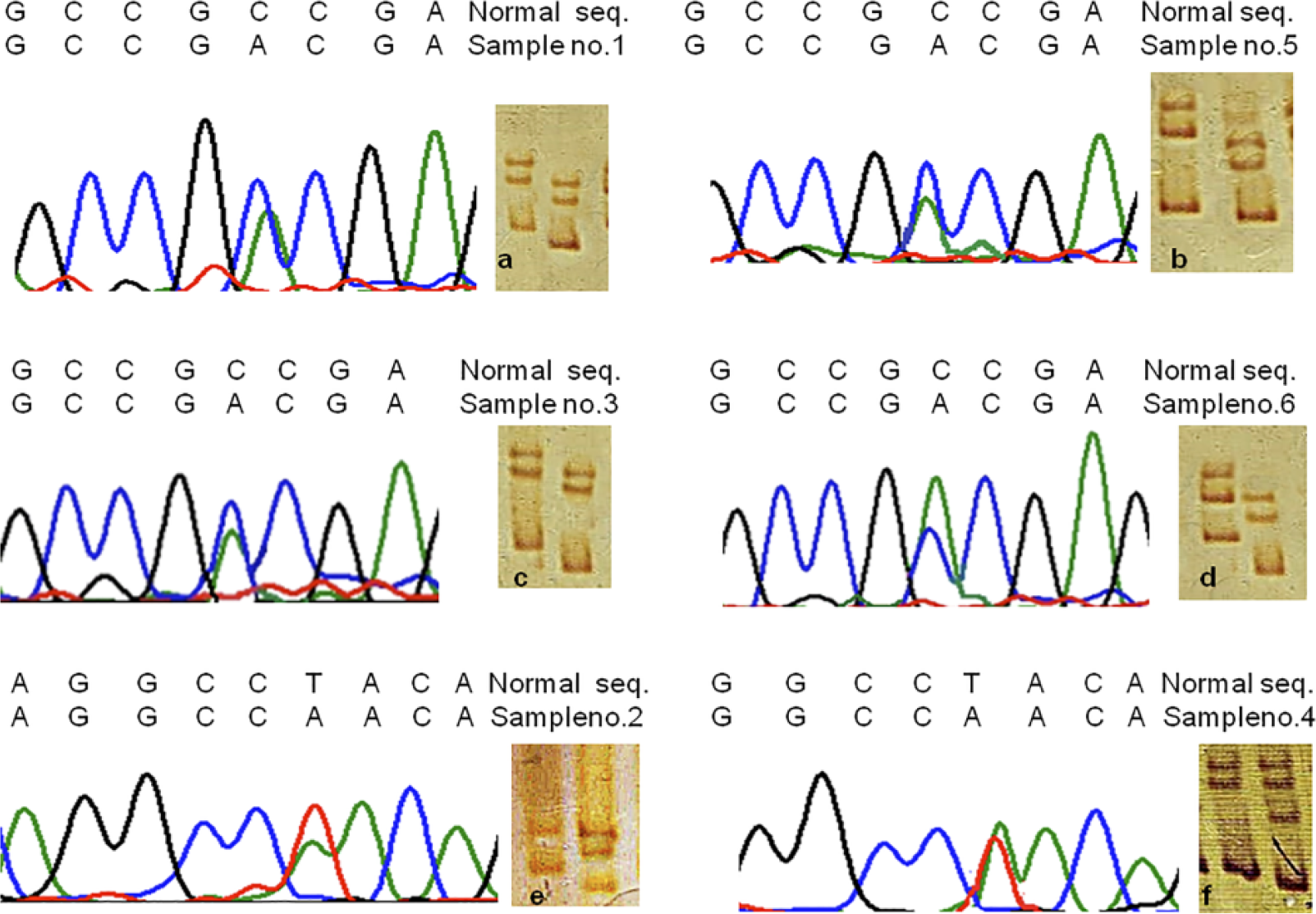
Gel picture representing SSCP and sequencing histogram for positive samples. In the gel picture, the first lane represent for normal tissue and second lane for tumor tissue (From Left) (a–d) Transversion of C to A at nucleotide position 1294 leading to conversion of alanine to aspartate at codon 398. (e and f) Transversion of T to A at nucleotide position 1272 leading to conversion of tyrosine to asparagine at codon 391.
Molecular dynamics simulations
We performed molecular dynamics simulations for wild type and other two substitutions (Y391N and A398D) to understand the internal motions and assess the conformational changes within the nanosecond time scale for these structures. The energies were calculated for NVT and NPT to confirm that the system is equilibrated. The simulation data were also subjected to energy calculation which showed that the system is equilibrated within 100 ps of NVT and NPT. The temperature for 100 ps NVT equilibration and the pressure graph for 100 ps NPT showed that temperature of the systems attained the value (300 K) quickly and remained stable in wild and both mutants PARK-2 (Figures S1 and S2). The density was also stable, indicating that the system was equilibrated within 100 ps of NVT and NPT each, with respect to temperature, pressure, and density (Figure S3). The structure properties, such as RMSD of the backbone and Cα atoms were calculated for wild type and the two mutant types of PARK-2 as function of time (Figure 2(a) and (b)). The mutant A398D showed high deviation in the structure compared to wild and another mutant Y391N. The structure of Y391N showed deviation higher than the wild type after 5 ns. The results suggested that both A398D and Y391N were deviated from wild type, and structures were stable after 5 ns. The average value of RMSD for backbone atoms was calculated from 5–10 ns Molecular Dynamics simulation data. The average value of wild type PARK-2 was 0.3831 nm, whereas, the average value of RMSD for backbone atoms in mutants Y391N and A398D was 0.392 nm and 0.4519 nm, respectively (Figure 2(a)). The plot of radius of gyration, Rg, (protein) versus time (Figure 2(c)) showed that mutants were more compact than wild type. The average values of radius of gyration (Rg-protein) of wild type, Y391N, and A398D mutants were 2.698, 2.680, and 2.646 nm, respectively (Figure 2(c)). SASA was calculated for wild type and mutants trajectories (Figure 2(d)). The average SASA of wild, A398D, and Y391N was 256.5, 261.5, and 257.4 nm2, respectively. This showed that wild type PARK-2 is more buried inside than both the mutants. RMSF of each amino acid residue (Figure 3) showed the difference in the dynamics of the core of wild type and mutant forms of PARK-2, where the overall flexibility of both the mutants PARK-2 decreased when compared to wild type PARK-2. THR-125 displayed a large flexibility mainly for Y391N and followed by A398D, but wild type showed very less flexibility for the particular region. In case of LEU-107, wild type showed high flexibility followed by A398D. Both these residues were present in the loop region of the PARK-2. The THR-125 residue fluctuated highly in mutants (Y391N: 0.9325 nm and A398D: 0.5121 nm) as compared to wild type (0.3899 nm). The LEU-107 residue fluctuated highly in wild type PARK-2 (0.7586 nm) as compared to mutants (A398D: 0.5971 nm and Y391N: 0.3182). From the RMSF analysis, we found that THR-125 is important for the role of both the mutants PARK-2.

Structural properties of wild type and mutant PARK-2. (a) Backbone RMSD of PARK-2. (b) C-alpha RMSD of the PARK-2. (c) Radius of gyration of protein. (d) Solvent-accessible surface areas (SASAs) of PARK-2.

Root mean square fluctuation of wild type and mutant PARK-2. (a) C-alpha RMSF of PARK-2. (b) PARK-2 structure highlighting two fluctuating residues.
PARK-2 promoter hypermethylation and correlation with clinicopathological parameters
Promoter hypermethylation of PARK-2 was detected in 48 of 168 (29%) tumor samples of cervical cancer, and none in the normal tissue samples. Result of PARK-2 promoter methylation is represented by gel image (Figure 4), indicating PCR amplified products obtained using the respective primers of methylated and unmethylated alleles. Furthermore, we evaluated statistical correlation of PARK-2 promoter hypermethylation with clinical factors (Table 4). However, PARK-2 promoter hypermethylation was not found to be associated with any clinical markers, such as patient age, tumor stage, tumor size, lymph node status, histological grade, and menopausal status.

MS-PCR gel picture representing promoter hypermethylation of PARK-2 in cervical tumors—T: tumor specimens, PC: positive control, NC: negative control; L: ladder (100 bp), M: methylation, and UM: unmethylation.
Correlation of PARK-2 promoter hypermethylation with clinical parameters in cervical cancer specimens.

OR: odds ratio; CI: confidence interval; WD: well differentiated; MD: moderately differentiated; PD: poorly differentiated.
p value was calculated by Fisher’s exact test.
Expression analysis of PARK-2 protein in cervical cancer patients
Overall, 78 out of 168 (46.43%) tumors exhibited negative expression of PARK-2 protein in the total sample studied. Results of PARK-2 protein expression are shown in Figure 5. While analyzing statistical correlation with clinical parameters, we found a significant association between the PARK-2 negative expression with the advanced tumor stage (PARK-2 protein negative expression in tumor stage I + II was 32.89% (25/76) whereas in tumor stage III + IV was 57.61% (53/92), p = 0.002) and higher histological grade (PARK-2 protein negative expression in well differentiated (WD) histological grade was 34.92% (22/63) whereas in moderately differentiated (MD) and poorly differentiated (PD) histological grade was 53.33% (56/92), p = 0.025) as shown in Table 5. No statistical correlation was found with other clinical parameters (p > 0.05). In addition, during the course, we could not establish any significant association with PARK-2 promoter hypermethylation and PARK-2 negative expression (p = 0.06) as shown in Table 6. However, we reported increased frequency of methylation in PARK-2 negative tumors 28/78 (35.9%) as compared to low frequency in PARK-2 positive tumors 20/90 (22.22%). A graphical representation depicting the association of PARK-2 expression with promoter hypermethylation and clinical parameters (clinical stage and histological grade) is presented in Figure 6.

Representative pictures of IHC on PARK-2 protein expression in cervical squamous carcinoma at 400× magnification. Positive expression in (a) normal and (b) cancerous tissue and negative expression in (c) cancerous tissue (scale bar: 1000 µm).
Correlation of PARK-2 expression with clinical parameters in cervical cancer specimens.

OR: odds ratio; CI: confidence interval: WD: well differentiated; MD: moderately differentiated; PD: poorly differentiated.
p value was calculated by Fisher’s exact test.
Bold represents the p Value 0.002 is significant even after Bonferroni correction.
Bonferroni significance level p < 0.007.
Association of PARK-2 promoter hypermethylation with its protein expression.

P value (Fisher’s exact test), OR odd ratio, CI confidence interval
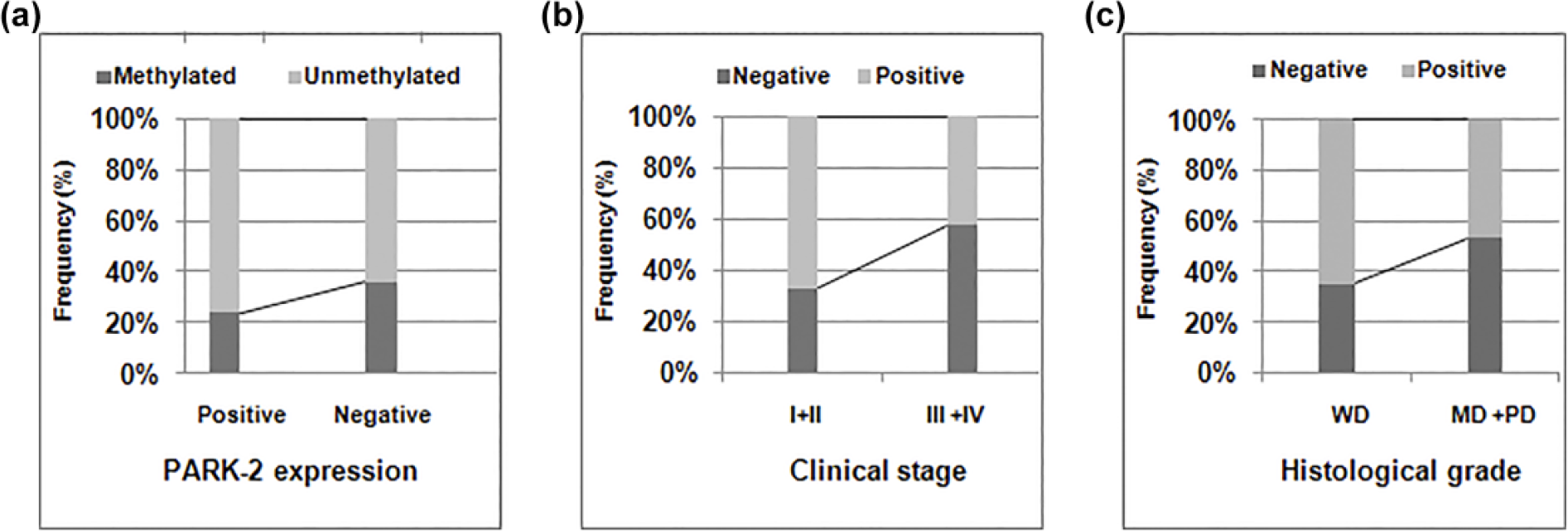
Frequency distribution of PARK-2 promoter hypermethylation with (a) protein expression and (b) frequency distribution of PARK-2 expression with (b) tumor stage and (c) histological grade.
Discussion
Uprising research on cancer has defined the genetic and epigenetic basis as fundamental but complex phenomenon that interacts during the development and progression of malignant cells into invasive tumors. 43 Though numerous population based studies have been performed but due to poorly understood biology, multiple risk factors, 44 molecular abnormalities, 45 as well as genetic diversity necessitated continuous investigation of novel targets in cervical carcinogenesis. This study, therefore, aimed at determining the contribution of genetic and epigenetic aberrations among novel and critical tumor suppressor component (genes), such as PARK-2 linking it not only with cervical carcinogenesis but their association with pathological markers which could be a promising approach in combating the fatal disease in future.
Worldwide, there are little studies made on the involvement of PARK-2 with the cancer of uterine cervix. Recently, our laboratory has reported significant association of LOH at PARK-2 locus with cervical carcinoma in North Indian population 29 in addition to ovarian cancer. 46 Hence, we intended to investigate the presence of tumor-specific mutations, promoter hypermethylation status, and expression pattern of PARK-2 in addition to the association of these variables with the clinical parameters among North Indian cervical cancer patients.
To date, somatic mutations of PARK-2 in cervical carcinoma have not been reported anywhere in the world. Here, we report novel tumor-specific missense mutations Ala398Asp and Tyr391Asn in exon 11 of PARK-2 in cervical cancer patients. Both these mutations lie on the linker helix of PARK-2 adjacent to RING2 domain at C terminal. 47 Reliable information regarding genotypic to phenotypic effect of such mutations in proteins can be accurately predicted using in silico analysis. Certain amino acid substitutions in protein sequences are more deleterious than others, eventually altering a protein structure drastically to affect its function negatively. 48 Point mutation at conserved locus Ala398 has been previously reported in Parkinson’s disease (PD) patients where it codes for different amino acid. 49 The missense mutation Tyr391Asn has not been reported earlier in cancer or in PD patients, and it was reported that this region from 391 to 396, N-terminal to linker helix, plays crucial role in PARK-2 functioning. 47 Therefore, identification of mutation at this spot of linker helix in our studied population may affect PARK-2 interaction with its substrates promoting tumorogenesis. 47 We analyzed both the wild type and mutant protein structures from the data obtained using RMSD, RMSF, and SASA analysis. RMSD results showed that the structural deviations were high in both A398D and Y391N mutants, wherein comparison to wild type the structural deviation was higher in A398D mutant than in Y391N. Since proteins are highly dynamic molecules and their inherent movement is very crucial for their proper functioning, these structural deviations found in mutant PARK-2 can thereby greatly influence normal functioning of PARK-2 protein. RMSF of each amino acid showed increase in structural rigidity in both the mutants. Minor decrease in flexibility was observed near the mutation sites but THR-125 position became highly flexible in both the mutants compared to wild type. Thus, an important role of THR-125 may be possible in PARK-2 protein because of these mutations. Studies have documented that tumor specific mutations affect the ligase activity of PARK-2 and its ability to ubiquitinate substrate in cancer cells. 21 Based on our in silico results of structural deviations in PARK-2, we therefore, suggest that the mutations identified in this study might show similar effects. Previously, in silico studies have successfully demonstrated how these predictions affected function of proteins both in vitro and in vivo.50,51 Significance of any of the identified mutations in relation to any clinical factor was not determined in this study.
In promoter hypermethylation analysis, incidence of PARK-2 promoter hypermethylation in cervical carcinoma accounted by our study is in conformity with earlier study where comparable hypermethylation frequency in acute lymphoblastic leukemia and chronic myeloid leukemia was reported 37 while low frequency was stated in colon cancer. 22 However, in stratified analysis, association of promoter methylation with any of the clinical parameters was not found to be significant.
We further analyzed PARK-2 expression in a series of North Indian cervical cancer patients and hereby report negative expression of PARK-2 in 43% tumors. This finding is in consonance with some previous reports exhibiting downregulated expression of PARK-2 in other malignancies such as breast, 33 ovarian,31,46 lung22,31 and colon 22 cancer. Unlike hypermethylation of PARK-2 promoter, we observed statistically significant association of PARK-2 expression loss with tumor stage and histological grade; this kind of association appears to be novel and has not been reported previously. Overall, our findings indicate correlation of loss of expression of PARK-2 with aggressiveness of tumor. The progression of uterine cervical carcinoma has been associated with distinct and sharply defined clinical stages. 52 In our study, association of advanced clinical stage with PARK-2 expression loss was found to be significant even after Bonferroni correction (p < 0.007). This underlies its relevance to cervical cancer pathogenesis among North Indian women. In this study, we tried to figure out a trend of association of PARK-2 promoter hypermethylation and its expressional loss, but there did not appear any statistical significance. This indicated that promoter hypermethylation is only partly responsible for its downregulation and other molecular mechanisms such as LOH16,29 might also be accountable for biallelic inactivation of PARK-2 in cervical cancer. Parkin loss of expression has been related with loss of ubiquitination of cell cycle components such as CYC E. The accumulation due to its constitutive expression could lead to mitotic instability and cell cycle deregulation eventually leading to tumor progression. This finding has been illustrated in colorectal cancers. 22 PARK-2 depletion increased the proportion of cells in the S and G2–M phases and led to increased frequency of abnormal mitosis, multipolar spindles, and genomic instability. 23 Furthermore, PARK-2 loss is predicted to lead to accumulation of defective mitochondria and possibly altered susceptibility to apoptosis since PARK-2 has been suggested to contribute in P53 signaling pathway.22,28 Besides, factors such as posttranslational modifications (biochemical and protein–protein interaction) could also lead to loss of its catalytic activity. 53 Although these findings still did not indicate so strongly about the exact mechanism and regulation of PARK-2 in tumor tissues; however, it can be concluded that PARK-2 is involved in multiple functions and is regulated through varied factors which could lead to its differential activity and ectopic expression in tumor cells.
To the best of our understanding, we are the first to report the evaluation of mutations and its in silico analysis along with promoter hypermethylation of PARK-2, protein expression in addition to the correlation with relevant clinicopathological profiles of cervical cancer patients. However, our findings need to be confirmed on larger data sets in multiple populations of the world. Moreover, for the North Indian patients where we have reported its relevance with the clinical stage (late) may provide some insights toward the development of battery of a test system that could ultimately help in the disease management.
Conclusion
Our observations have made us to believe that PARK-2 alterations during the course of cervical carcinogenesis are highly relevant and crucial events that ultimately established PARK-2 as a putative biomarker. Furthermore, keeping the pattern of epigenetic silencing and loss of protein expression in view, this gene is understood to offer an array of prognostic, therapeutic, and management strategies/designs which could be exploited by the researchers in future for the greater benefits of cancer patients.
Footnotes
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was funded by University Grants Commission (UGC), Government of India, New Delhi.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
