Abstract
There is an increasing body of evidence which highlights the critical functions of long non-coding RNAs in the carcinogenicity mechanism of a variety of cancers. It has been reported that HOX transcript antisense intergenic RNA, a member of long non-coding RNA family, increases breast cancer risk. To date, no data regarding the association between HOX transcript antisense intergenic RNA polymorphisms and the risk of breast cancer development has been reported in Iran. Here, we examine the possible association between HOX transcript antisense intergenic RNA gene polymorphisms and breast cancer in a sample of southeast Iranian female population. The HOX transcript antisense intergenic RNA rs920778, rs12826786, rs4759314, and 1899663 gene polymorphisms were genotyped in 220 cases and 231 controls by polymerase chain reaction–restriction fragment length polymorphism. Our findings indicated that rs920778 polymorphism has significant positive association with breast cancer; rs12826786 and rs1899663 polymorphisms demonstrated significant negative association with breast cancer; and the rs4759314 variant was not associated with breast cancer risk. Haplotype analysis revealed that TGAC, CTAT, and TTAT haplotypes significantly decreased the risk of breast cancer compared with rs920778T/rs1899663G/rs4759314A/rs12826786T haplotype. In conclusion, we investigated only four variants of HOX transcript antisense intergenic RNA gene, and the findings suggest that HOX transcript antisense intergenic RNA rs920778, rs12826786, and rs1899663 polymorphisms may be associated with breast cancer risk in a sample of southeast Iranian population. Further replication studies with other polymorphisms of HOX transcript antisense intergenic RNA gene involving a greater sample size and different ethnicities are necessary to verify our findings.
Introduction
Breast cancer (BC) is the most common cancer among females, and the second leading cause of cancer-related mortality in women worldwide. 1 Both environmental and genetic risk factors contribute to the occurrence of BC. 2 Single nucleotide polymorphisms (SNP) in the DNA sequence—involved in creation of different genotypes—may be associated with the risk of BC. 3
MicroRNAs (miRNAs) and long non-coding RNAs (lncRNAs) are considered as the two major families of non-coding RNAs (ncRNAs). LncRNAs are a main class of non-protein-coding RNA molecules greater than 200 nucleotides in length.4,5 Growing evidence suggests that lncRNAs could regulate gene expression at transcriptional and post-transcriptional levels, via interaction with DNA, RNA, and protein.6–8 It has been proposed that lncRNAs, functioning as oncogenes or tumor suppressors, play critical roles in a wide range of biological processes including differentiation, proliferation, and metastasis.9,10 Altered expression of lncRNAs is shown to be related to carcinogenicity and progression of many human cancers.6,11,12
HOX transcript antisense intergenic RNA (HOTAIR) has been recently identified as an oncogene. It plays a critical role in gene regulation and chromatin dynamics and appears to be persistently overexpressed in a variety of cancers. HOTAIR overexpression is associated with tumor invasion, progression, metastasis, and poor prognosis of corresponding cancers.13,14 HOTAIR gene, a 2158 nucleotides in length lncRNA, is transcribed from the HOXC locus located on chromosome 12q13.13 15 and has an important role in gene regulation via the alternation of chromatin structure (histone modifications).15,16
Several studies investigated the impact of HOTAIR gene polymorphisms on the incidence risk of various cancers including BC, gastric cancer, prostate cancer, and ovarian cancer.17–25 However, the data from these studies remain controversial and inconclusive. Furthermore, few studies have been conducted regarding the association of HOTAIR gene polymorphisms on BC risk.17–19 To the best of our knowledge, there have been no reports regarding the association of HOTAIR polymorphisms and BC risk in the Iranian population. We therefore performed a case-control study to evaluate the possible association between HOTAIR rs920778, rs1899663, rs4759314, and rs12826786 polymorphisms and BC in a sample of the Iranian population.
Materials and methods
Patients
A total of 220 women diagnosed with BC and 231 healthy women as controls were enrolled in this case-control study in Zahedan, southeast Iran. The study design and enrollment procedure have been defined in advance.26–29 Local ethics committee of Zahedan University of Medical sciences approved the project (IR.ZAUMS.Rec.1395.103), and informed consent was obtained from cases and controls. Venous blood from each participant was taken in ethylenediaminetetraacetic acid (EDTA) containing tube. Genomic DNA was extracted by salting out method and then stored at −20°C until use.
Genotyping
Genotyping of HOTAIR rs920778, rs1899663, rs4759314, and rs12826786 polymorphisms was determined by polymerase chain reaction–restriction fragment length polymorphism (PCR-RFLP) method. The primers are listed in Table 1. Each 0.20 mL of PCR reaction tube contained 1 µL of genomic DNA (~100 ng/mL), 1 µL of each primer, 8 µL of master mix, and 6 µL of double-distilled water (ddH2O). The PCR conditions for each SNPs were set as follows: 95°C for 6 min; 30 cycles of 95°C for 30 s; 60°C for rs920778, 66°C for rs1899663, 58°C for rs4759314, and 68°C for rs12826786 for 30 s; 72°C for 30 s; and a final extension step of 72°C for 5 min. A volume of 10 µL of PCR product is subsequently digested by an appropriate restriction enzyme (Table 1) and electrophoresed onto agarose gels, containing 0.5 µg/mL of ethidium bromide. It was then visualized under ultraviolet (UV) light (Figure 1). To confirm the genotyping quality, we randomly regenotyped approximately 20% of all samples and the genotypes were 100% concordant.
The primers used for detection of HOTAIR polymorphisms using PCR-RFLP methods.

HOTAIR: HOX transcript antisense intergenic RNA; PCR-RFLP: polymerase chain reaction–restriction fragment length polymorphism; F: forward; R: reverse.
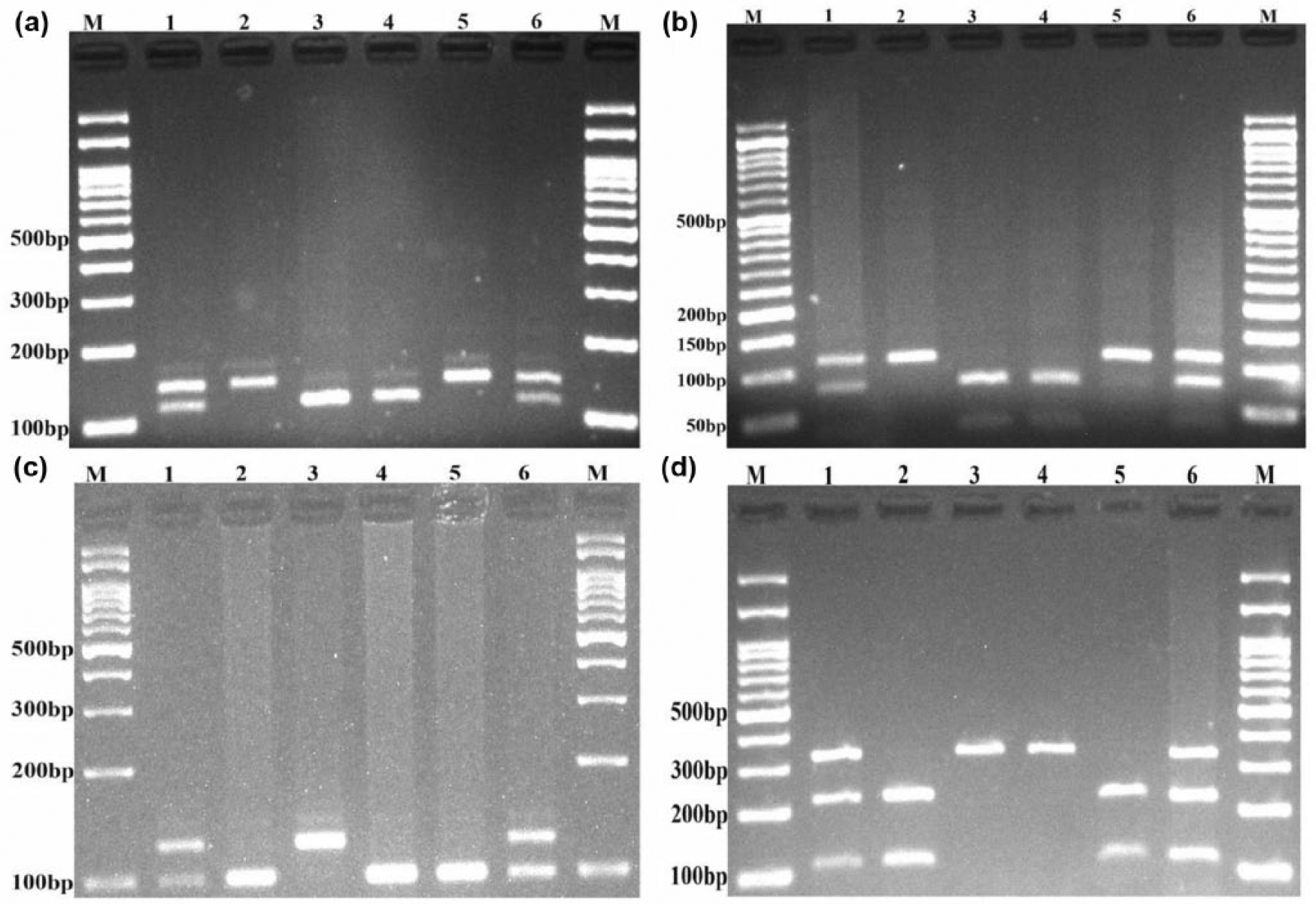
Electrophoresis pattern of polymerase chain reaction–restriction fragment length polymorphism (PCR-RFLP) for detection of HOTAIR (a) rs920778 T>C, (b) rs1899663 G>T, (c) rs4759314 A>G, and (d) rs12826786 C>T) polymorphisms. (a) For rs920778 T>C—lanes 1 and 6: TC; lanes 2 and 5: TT, and lanes 3 and 4: CC. (b) For rs1899663 G>T—lanes 1 and 6: GT; lanes 2 and 5: TT; and lanes 3 and 4: GG. (c) For rs4759314 A>G—lanes 1 and 6: AG; lanes 2, 4, and 5: AA; and lane 3: GG. (d) For rs12826786 C>T—lanes 1 and 6: CT; lanes 2 and 5: AA; and lanes 3 and 4: CC. M: DNA marker.
Statistical analysis
All statistical analysis was performed by software. The data were analyzed by independent sample t test and χ2 test. The logistic regression analysis was employed to calculate the odds ratios (ORs) and 95% confidence intervals (95% CIs). Haplotype analysis was performed using SNPSStats software. 30 p value <0.05 was considered to be statistically significant. Power of the study was calculated by using STATA 10 software, which is shown in Table 2.
Genotypic and allelic frequencies of HOTAIR variants in BC women.

HOTAIR: HOX transcript antisense intergenic RNA; BC: breast cancer; OR: odds ratio; CI: confidence interval.
Results
The study group comprised 220 pathologically confirmed BC patients, with mean age of 48.1 ± 10.7 years, and 230 healthy women, with mean age of 48.5 ± 11.4 years. No significant difference was found between the groups regarding age (p = 0.692).
Genotypic and allelic frequencies of HOTAIR variants in BC patients and controls are shown in Table 2. The findings suggest that rs920778 T>C variant of HOTAIR gene significantly increased the risk of BC; codominant (OR = 2.23, 95% CI = 1.50–3.33, p < 0.0001, TC vs TT; OR = 7.58, 95% CI = 3.32–17.34, p < 0.0001, CC vs TT), dominant (OR = 2.64, 95% CI = 1.79–3.88, p < 0.0001, TC + CC vs TT), recessive (OR = 4.92, 95% CI = 2.22–10.91, p < 0.0001, CC vs TT + TC), overdominant (OR = 1.60, 95% CI = 1.10–2.32, TC vs TT + CC), log-additive, and allelic (OR = 2.22, 95% CI = 1.67–2.94, p < 0.0001, C vs T) inheritance models were utilized. The rs1899663 G>T variant significantly decreased the risk of BC in codominant (OR = 0.18, 95% CI = 0.11–0.29, p < 0.0001, GT vs GG), dominant (OR = 0.19, 95% CI = 0.12–0.32, p < 0.0001, GT + TT vs GG), overdominant (OR = 0.20, 95% CI = 0.12–0.31, GT vs GG + TT), log-additive (OR = 0.38, 95% CI = 0.25–0.56, p < 0.0001), and allelic (OR = 0.61, 95% CI = 0.47–0.80, p < 0.001, T vs G) models. Furthermore, the rs12826786 T>C polymorphism of HOTAIR gene was significantly associated with protection against BC in codominant, dominant, recessive, overdominant, log-additive, and allele inheritance model tests (Table 2). Our findings did not support an association between rs4759314 A>G polymorphism and risk/protection against BC.
We performed haplotype analysis, and the findings showed that TGAC, CTAT, and TTAT haplotypes significantly decreased the risk of BC compared with rs920778T/rs1899663G/rs4759314A/rs12826786T haplotype (Table 3).
Haplotype analysis of HOTAIR gene polymorphisms and their association with breast cancer risk.

HOTAIR: HOX transcript antisense intergenic RNA; BC: breast cancer; OR: odds ratio; CI: confidence interval.
Association between HOTAIR gene polymorphisms and clinicopathological characteristics of BC patients such as age, tumor size, histologic type, lymph node involvement, grade, stage, estrogen receptor (ER) status, progesterone receptor (PR) status, and human epithelial growth factor receptor 2 (HER2) status is presented in Table 4. The findings showed that rs1899663 and rs4759314 variants were not associated with any clinicopathological characteristics, while the rs920778 and rs12826786 variants were only significantly associated with ER status (Table 4).
Association between HOTAIR gene polymorphisms and clinicopathological characteristics of BC patients.
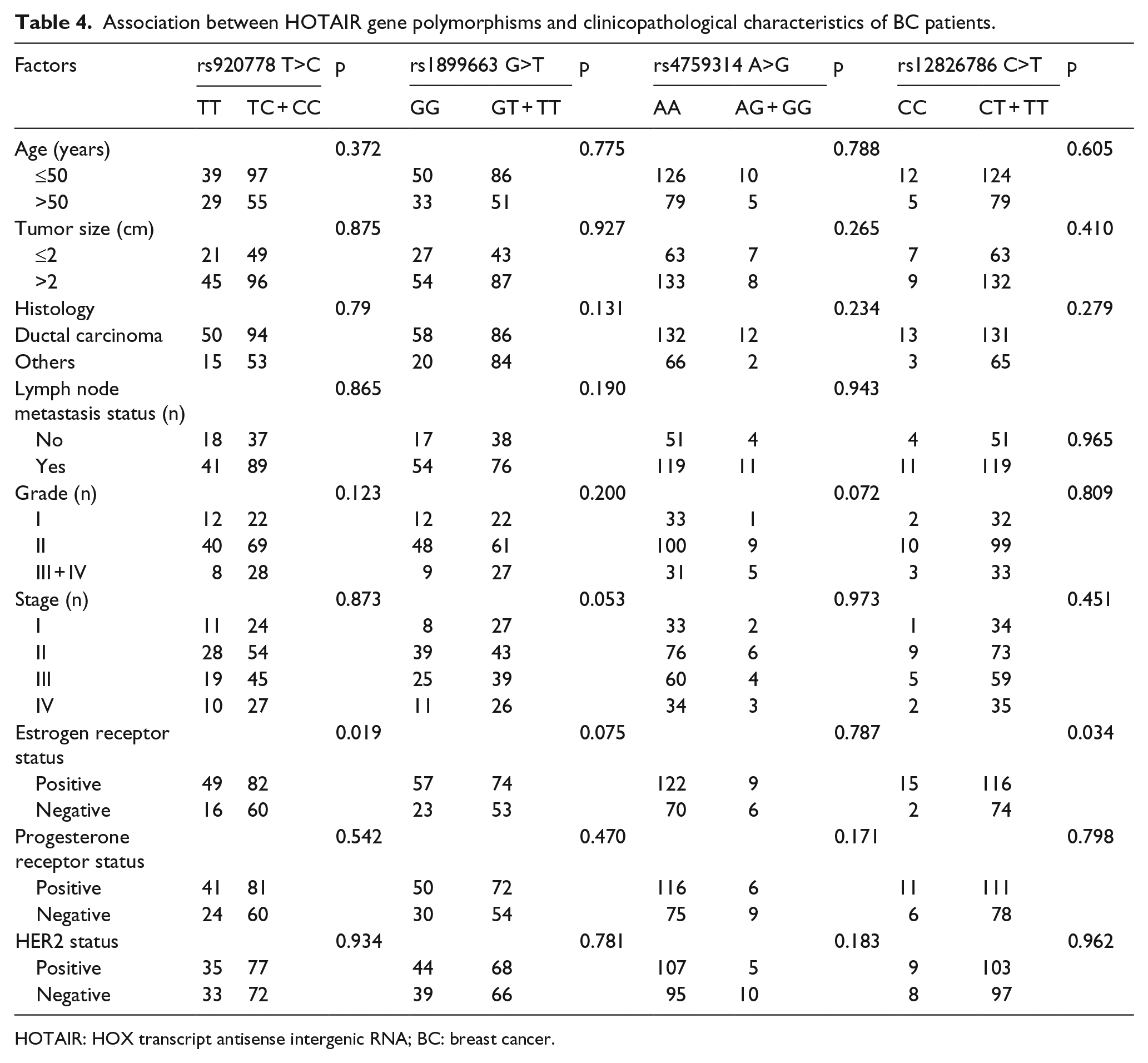
HOTAIR: HOX transcript antisense intergenic RNA; BC: breast cancer.
Discussion
As a major class of RNAs, lncRNAs are RNA molecules that are not translated into proteins. They play an important role in malignancy.6,7,10,31,32 HOTAIR, a trans-acting lncRNA, acts as a molecular scaffold by recruiting polycomb repressive complex 2 (PRC2) 14 and lysine-specific demethylase 1 (LSD1) complexes 14 to target genes which result in H3K27 methylation and H3K4 demethylation leading to epigenetic silencing of related genes. Aberrant expression of HOTAIR in various cancers indicates its contribution in their development. 33
Genetic variations could affect the function and expression of the relevant genes.34,35 Here, we tested the impact of HOTAIR polymorphisms on BC risk in a sample of southeast Iranian female population. We found that rs920778 and rs12826786 polymorphisms were significantly associated with increased BC risk; rs1899663 polymorphism displayed significant negative association; and the rs4759314 variant showed no association with BC risk.
Consistent with our findings, Bayram et al. 36 reported that rs12826786 C>T polymorphism increased the risk of BC in the Turkish population. Stratification by clinicopathological characteristics showed that the rs12826786 variant was significantly associated with advance stage, larger tumor size, presence of distant metastasis, and poor histological grade in BC patients. 36 In another study, Bayram et al. 17 found that rs920778 variant significantly increased the risk of BC in the Turkish population. They indicated that HOTAIR rs920778 polymorphism was associated with advanced stage, larger tumor size, presence of distant metastasis, and poor histological grade in BC patients. 17 Yan et al. 19 investigated the impact of rs1899663, rs4759314, and rs920778 polymorphisms of HOTAIR gene on BC risk in the Chinese population. Their findings indicated that the T allele of rs920778 variant was a risk factor for predisposition to BC. Haplotype analysis showed that rs1899663 G, rs4759314 A, and r920778 T (GAT) displayed positive association with BC risk. The rs1899663 and rs475314 variants displayed no association with BC risk in the Chinese population. 19 BC risk reduction was limited to subgroups of age: menarche >14 and number of pregnancies >2 for GT + TT rs1899663 and menopause ≤50 for AG + GG rs4759314. In our study, we did not observe significant association between rs1899663 and rs4759314 variants and clinicopathological characteristics of BC patients. The rs920778 and rs12826786 variants were only significantly associated with ER status.
The findings of the recent meta-analysis 23 suggest a correlation between rs920778 and rs7958904 polymorphisms of HOTAIR gene with overall cancer risk. Another meta-analysis performed by Chu et al. 24 revealed that HOTAIR rs920778 variant increases susceptibility to cancer. Their findings did not support an association between rs1899663 and rs4759314 variants and the risk of cancer. A meta-analysis performed by Ge et al. 25 showed that HOTAIR rs7958904 variant significantly decreased and rs920778 significantly increased the risk of cancer. No significant association between cancer susceptibility and rs874945, rs4759314, and rs1899663 polymorphisms was found.
HOTAIR functions as an oncogene. The overexpression of HOTAIR has been observed in various cancers, including BC,14,37–39 thyroid cancer, 40 hepatocellular carcinoma,41,42 cervical cancer,43,44 pancreatic cancer,45,46 and colorectal cancer.47,48 It has been proposed that the overexpression of HOTAIR in BC reduces the apoptotic rate while increasing cell growth, migration, and invasion.15,16 It has been recently proved that HOTAIR functions as an oncogene, overexpression of which induces invasiveness of cancer cell lines in vitro and initiates tumor growth and metastasis in lung model in vivo. 14 It has been reported that HOTAIR depletion in cancer cells causes a significant reduction in their invasiveness of cells that express a high level of polycomb proteins (PRC2). 14 These findings suggest that ncRNA-mediated targeting of polycomb complexes is possibly a determining step in breast tumorigenesis. Specifically, HOTAIR is involved in modification and regulation of epigenesis in the cells via targeting chromatin-modifying complex occupancy/localization/enzymatic activity in trans. HOTAIR is found at high levels in ER-positive (ER+) patients; it might even serve as a marker. 39 17β-estradiol (E2) increases HOTAIR expression in ER+ BC. In patients with triple-negative BC (TNBC), E2 binds to G-protein coupled estrogen receptor 1 (GPER), causing decreased expression of mir-148a. Mir-148a normally reduces the expression of HOTAIR. Therefore, reduction in mir-148 expression will be accompanied by an increase in HOTAIR expression. 49
There are few important limitations in our study including numbers of SNPs that we have investigated for the HOTAIR gene and limitation about the lack of the information regarding the known risk factors for BC (e.g. parity, menstrual patterns, hysterectomy, oral contraceptive or hormone therapy use, breastfeeding, smoking, and alcohol intake).
In conclusion, our findings suggest that HOTAIR variants were associated with BC risk in a sample of the Iranian population. They could serve as useful genetic biomarkers for the estimation of BC susceptibility and prognosis. Additional studies with larger sample sizes and different ethnicities are required to further confirm our results.
Footnotes
Acknowledgements
The authors wish to acknowledge deputy of research of Zahedan University of Medical Sciences for supporting dissertation grant. The authors also acknowledge Amir Vosoughi, University of Manitoba, Faculty of Science for English editing of the manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by a dissertation grant (MSc thesis of SHZ #7831) from Zahedan University of Medical Sciences.
