Abstract
HOTTIP (HOXA transcript at the distal tip), a newly discovered cancer-associated long non-coding RNA, appears to serve as a novel potential prognostic biomarker for patients with different types of cancers. However, most studies reported that the prognostic role of HOTTIP in cancers are limited by sample size and discrete outcomes. Therefore, a meta-analysis was performed to investigate the prognostic value of HOTTIP expression in various cancers. A total of 800 patients from nine studies were included in this meta-analysis. The results found that elevated HOTTIP expression predicted a poor outcome for overall survival in seven types of cancer (pooled hazard ratio = 2.11, 95% confidence interval = 1.51–2.71) with no heterogeneity (I2 = 0.0%, p = 0.879). Furthermore, subgroup analysis showed a more significant positive relationship between HOTTIP and overall survival in the studies with digestive system cancers (hazard ratio = 2.07, 95% confidence interval = 1.45–2.68), Chinese subjects (hazard ratio = 2.34, 95% confidence interval = 1.66–3.02), and sample size >100 subjects (hazard ratio = 2.36, 95% confidence interval = 1.21–3.51). In addition, the results found that the level of HOTTIP expression was positively associated with tumor size (relative ratio = 1.55, 95% confidence interval = 1.32–1.82, p < 0.001), tumor–node–metastasis stages (relative ratio = 1.81, 95% confidence interval = 1.51–2.16, p < 0.001), lymph node metastasis (relative ratio = 1.20, 95% confidence interval = 1.05–1.37, p = 0.01), and distant metastasis (relative ratio = 2.53, 95% confidence interval = 1.55–4.15, p < 0.001). In conclusion, our results suggested that elevated HOTTIP expression was correlated with poor clinical outcomes and may serve as a novel potential biomarker for patients’ prognosis in various cancers.
Introduction
Cancer has already become one of the leading causes responsible for millions of deaths each year worldwide.1,2 Despite current advances in cancer therapy, cancer patients have comparatively poor prognosis due to frequent cancer metastasis, recurrence, and a lack of curative treatment.2,3 Early diagnosis and treatment remain the most effective ways to improve clinical outcomes of cancer patients. 3 Thus, it is essential to identify more and better tumor molecular biomarkers to enhance the accuracy and utility of cancer prediction models.
Recently, numerous studies have focused on the involvement of long non-coding RNAs (lncRNAs) in the development and progression of various cancers.4–7 LncRNAs are a novel group of evolutionarily conserved non-protein coding RNAs that are longer than 200 nucleotides in length. 8 Accumulating evidence has suggested that most lncRNAs are dysregulated in a variety of cancers and involved in diverse cancer-related processes, such as modulation of cell proliferation and apoptosis, drug resistance, tumor invasion, and metastasis.9–11 Among these cancer-related lncRNAs, a new lncRNA termed “HOXA transcript at the distal tip” (HOTTIP) has drawn increasing attention.12,13
HOTTIP, a functionally characterized antisense lncRNA, is located at the 5′-end of the HOXA gene cluster (chromosome 7p15.2) that encodes the 3764 bp transcript.14,15 HOTTIP directly interacts with the adaptor protein WDR5 and targets the WDR5/MLL complex to drive H3K4 methylation and thus transcriptionally activate the distal of HOXA gene. 14 HOTTIP has been functionally characterized as one of the most familiar onco-lncRNAs in various cancers. The aberrant expression of HOTTIP has been found in different types of cancers, including hepatocellular carcinoma (HCC),16,17 colorectal cancer (CRC), 18 gastric cancer (GC), 19 pancreatic cancer (PC), 20 osteosarcoma, 21 tongue squamous cell carcinoma (TSCC), 22 and breast cancer (BC). 23 Furthermore, HOTTIP might act as a novel effective prognostic biomarker and a therapeutic target for cancers. A recent study revealed that overexpression of HOTTIP correlated with larger tumor size, deeper invasion depth, positive lymph node metastasis (LNM), advanced tumor–node–metastasis (TNM) stages, and shorter overall survival (OS) in GC patients. 19 Quagliata et al. 16 found that high HOTTIP expression was associated with metastasis formation and poor survival for HCC patients. Li et al. 24 suggested that the upregulation of HOTTIP expression is markedly correlated with advanced clinical stage, distant metastasis, and poor prognosis of patients with osteosarcoma. Increased HOTTIP expression was reported to be positively correlated with T stage, clinical stage, and distant metastasis and played as an unfavorable prognosis predictor for TSCC patients. 22 Together, HOTTIP may act not only as a novel prognostic biomarker but also as a potential therapeutic target in human cancers.
Although a significant relationship between HOTTIP expression and clinical outcomes was found in a variety of cancers, most studies reported that the prognostic role of HOTTIP in cancers is limited by sample size and discrete outcomes. Therefore, we conducted a systematic review and quantitative meta-analysis to investigate the clinical value of HOTTIP as a prognostic biomarker in various cancers.
Materials and methods
Search strategy
We searched potentially eligible studies though PubMed, EMBASE, Web of Science, Ovid, and Cochrane library databases for eligible articles on the prognostic value of HOTTIP expression in cancers from inception until February 2017. The following search terms were included: “HOTTIP,” “HOXA transcript at the distal tip,” “lincRNA,” “lncRNA,” “long intergenic noncoding RNA,” “noncoding RNA,” “cancer,” “carcinoma,” “neoplasm,” “outcome,” “prognosis,” and “survival.” The reference lists of the retrieved articles were searched manually. The literature search was performed by two independent researchers and was limited to the English language.
Inclusion and exclusion criteria
The studies were considered eligible if they met the following criteria: any type of human cancer was studied, studies investigating the prognostic role of HOTTIP expression in cancers, the levels of HOTTIP expression in cancerous tissues were detected, patients were grouped according to the levels of HOTTIP expression, a link between HOTTIP expression and clinicopathological parameters was included, studies providing sufficient data to estimate the hazard ratios (HRs) with corresponding 95% confidence interval (CI) for OS, and studies were published in English. Exclusion criteria were as follows: letters, editorials, expert opinions, case reports, and reviews; studies only investigated the molecular structure and functions of HOTTIP; studies without usable data for further analysis; and duplicate publications.
Data extraction
Two investigators (S.-K.Z. and Y.Z.) independently evaluated and extracted the data from each eligible study. A consensus was achieved for disagreements by a third investigator (Y.Z.). According to the above inclusion and exclusion criteria, the following elements were extracted: first author, publication date, country, tumor type, TNM stages, sample size, cutoff value, follow-up period, detection method, survival analysis methodology, HRs with corresponding 95% CIs for OS and disease-free survival (DFS), and other clinicopathological parameters, including age, gender, tumor size, differentiation, TNM stages, LNM, and distant metastasis. HRs with corresponding 95% CIs were preferentially extracted from univariate or multivariate analyses. If these were not available, the HR estimates were calculated from Kaplan–Meier survival curves using Engauge Digitizer V4.1 (http://sourceforge.net).
Quality assessment
The quality of included studies was assessed based on the Newcastle–Ottawa Scale (NOS). The NOS criteria applies a “star” rating system, which ranges from 0 to 9 stars for the judgment of methodological quality based on selection (0–4 stars), comparability (0–2 stars), and outcome (0–3 stars). 25 Based on previous recommendations, studies with 5 points were considered to be of high quality. The two aforementioned investigators independently evaluated the quality of each study. Conflicting evaluations or inconsistent data were resolved through discussion with the third investigator (Y.Z.).
Statistical analyses
All data analyses were performed with STATA statistical software version 14.0 (Stata Corporation, College Station, TX, USA). Pooled HRs with 95% CIs were used to estimate the prognostic role of HOTTIP in various cancers. Pooled relative ratios (RRs) with 95% CIs were used to estimate the association between HOTTIP expression and clinicopathological characteristics.
Results
Study selection and characteristics
A total of 37 relevant articles were identified based on electronic databases, including PubMed, EMBASE, Web of Science, Ovid, and the Cochrane library. First, 1 duplicate article was excluded. After careful title and abstract screening, 20 irrelevant articles were excluded. After being reviewed in detail, 7 full-text articles were excluded due to insufficient data and non-comparative studies. Finally, 9 eligible studies were included in this meta-analysis (Figure 1).

The flow diagram of the meta-analysis.
The characteristics of the included studies are summarized in Table 1. A total of 800 patients from eight studies between 2014 and 2017 were included. The regions of these studies included Switzerland (n = 1) and China (n = 8). Among these studies, the sample size ranged between 48 and 156, and three studies enrolled more than 100. Seven types of cancers were recorded, including HCC (n = 2), osteosarcoma (n = 1), CRC (n = 2), pancreatic ductal adenocarcinoma (n = 1), TSCC (n = 1), GC (n = 1), and BC (n = 1). The levels of HOTTIP expression were detected by quantitative reverse transcription polymerase chain reaction (qRT-PCR) in all included studies. All HRs with the corresponding 95% CIs for OS and DFS were extracted from the original data, including three studies that calculated Kaplan–Meier curves.
The main characteristics of the included studies in the meta-analysis.
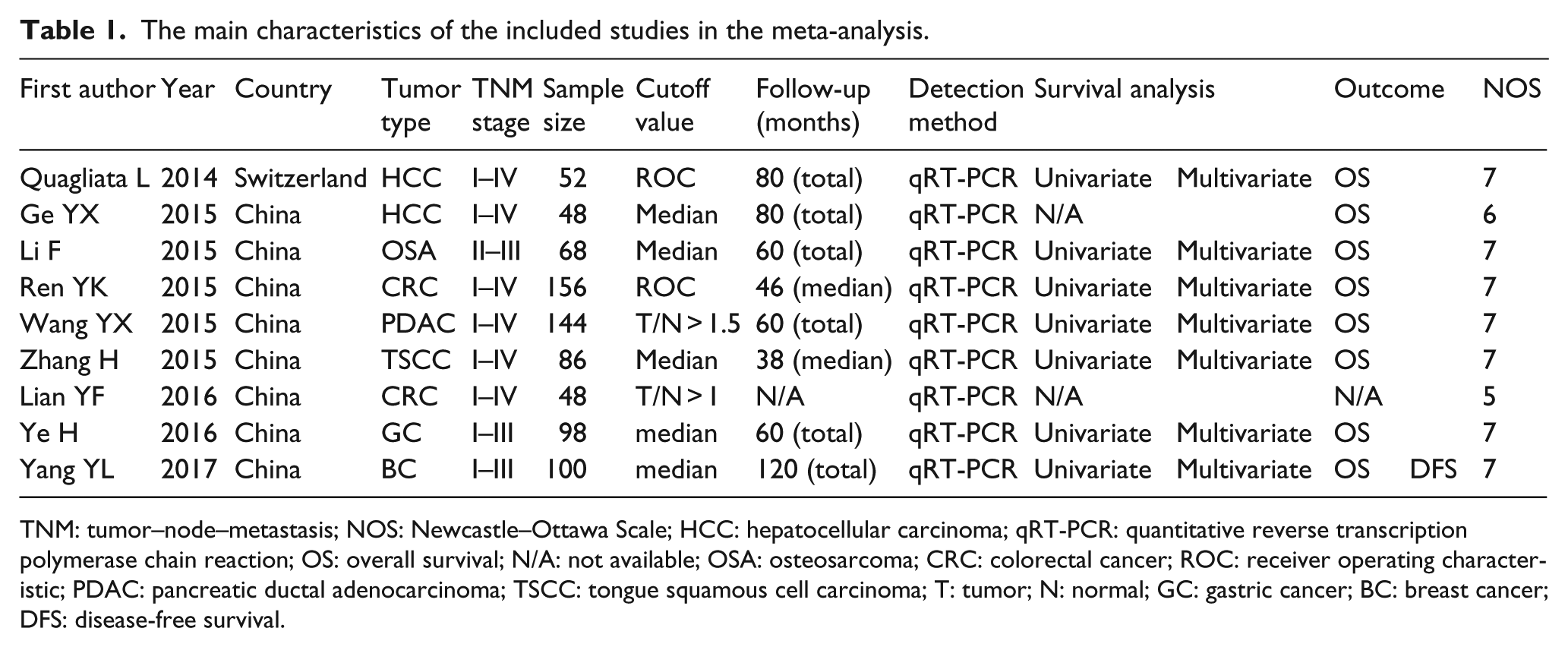
TNM: tumor–node–metastasis; NOS: Newcastle–Ottawa Scale; HCC: hepatocellular carcinoma; qRT-PCR: quantitative reverse transcription polymerase chain reaction; OS: overall survival; N/A: not available; OSA: osteosarcoma; CRC: colorectal cancer; ROC: receiver operating characteristic; PDAC: pancreatic ductal adenocarcinoma; TSCC: tongue squamous cell carcinoma; T: tumor; N: normal; GC: gastric cancer; BC: breast cancer; DFS: disease-free survival.
Association between HOTTIP expression and OS in various cancers
A total of eight studies, including 752 patients, were recruited to assess the effects of HOTTIP overexpression on OS in various cancers. The results found that elevated HOTTIP expression predicted a poor outcome for OS in seven types of cancers (pooled HR = 2.11, 95% CI = 1.51–2.71; Figure 2). There appeared to be no heterogeneity between the HRs of HOTTIP among these studies (
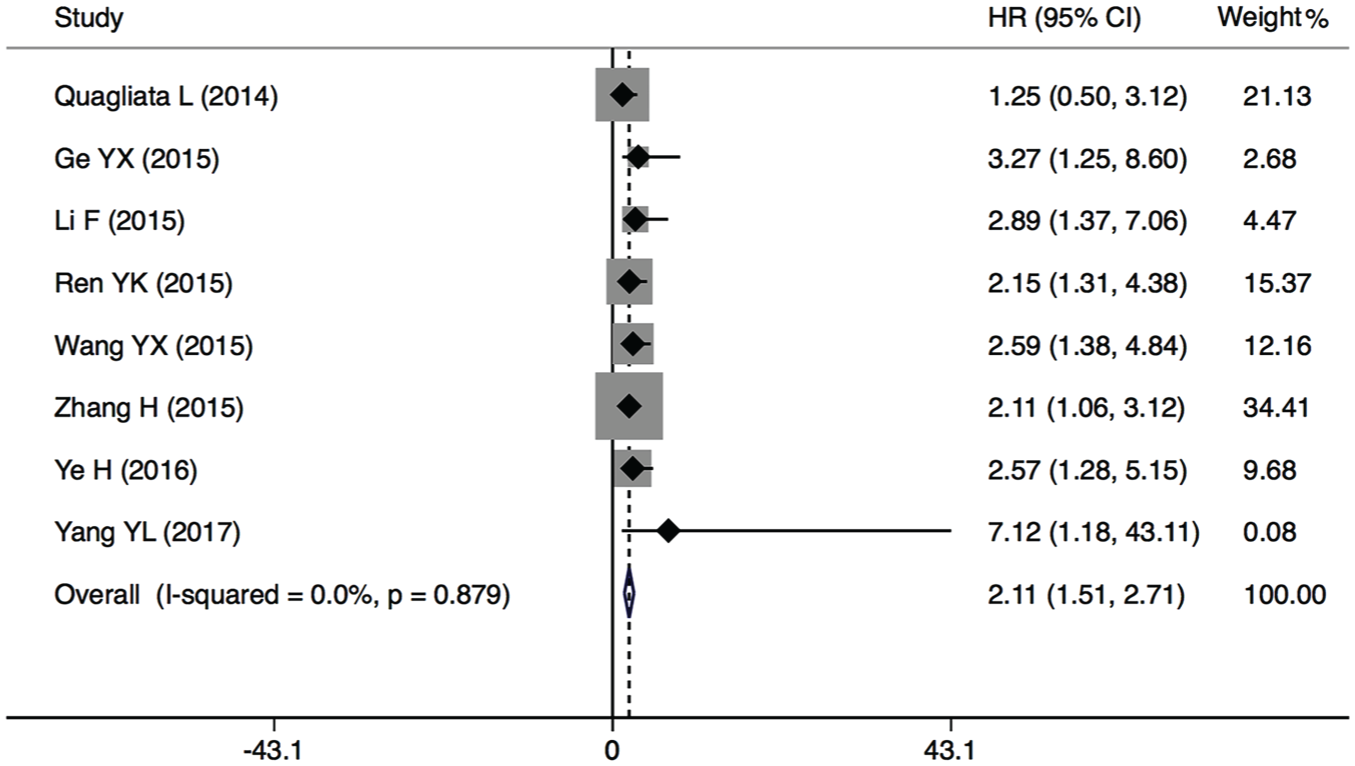
Forest plots of the included studies evaluating the correlation between HOTTIP expression and OS.
Association between HOTTIP and the clinicopathological characteristics of cancers
The association between HOTTIP expression and clinicopathological characteristics was examined in eight studies with seven types of cancer (Table 2). To study the association between HOTTIP expression and tumor size, seven studies, reporting a total of 700 cancer patients, were included. Analysis revealed that a higher HOTTIP expression was predictive of a larger tumor size (pooled RR = 1.548, 95% CI = 1.320–1.815,
Correlation between HOTTIP expression and clinicopathological characteristics of cancers.
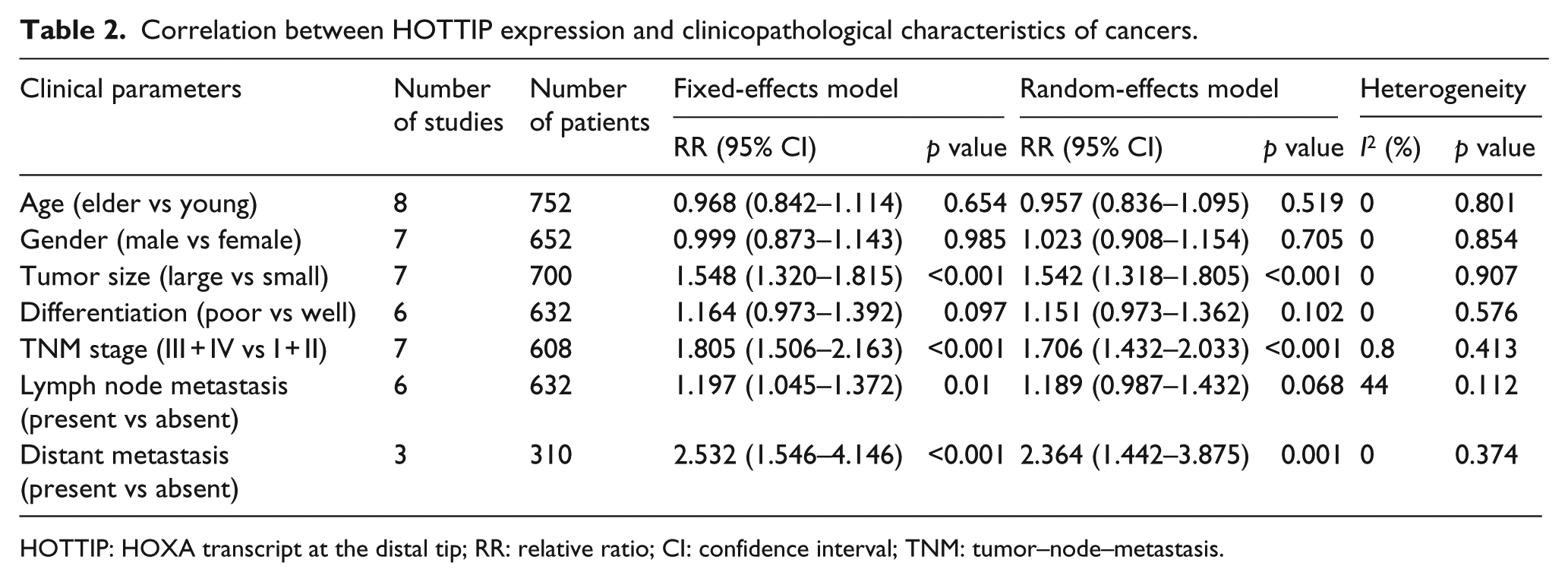
HOTTIP: HOXA transcript at the distal tip; RR: relative ratio; CI: confidence interval; TNM: tumor–node–metastasis.

Forest plots of the included studies evaluating the correlation between HOTTIP expression and clinicopathological characteristics including (a) tumor size, (b) TNM stages, (c) lymph node metastasis, and (d) distant metastasis.
Publication bias and sensitivity analysis
To evaluate the publication bias in this meta-analysis, Begg’s funnel plots and Egger’s linear regression tests were applied. Both plots were also asymmetrically inverted funnels (Figure 4(a) and (b)). Subsequently, the results of Egger’s linear regression test revealed that there was no publication bias for the analysis of OS (
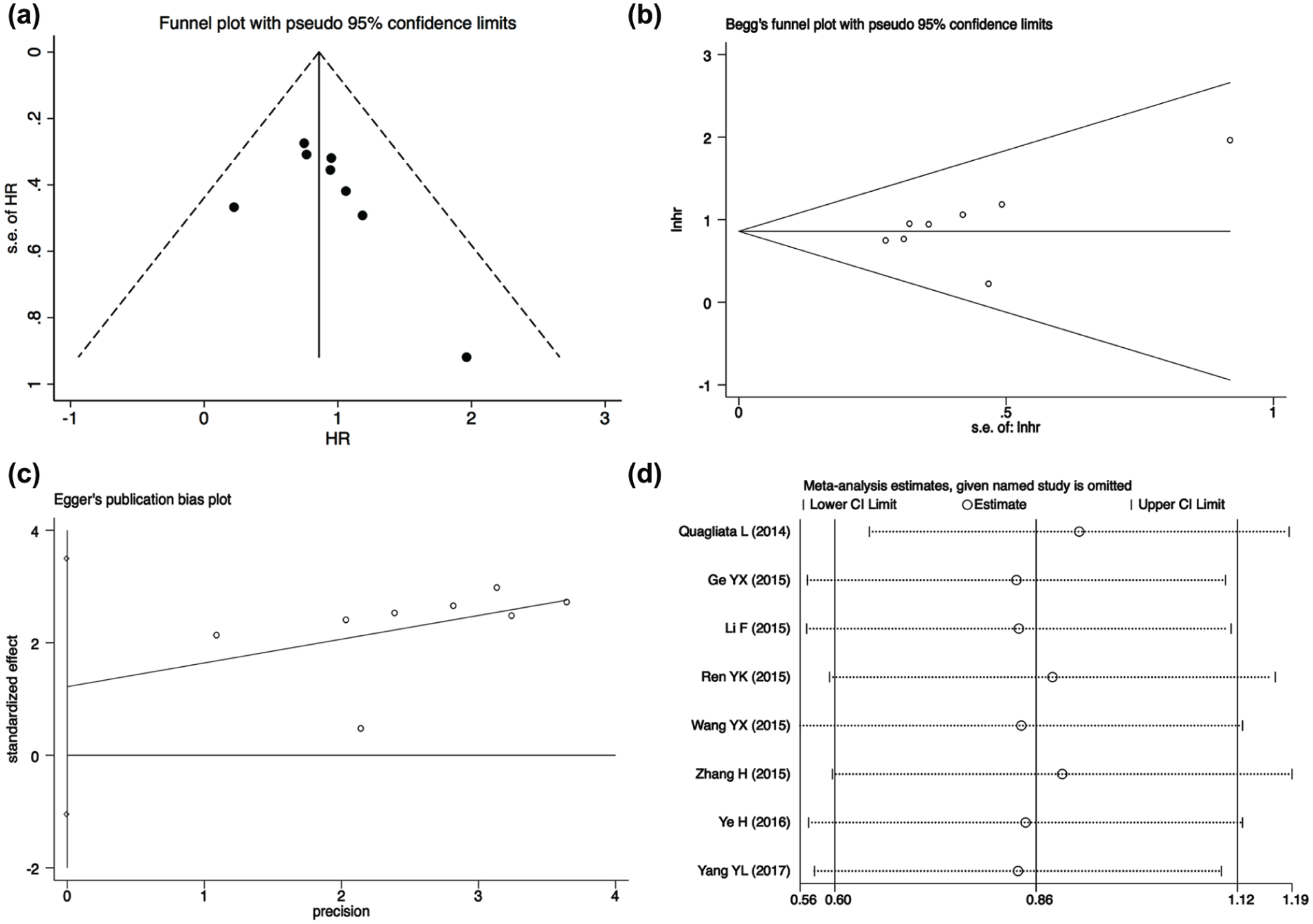
Publication bias and sensitivity analysis in the meta-analysis. (a) Funnel plots of the included studies for overall survival. (b) Begg’s funnel plots of the included studies for overall survival. (c) Egger’s funnel plots of the included studies for overall survival. (d) Sensitivity analysis of the included studies for overall survival (HR: hazard ratio; CI: confidence interval; SE: standard error).
Discussion
As a novel molecular basis, many studies have focused on the involvement of lncRNAs in the development and the progression of various tumors, providing a new insight into cancer therapeutic strategy. 11 Recent studies have demonstrated that lncRNAs may play an important regulatory role in chromatin modification, X-chromosome inactivation and transcription, genetic imprinting, dosage compensation, and the regulation of protein activity and RNA alternative splicing, which may substantially contribute to cancer development.4,26
HOTTIP, regarded as one of the most familiar oncogenic lncRNAs, was aberrantly expressed in different types of cancers.12,27 HOTTIP may play important roles in tumor growth, invasion, and metastasis.12,24,28–30 Li et al. 31 revealed that HOTTIP could promote progression and gemcitabine resistance by regulating HOXA13 in PC. Cheng et al. 32 found that silencing of HOTTIP expression decreased PC cell proliferation, induced apoptosis, and decreased migration. HOTTIP silencing also resulted in cell proliferation arrest by altering cell cycle progression and impaired cell invasion by inhibiting epithelial–mesenchymal transition in esophageal squamous cell carcinoma. 33 Furthermore, the knockdown of HOTTIP expression impaired CRC cell proliferation and induced apoptosis, which was partly dependent on repressing p21 expression. 34 In addition, recent studies found that miR-125b may serve as a posttranscriptional regulator of HOTTIP in HCC, and the loss of miR-125b expression might contribute to the frequent upregulation of HOTTIP. 35 Intriguingly, Ge et al. 17 identified that HOTTIP acted as a novel target of miR-192 and miR-204 and HOXA genes.
Recent studies found that the upregulation of HOTTIP expression was correlated with poor prognosis of patients with nearly all types of cancer. However, perplexity and inconsistency arise from a wide range of studies due to heterogeneity. A meta-analysis of nine studies, including 800 patients, was undertaken to assess the prognostic role of HOTTIP expression in several types of cancers and provided sufficient evidence for the association between HOTTIP expression and clinicopathological characteristics of cancers. In this meta-analysis, we found that elevated HOTTIP expression predicted a poor outcome for OS in seven types of cancer, indicating that HOTTIP may be a promising prognostic biomarker for cancer patients. Moreover, subgroup analysis was also conducted to investigate the association between HRs and these variables, including cancer type, country, and sample size. Stratified analysis indicated that abnormal HOTTIP expression was strongly correlated with poor survival in studies with digestive system cancers, Chinese patients, or sample size larger than 100. In addition, we also examined the association between HOTTIP expression and the clinicopathological characteristics of cancers. The results demonstrated that HOTTIP expression was significantly associated with clinicopathological parameters, including tumor size, TNM stage, LNM, and distant metastasis. Furthermore, both Begg’s and Egger’s tests revealed no significant publication bias regarding the prognostic role of HOTTIP expression in cancer. Therefore, this meta-analysis provided convincing evidence that HOTTIP may be used as a negative, unfavorable prognostic biomarker for various cancers.
Some limitations of this meta-analysis should be acknowledged due to the discrete data across these clinical studies. The criteria for calculating the cutoff value were not the same in different studies. The inclusion of studies that only reported HR or survival curves might have resulted in the potential for selection bias. Furthermore, some of the HRs were calculated by reconstructing survival curves rather than directly obtained from the primary studies; thus, calculation bias might be present. In addition, the inclusion of a relatively small number of studies in different regions (Switzerland and China) might have decreased the applicability of our results across different ethnicities. Data collection may be incomplete because data from non-English language articles were not included. In this meta-analysis, only summarized data rather than individual patient data were used. Therefore, it is possible that our results might overestimate the prognostic effects of abnormal HOTTIP expression on survival in different types of cancers. We believe that more clinical studies should be conducted to evaluate the potential prognostic role of HOTTIP in other types of cancers that have not been included.
Conclusion
In conclusion, this meta-analysis showed that elevated HOTTIP expression was significantly associated with poor survival in cancer patients and might act as a novel effective prognostic biomarker and therapeutic target for cancers. Furthermore, the functional role of HOTTIP in the regulation of cell proliferation, apoptosis, and metastasis suggested that HOTTIP may play a key role in tumor progression. Thus, HOTTIP may potentially be used as a novel biomarker for predicting cancer patient survival. More clinical studies on other different types of human cancers that have not yet been investigated need to be conducted.
Footnotes
Acknowledgements
S.K-.Z. and Y.Z. conceived and designed the study, collected and analyzed the data, and wrote the manuscript. All authors reviewed and approved the final manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by a grant from the National Natural Science Foundation of China (No. 81402029).
