Abstract
The aberrant dysregulation of taurine upregulated 1, a novel discovered long non-coding RNA, was ubiquitous in different human solid tumors. Accumulating researches have indicated that taurine upregulated 1 is an independent prognostic indicator in cancer patients. This investigation aimed to further explore the prognostic and clinical significance of taurine upregulated 1 in various types of cancers. Eligible studies were systematically searched in PubMed, Embase, Medline, and Web of Science databases. A total of 12/14 studies with 1303/1228 individuals were included to evaluate the association of taurine upregulated 1 with overall survival and clinicopathological features by pooled hazard ratio and odds ratio in malignancies. The meta-analysis suggested overexpression of taurine upregulated 1 was significantly correlated with unfavorable overall survival in patients with cancer (pooled hazard ratio = 1.63, 95% confidence interval: 1.29–2.06). There was also a significantly positive correlation between high level of taurine upregulated 1 and high pathological grade carcinoma (pooled odds ratio = 4.41, 95% confidence interval: 3.07–6.43) and positive lymphatic metastasis (pooled odds ratio = 2.00, 95% confidence interval: 1.31–3.06). In summary, upregulated taurine upregulated 1 is correlated with more advanced clinicopathological characteristics and poor prognosis, suggesting that taurine upregulated 1 may serve as a novel predictive biomarker of patients with numerous tumors.
Introduction
With the development of high-throughput sequencing technology, over 90% non-coding RNAs (ncRNAs) were certificated through large-scale mapping of transcriptomes.1,2 Long non-coding RNAs (LncRNAs) are one of major group regulatory ncRNAs longer than 200 nucleotides including intergenic lncRNAs (lincRNAs) and circular RNAs (circRNAs).3,4 Increasing evidence have illustrated lncRNAs play important roles in transcription factor and coactivator recruitment, chromatin remodeling, histone modification, pre-mRNA splicing, molecular sponge, and scaffold during transcriptional and post-transcriptional progression of carcinogenesis and cancer aggression.5–7
Taurine upregulated 1 (TUG1) located on 22q12.2 covering four exons encodes three transcript variants which was initially identified through a genomic scanning in
Moreover, overexpression of TUG1 is associated with high risk of poor prognosis and pathological outcome of cancer patients including non–small-cell lung carcinoma, bladder cancer, cervical cancer, esophageal squamous cell carcinoma, osteosarcoma, colorectal cancer, gastric cancer, and renal cell carcinoma. Therefore, we qualified and evaluated recent studies to explore the prognostic and clinicopathological role of TUG1 in patients with different types of tumors.
Materials and methods
Search strategy and study selection criteria
Literature searching was conducted in PubMed, Embase, Medline, and Web of Science databases (up to February 8, 2017) by the following searching strategy: “TUG1” OR “TUG-1” OR “taurine up-regulated 1” OR “TI-227H” OR “LINC00080” OR “NCRNA00080”) AND (“cancer” OR “tumor” OR “tumour” OR “carcinoma” OR “neoplasm” OR “adenoma” OR “sarcoma”). The retrieved references were excluded from duplicated first. Then, titles, abstracts, and full texts of potential retrieved studies were carefully scanned to eliminate irrelevant studies with the following eligibility criteria: (1) evaluated the relationship of dichotomous TUG1 level with prognosis and clinicopathological parameters of any type of cancer, (2) calculated hazard ratios (HRs) and 95% confidence intervals (CIs) for overall survival (OS), (3) were published in English, and (4) performed in laboratory studies (mechanistic or cell line studies) of human research.
Data extraction
To each study, we deliberatively extracted the following information: first author, journal, year of publication, country, ethnicity, specimen and carcinoma type, number of patients, detection method, cut-off value, follow-up, quality score, and HRs and 95% CIs for OS. Quality assessment was performed according to reporting recommendations for tumor marker prognostic studies (REMARK) guideline. 13 HR was, respectively, extracted from multivariable analysis, univariate analysis and estimated from graphical survival plots by Engauge Digitizer 4.1 software as described previously, if only Kaplan–Meier curve existed.14,15 Clinical parameters with dichotomous TUG1 levels also were retrieved such as pathological stage and lymph node metastasis (LNM).
Statistical analysis
This meta-analysis was conducted with Stata software version 12.0 (Stata Corporation, College Station, TX, USA). Odds ratios (ORs) with corresponding 95% CIs were performed to analyze the association of TUG1 expression with pathological stage and lymphatic metastasis, and HRs with corresponding 95% CIs were utilized to estimate the relationship strength between TUG1 expression and OS of patients. The pooled HRs were calculated by a random effect model (DerSimonian–Laird). The heterogeneity tests of combined HRs and ORs were performed by Cochran’s Q test and Higgins I-squared statistic (PH < 0.1 and I2 > 50%). To further explore the potential heterogeneity in different studies, a subgroup analysis was applied into the meta-analysis. Publication bias was evaluated by an Egger’s linear regression test (p < 0.05).
Results
Selection and characteristics of included studies
By screening PubMed, Embase, Medline, and Web of Science databases, a total of 12 eligible OS articles including 1303 tissue samples from 246 records published from 2013 to 2017 were enrolled upon inclusion and exclusion criteria (Figure 1). Nine cancer types were evaluated in this meta-analysis. The main characteristics of all OS studies are summarized in Supplementary Table 1. Simultaneously, in total, 14 available clinicopathological studies containing 1228 tissue specimens were collected with clinical stage and LNM data, which consist of 12 kinds of tumors, as shown in Supplementary Table S3. Quality score based on REMARK guideline ranged from 35% to 75%, as shown in Supplementary Table S2. The expression of TUG1 in patient groups was separated by inconsistent cut-off values due to different detecting methods. HRs with 95% CIs were extracted from five multivariate analysis and seven univariate analysis groups.
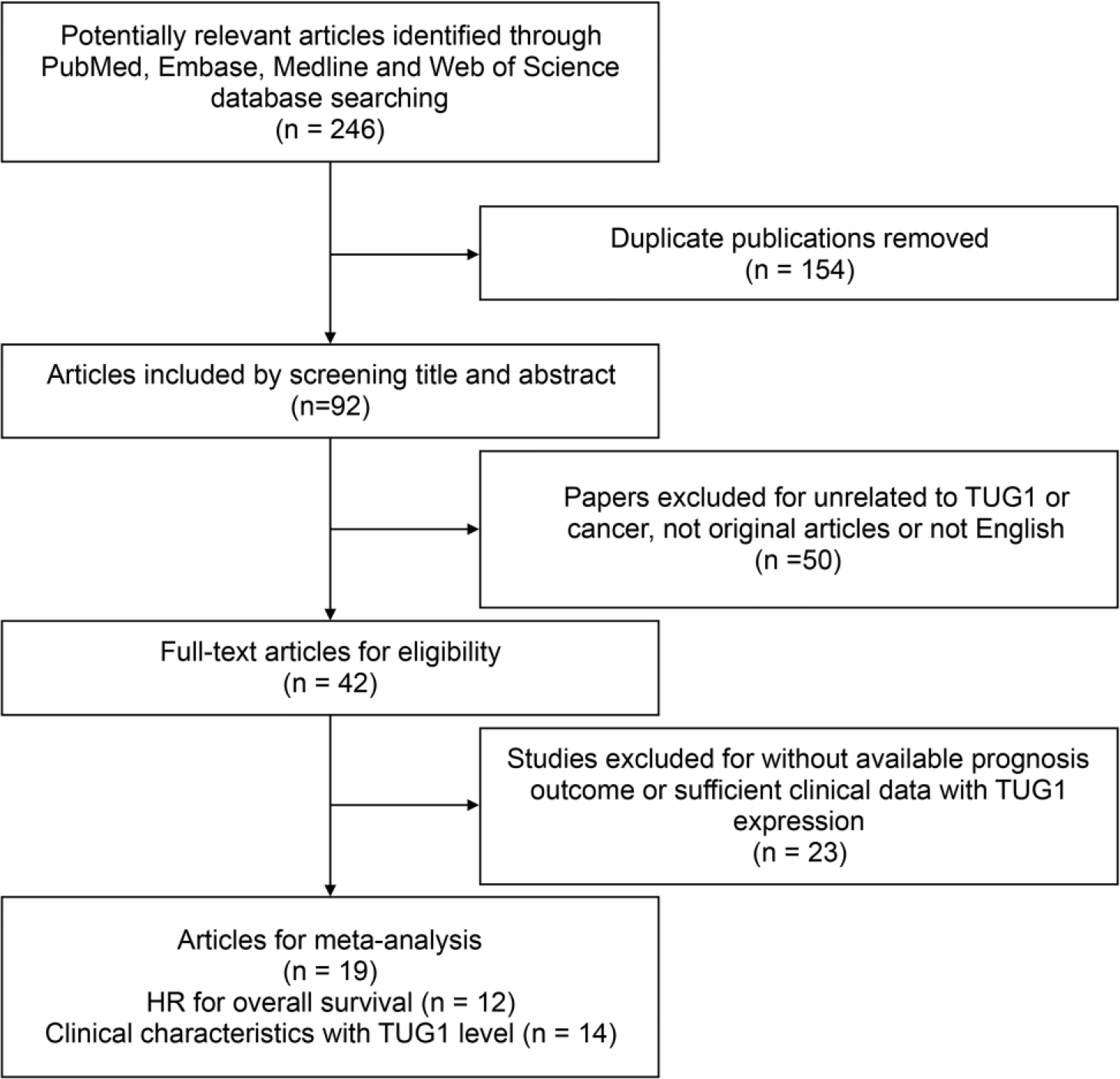
Flow diagram searching strategy and study selection progress in the meta-analysis.
Association between TUG1 expression and OS in human solid tumors
To assess the relationship of TUG1 expression with OS in cancer patients, a total of 1303 patients with HRs and 95% CIs were included. A significant correlation was found between high TUG1 level and unfavorable OS of patients (pooled HR = 1.63, 95% CI: 1.29–2.06), but an obvious heterogeneity existed across 12 studies under a random effect model (I2 = 87.8%, pH < 0.001), as shown in Figure 2(a).

Prognostic and clinical significance of TUG1 for cancer patients. (a) Forest plot of HR studies of TUG1 for OS in a random effect model. Each square represents a study and the center of which denotes the HR with a horizontal 95% CI lines. The diamond shows the combined overall HR. (b) Egger’s test for publication bias in OS analysis. Standard normal deviate (SND) is defined as HR divided by its standard error. (c) Forest plot of OR studies of TUG1 for cancer clinical stage in fixed and random effect models. (d) Forest plot of OR studies of TUG1 for lymph node metastasis (LNM) in fixed and random effect models.
The subgroup analysis in fixed and random effect models was conducted by cancer types, analysis methods, sample sizes, and quality scores for a subsequent investigation of potential heterogeneity in Table 1. In the subgroup of high-quality score (I2 = 34.1%, pH = 0.194), there was no statistical significance for heterogeneity test.
Subgroup analysis of HRs in different cancer type, analysis method, sample size, and quality score subgroups.

NSCLC: non–small-cell lung carcinoma; BDC: bladder cancer; CVC: cervical cancer; ESCC: esophageal squamous cell carcinoma; OSC: osteosarcoma; CRC: colorectal cancer; GC: gastric cancer; SCLC: small-cell lung cancer; RCC: renal cell carcinoma; HRs: hazard ratios.
Publication bias evaluated by Egger’s test indicated that there was evident asymmetry in this meta-analysis (pEgger’s = 0.023; Figure 2(b)). And subgroup analysis of publication bias showed that small-studies effects existed in multivariate analysis subgroup (pEgger’s = 0.022) in Supplementary Table S4. To further investigate the origin of publication bias, we conducted a contour-enhanced funnel plot, in which most of the estimated missing studies were in <5% area (Supplementary Figure S1). This result indicated that publication bias might originate from other factors such as heterogeneity. Sensitivity analysis showed an obvious asymmetry in pooled HR after removing Zhang et al.’s study (2016) in Supplementary Figure S2. Thus, publication bias mainly originated from heterogeneity instead of other factors.
Correlation of TUG1 level with advanced clinical characteristics in cancer patients
In 14 clinicopathological studies consisting of 1228 tumor samples with pathological grade and lymphatic metastasis, data were retrieved into OR analysis. We observed that high TUG1 expression was associated with high grade (pooled OR = 4.44, 95% CI: 3.07–6.43) and positive LNM (pooled OR = 2.00, 95% CI: 1.31–3.06), as shown in Figure 2(c) and (d).
The subgroup analysis stratified by cancer type uncovered the resource of heterogeneity under fixed and random effect models. There existed a significantly statistical heterogeneity across lymph node subgroup (I2 = 50.6%, pH = 0.040) in nine clinicopathological studies, as shown in Table 2. In lymphatic metastasis heterogeneity analysis, we detected a significant heterogeneity in digestive system subgroup (I2 = 71.0%, pH = 0.016). No evidence of statistical heterogeneity was detected in clinical grade stratification analysis (I2 = 46.4%, pH = 0.029) in Table 3. However, digestive system subgroup showed an obvious heterogeneity (I2 = 75.9%, pH = 0.002), as shown in Table 3.
Subgroup analysis of the association and heterogeneity between overexpression of TUG1 and lymph node metastasis.

CVC: cervical cancer; ESCC: esophageal squamous cell carcinoma; OC: ovarian cancer; BC: breast cancer; CRC: colorectal cancer; GC: gastric cancer; RCC: renal cell carcinoma; OSCC: oral squamous cell carcinoma; TUG1: taurine upregulated 1; OR: odds ratio; CI: confidence interval.
Subgroup analysis of the association and heterogeneity between overexpression of TUG1 and clinical stage.
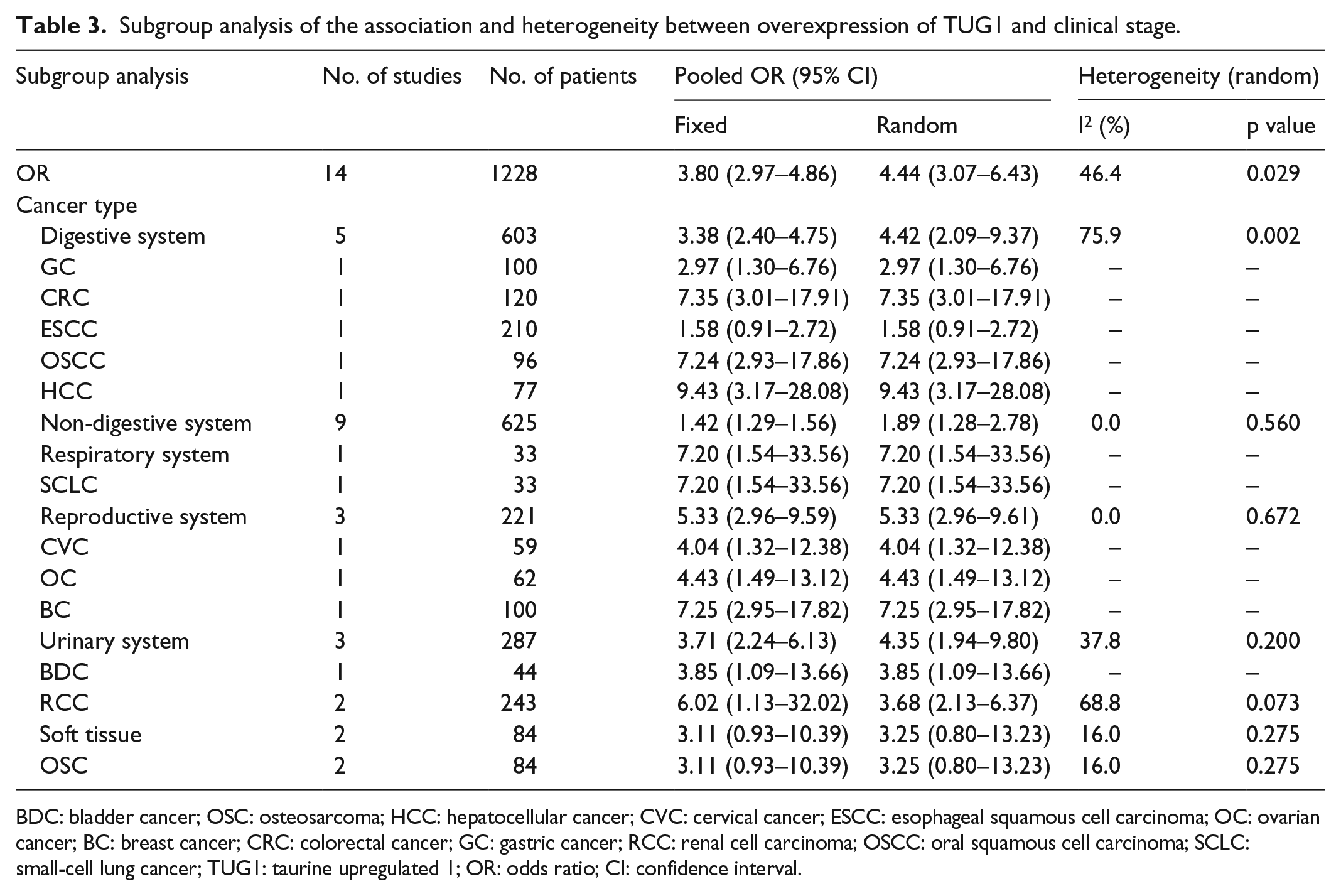
BDC: bladder cancer; OSC: osteosarcoma; HCC: hepatocellular cancer; CVC: cervical cancer; ESCC: esophageal squamous cell carcinoma; OC: ovarian cancer; BC: breast cancer; CRC: colorectal cancer; GC: gastric cancer; RCC: renal cell carcinoma; OSCC: oral squamous cell carcinoma; SCLC: small-cell lung cancer; TUG1: taurine upregulated 1; OR: odds ratio; CI: confidence interval.
Discussion
It has been estimated that approximately 3% protein-coding RNAs are transcribed by the eukaryotic genome compared with over 90% of ncRNAs used to be recognized as “biological noises” through the emerging next generation sequencing technology and large-scale transcriptome mapping.1,2 The enormous advances in these new technologies reflected the vital and complicated biological functions of lncRNAs in genetic and epigenetic regulation.6,7 By interaction with DNA, RNA, or proteins, lncRNAs account for cancer abnormal proliferation, potential invasion, replicative senescence, resistance to radiation and drugs, and reprogrammed energy metabolism.16,17 Recent accumulating studies indicate that lncRNAs act as diagnosis and prognosis biomarkers in human tumors due to tissue sensitivity and specificity, efficient and convenient determination in body fluids, and high stability. 18
Among all rising cancer-associated lncRNAs, TUG1has been documented in various malignancies.10–12,19–34 TUG1 via combination with polycomb repressive complex 2 (PRC2) trimethylates histone 3 (H3K27me3)repressed p15, Krüppel-like factor 2 (KLF2), and LIM-kinase 2b (LIMK2b) tumor suppressor genes in hepatocellular carcinoma, gastric cancer, and small-cell lung cancer.10,31,33 In addition, targeting TUG1 inhibited phosphoinositide 3-kinase (PI3K)/Akt, Notch, transforming growth factor β (TGF-β), and Wnt/β-catenin pathways in osteosarcoma, glioma, gallbladder carcinoma, and oral squamous cell carcinoma.12,35–37 TUG1 is also a key molecular sponge or competing endogenous RNA (ceRNA) for miR-144, miR-145, miR-26a, miR-9-5p, miR-34a-5p, miR-229, and miR-300 involved in radioresistance, carcinogenesis, invasion, angiogenesis, and blood–tumor barrier permeability.11,37–42 Although no direct evidence showed relationship between TUG1 and cachectic status, we suggest that TUG1 may be elevated in advanced clinical stage cancer patients due to cachexia. Furthermore, TUG1 was markedly increased in multiple sclerosis (MS), diabetes mellitus (DM), chronic kidney disease (CKD), and chronic obstructive pulmonary disease (COPD).43–46 TUG1 promoted peroxisome proliferator–activated receptor gamma (PPARG) coactivator 1α (PGC-1) transcription by binding to its promoter and improved mitochondrial bioenergetics in the podocytes of diabetic nephropathy and CKD.45,46 In mouse pancreas, enriched TUG1 facilitated β-cell proliferation, induced insulin secretion, and ameliorated pancreatic islet dysfunction.47,48
Here, we performed this meta-analysis to assess prognostic and clinicopathological significance of TUG1 for cancer patients. All of these results above suggest that elevated TUG1 may be regarded as an unfavorable predictor in different cancers (pooled HR = 1.63, 95% CI: 1.29–2.06). And the pooled clinicopathological studies also indicated that high TUG1 expression was significantly correlated with poor stage (pooled OR = 4.44, 95% CI: 3.07–6.43) and positive lymphatic metastasis (pooled OR = 2.00, 95% CI: 1.31–3.06).
Otherwise, we acknowledged that several potential limitations existed in this meta-analysis. First, more cancer types in large-scale and well-designed studies need to be included in analysis. Second, inconsistent cut-off values of TUG1 expression may result in some heterogeneity. Third, most of the HRs were indirectly obtained from survival curve calculation and differences of paper quality among studies, which may lead to bias in subgroup heterogeneous analysis. Finally, heterogeneity, only English papers and most of positive studies may generate publication bias. Thus, it may overestimate the predictive significance of TUG1 to some extent. In conclusion, the meta-analysis results suggest that TUG1 may serve as a candidate biomarker to predict unexpected prognosis and aggressive tumor progression of patients with cancer.
Footnotes
Acknowledgements
X.W., X.C., and D.Z. contributed equally to this work. S.Z. conceived and designed, coordinated, and directed this meta-analysis. X.W., X.C., and D.Z. searched the publications, analyzed the data, and wrote the manuscript. G.Y., D.Z., Z.Y., and Z.Y. extracted the data, judged and checked data synthesis results, and made figures and tables. All authors reviewed and approved the final manuscript.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The work was supported by Special Fund for Translational Research of Sino-Russia Medical Research Center in Harbin Medical University (CR201512 to S.Z.), Scientific Fund of Heilongjiang Province for Youth (QC2016124 to X.C.), and National Natural Science Foundation of China (81572482 to G.Y.).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
